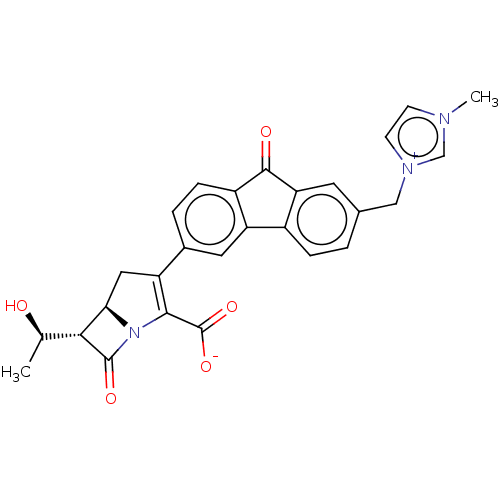
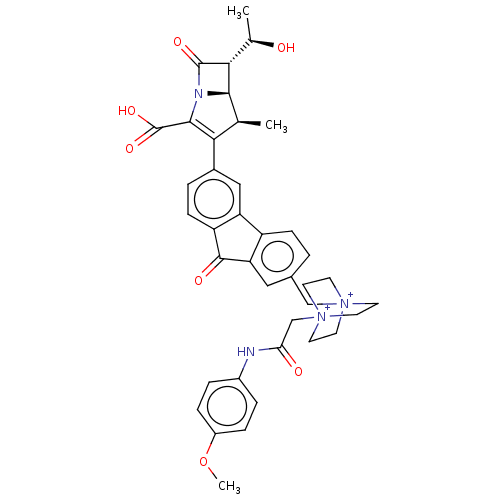
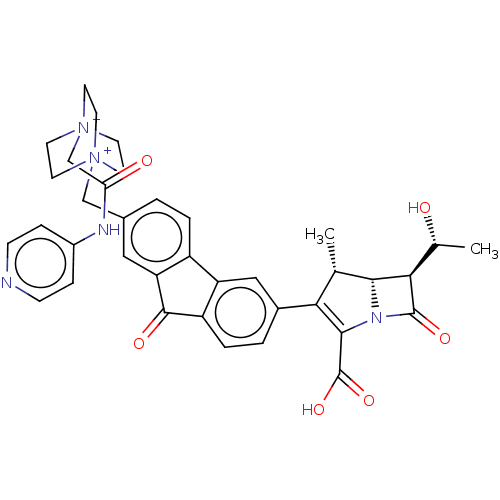
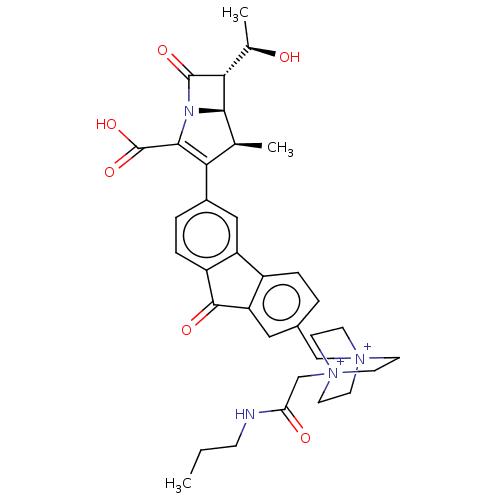
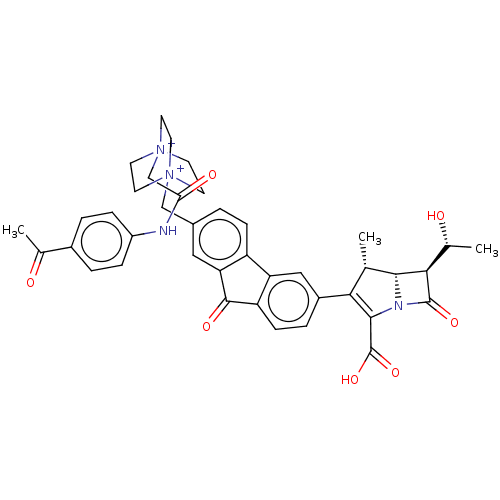
Report error Found 48 Enz. Inhib. hit(s) with all data for entry = 50048474
Affinity DataIC50: 275nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 306nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 416nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 430nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 455nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 463nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 467nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 474nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 501nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 566nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 573nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 610nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 652nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 729nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 752nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 853nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 867nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 879nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 906nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.18E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.31E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.47E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.96E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.66E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.97E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.15E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.25E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.43E+3nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.34E+5nMAssay Description:Binding affinity against PBP2a receptor by competition analysis with [3H]benzylpenicillin using cell membrane fractions from the MRSA COL strain.More data for this Ligand-Target Pair