Report error Found 82 Enz. Inhib. hit(s) with all data for entry = 50036792
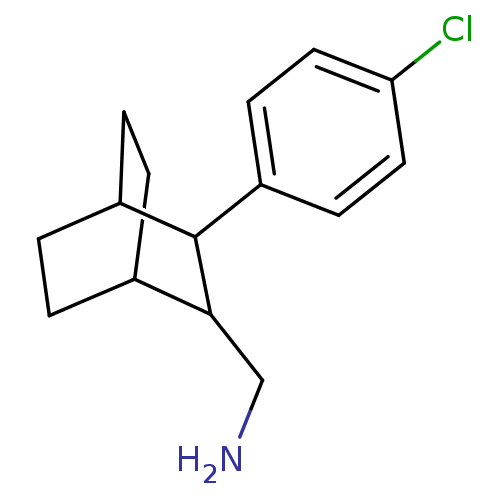
Affinity DataIC50: 12.9nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 12.9nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
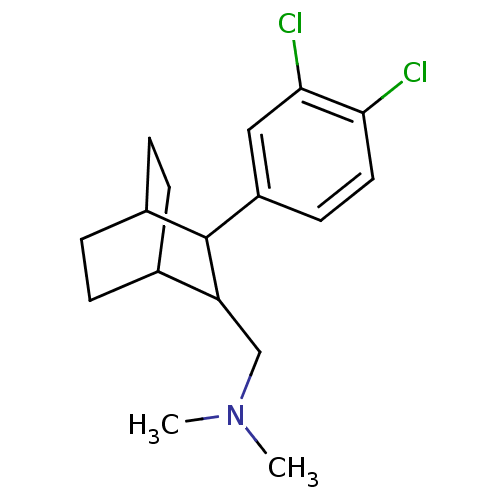
Affinity DataIC50: 14.2nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 14.2nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
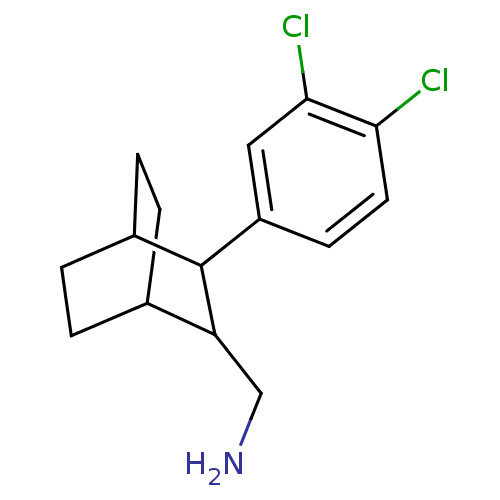
Affinity DataIC50: 22nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
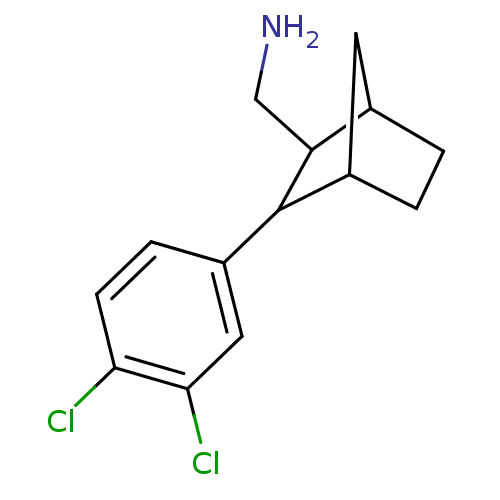
Affinity DataIC50: 29.3nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 29.3nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 68.3nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 68.3nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 94nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 94nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 94nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 191nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 191nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 204nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 204nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 207nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 207nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 207nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 207nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 239nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 239nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 249nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 283nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 283nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibitory binding activity against rat striatal tissue using [3H]WIN-35248 as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 404nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 593nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 593nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 593nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 704nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 758nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 780nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 840nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 840nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 875nMAssay Description:Inhibition of dopamine uptake into rat striatal tissue, using [3H]- dopamine as the radioligand.More data for this Ligand-Target Pair