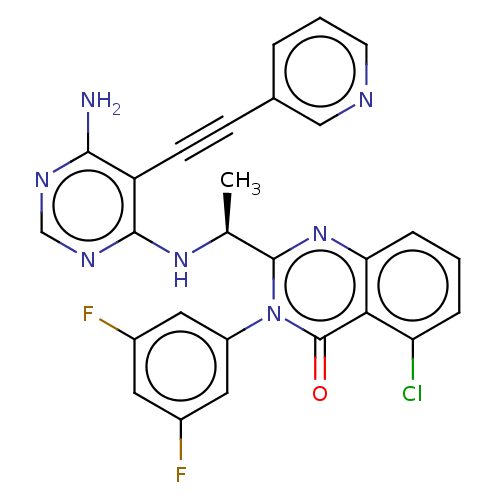
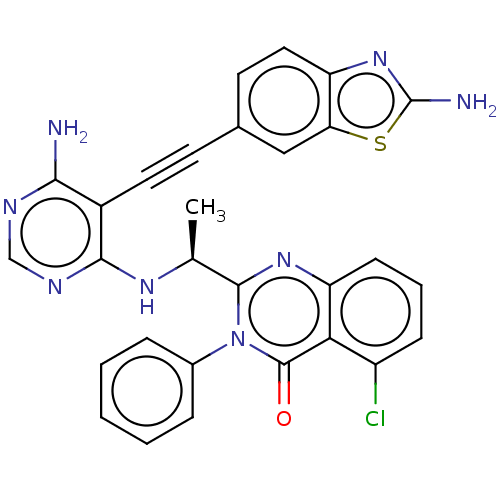
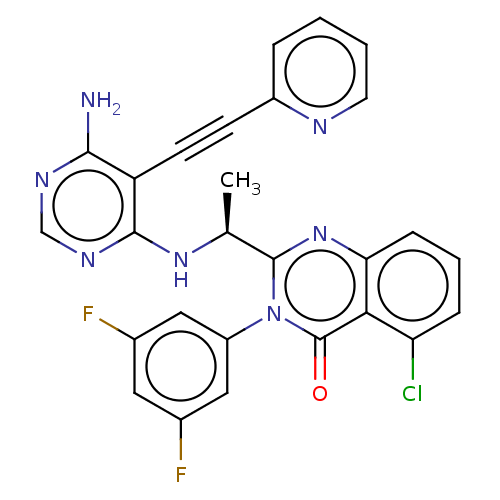
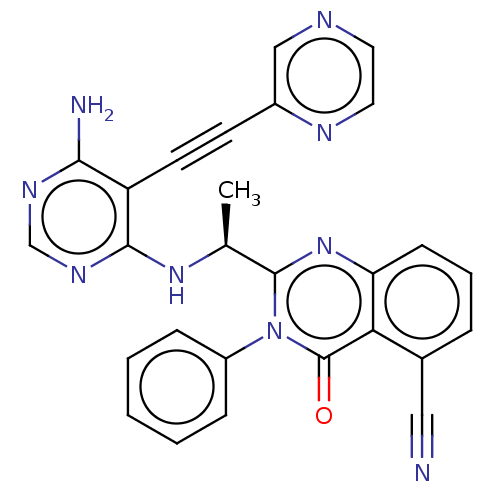
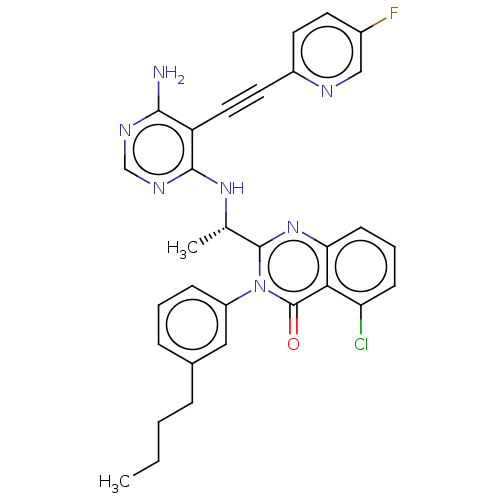
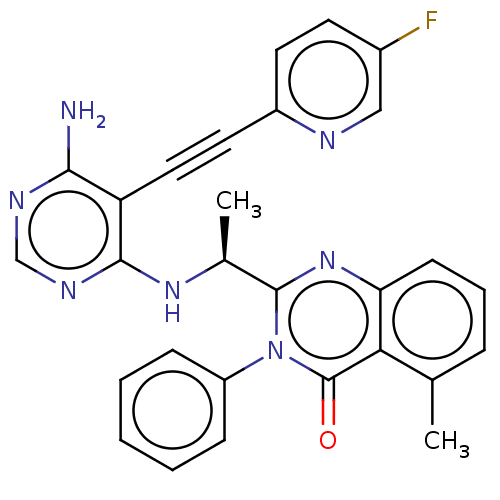
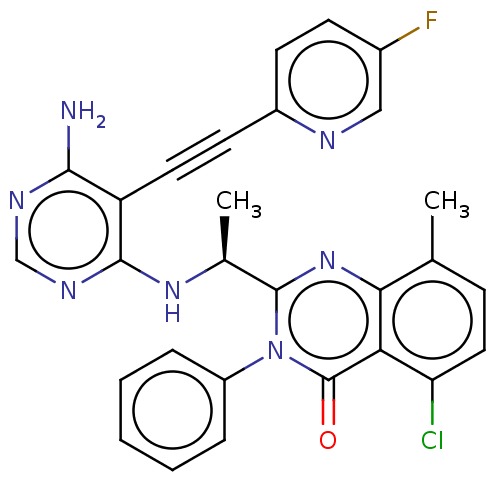
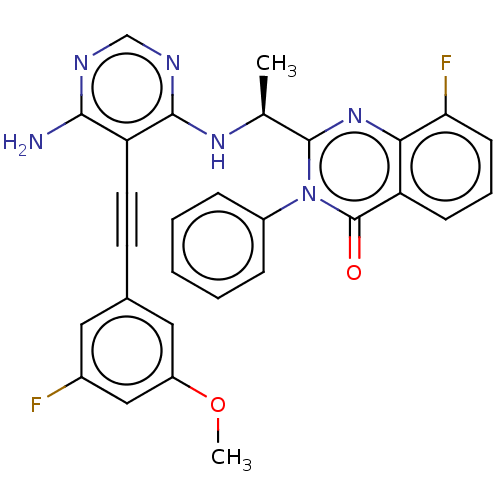
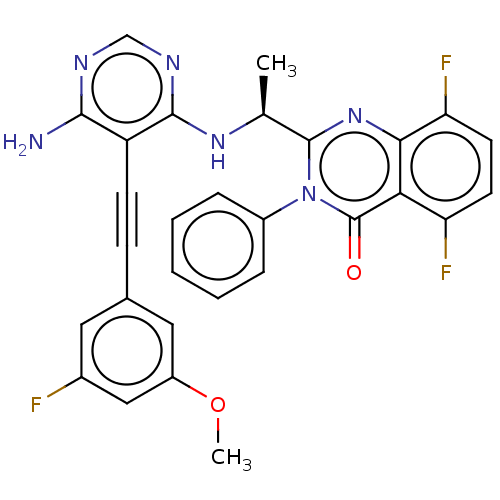
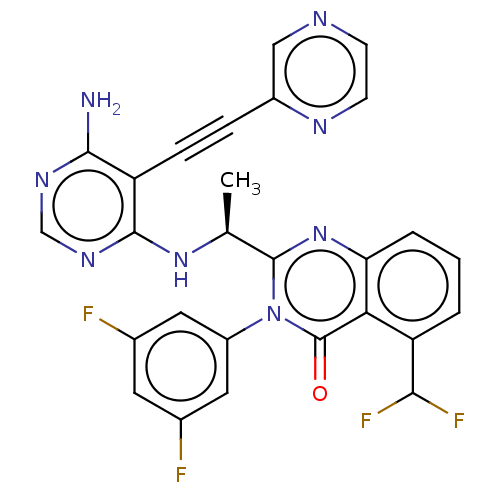
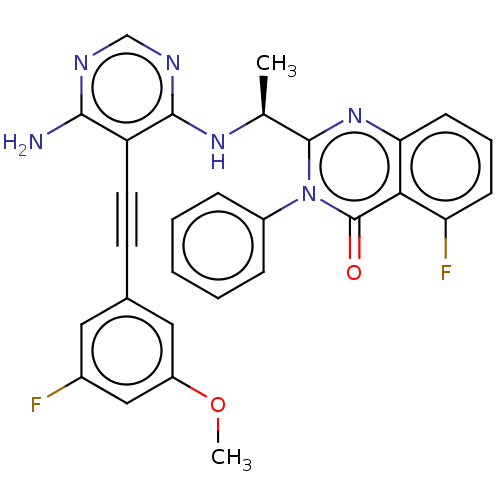
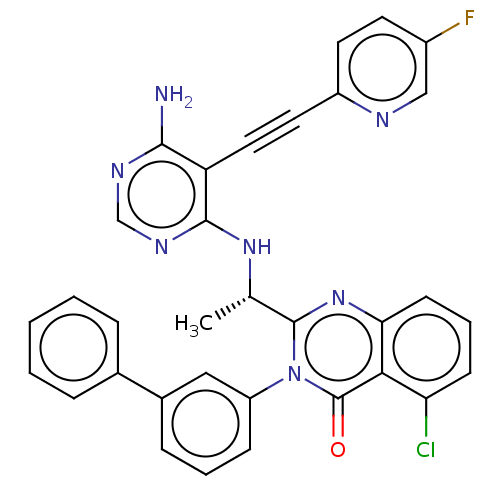
Report error Found 656 Enz. Inhib. hit(s) with all data for entry = 7653
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 1nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Gilead Calistoga
US Patent
Gilead Calistoga
US Patent
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:PI3K isoforms were assayed under initial rate conditions in the presence of 25 mM Hepes (pH 7.4), and 2xKm ATP (100-300 uM), 10 uM PIP2, 5% glycerol,...More data for this Ligand-Target Pair
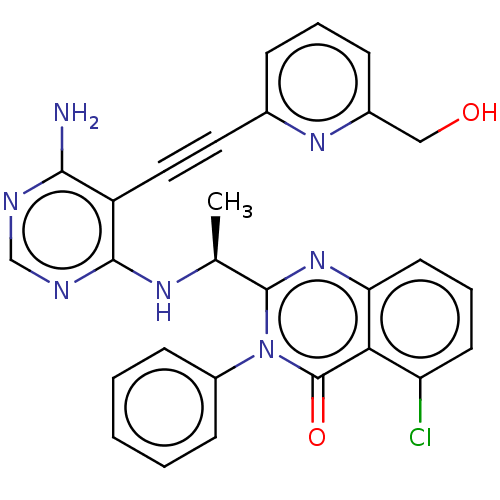
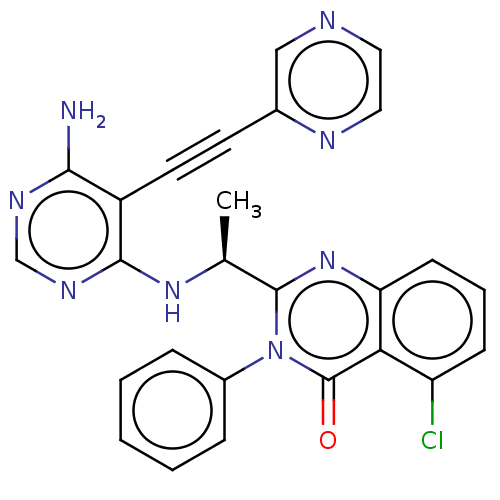
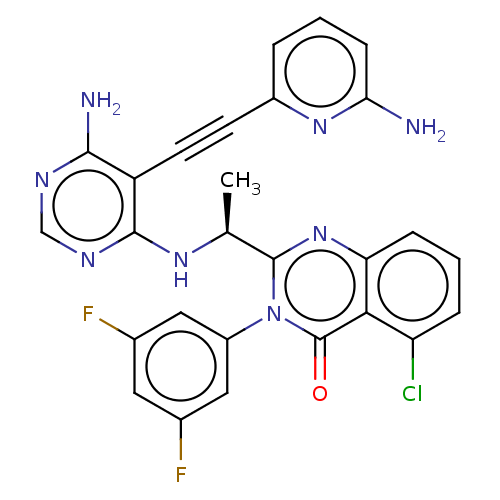

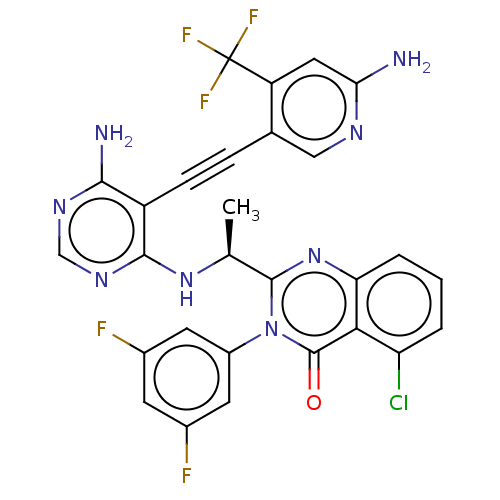
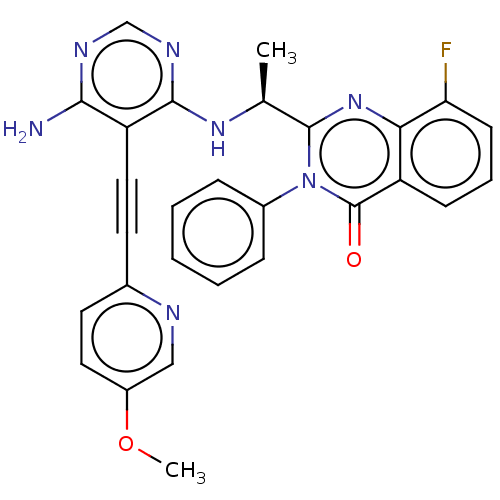
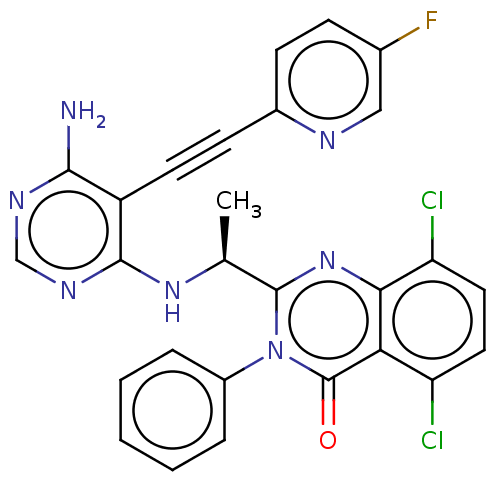
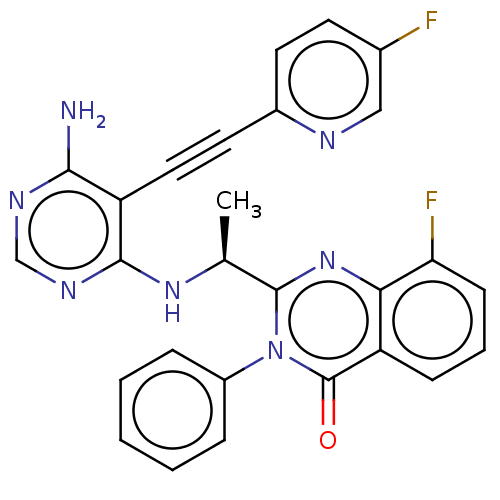
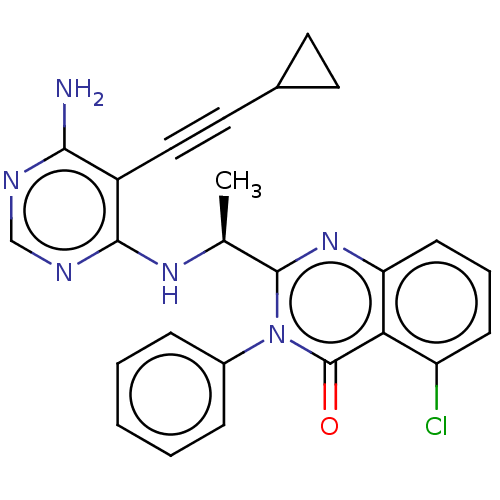
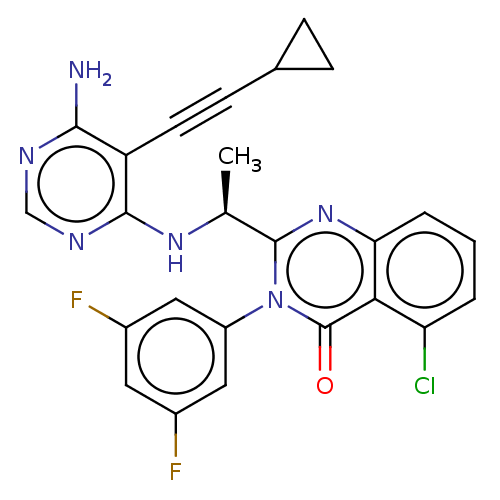
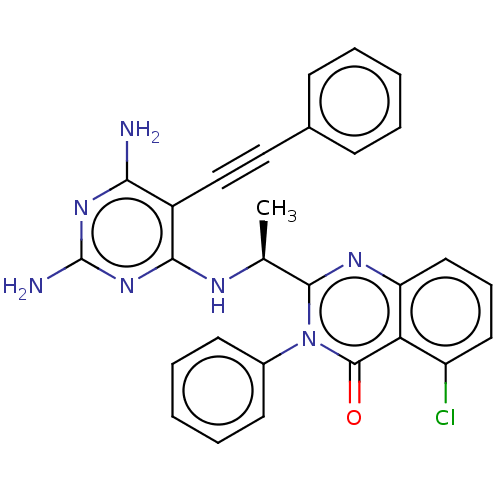
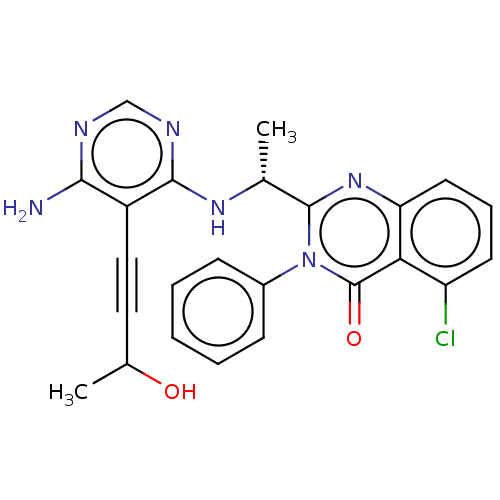
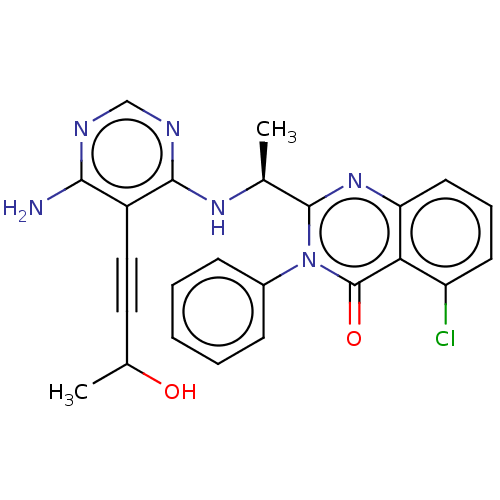
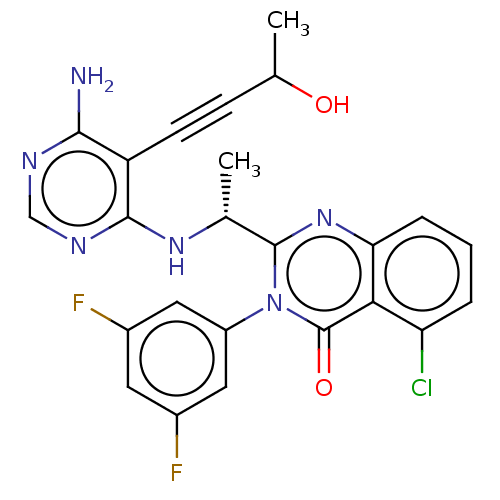
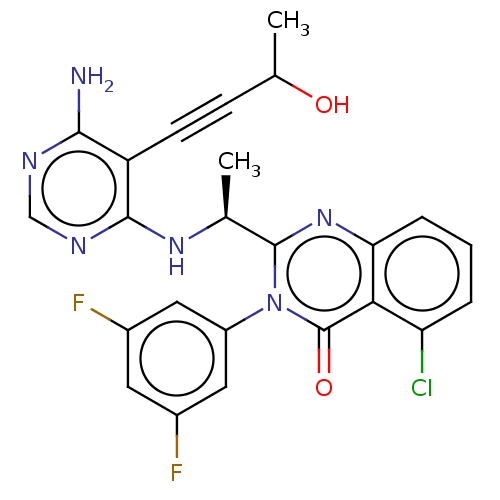
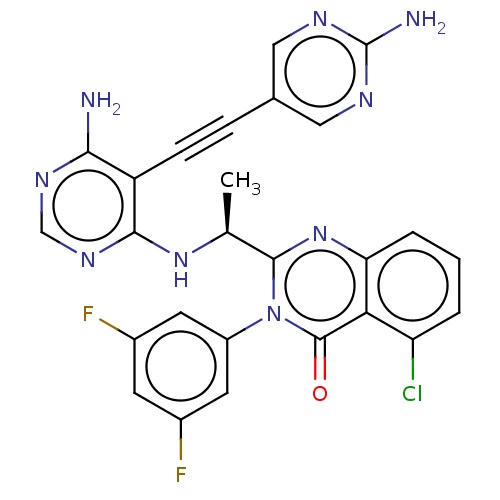
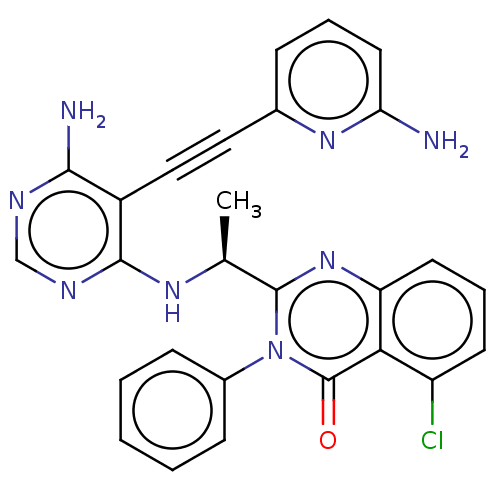
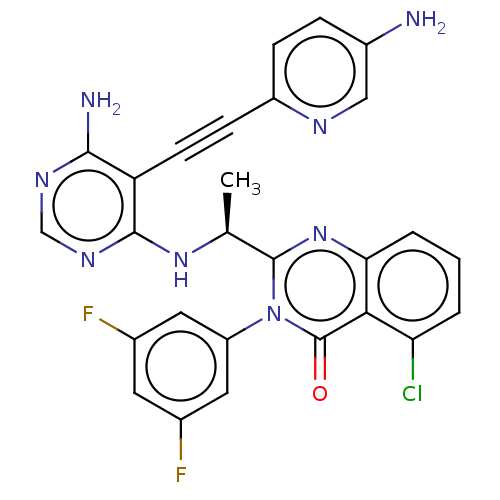
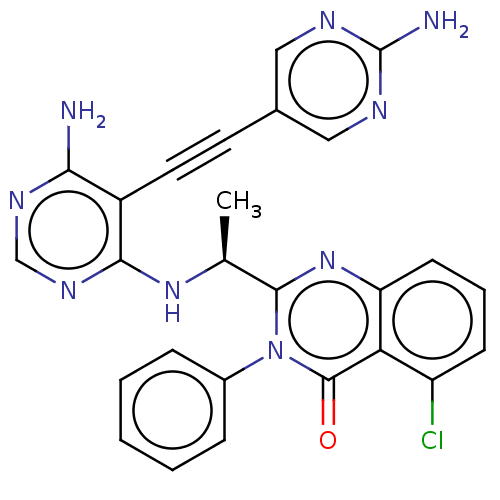
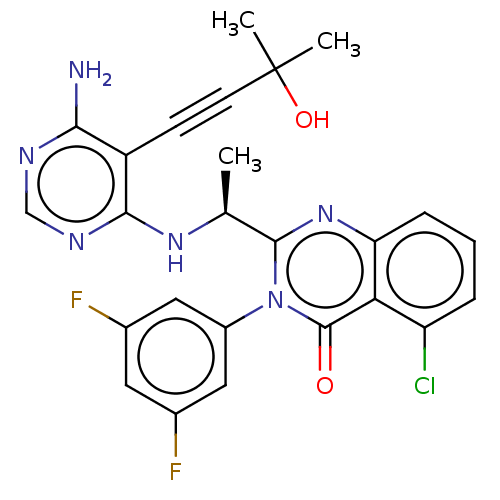

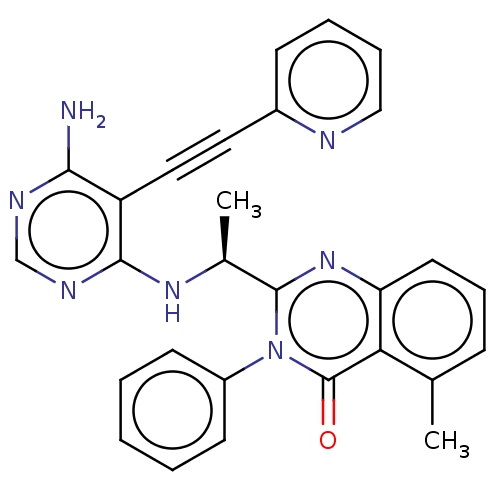
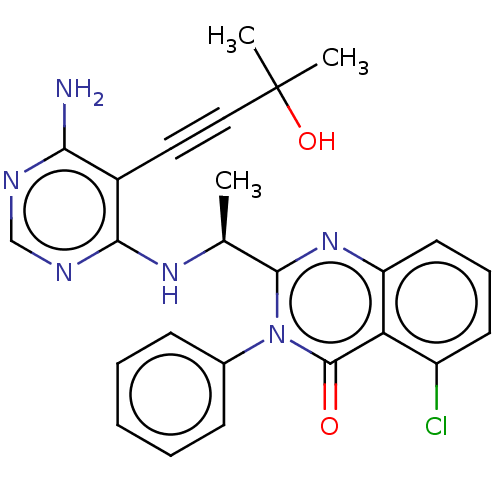
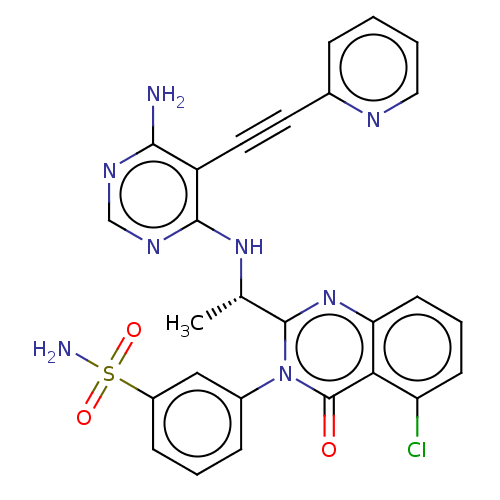
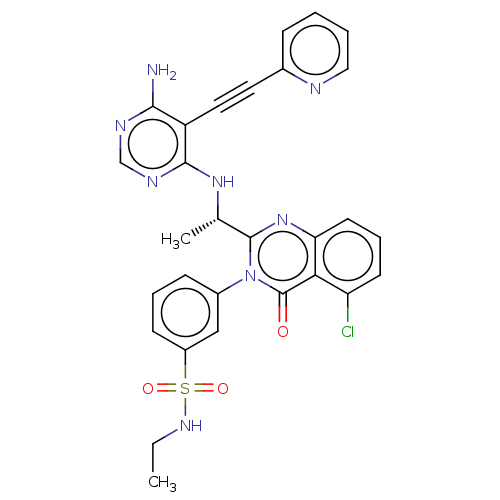
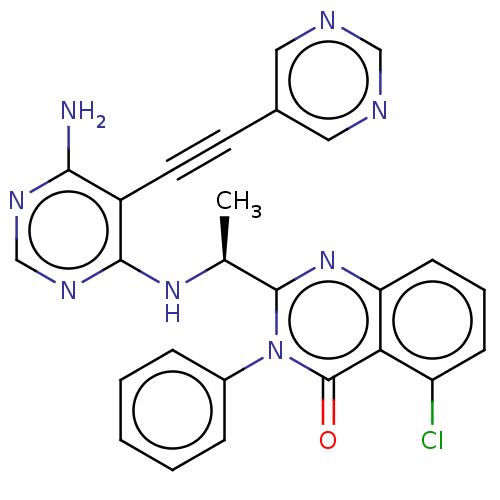
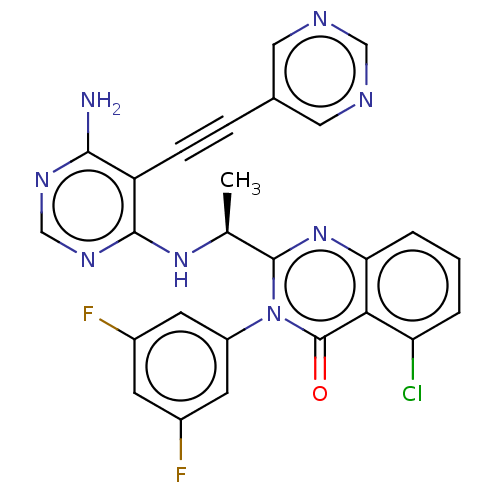
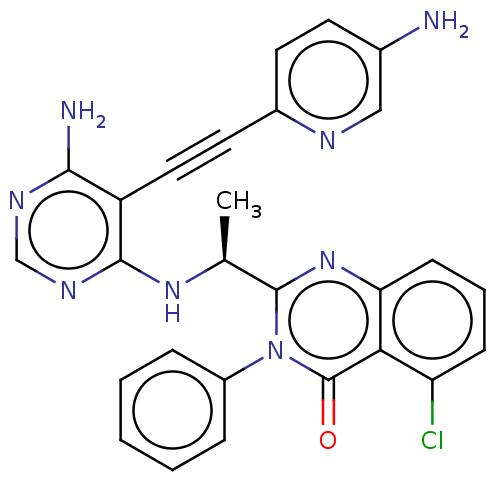
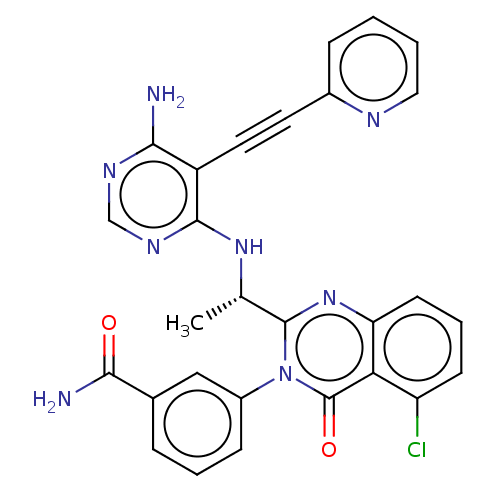
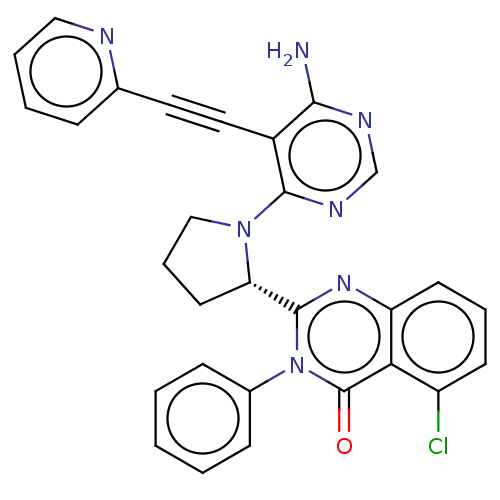
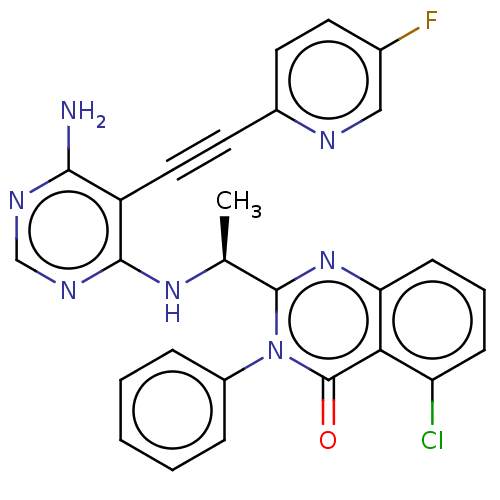
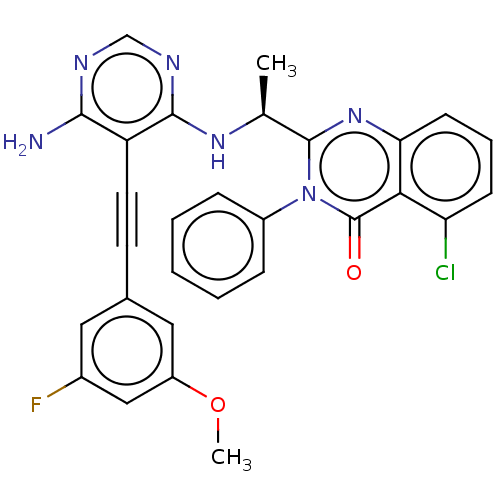
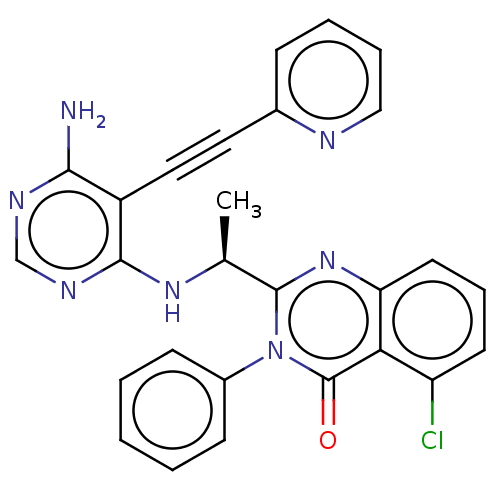
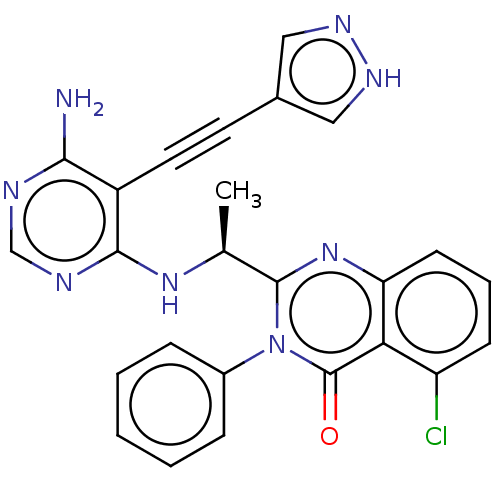
 3D Structure (crystal)
3D Structure (crystal)