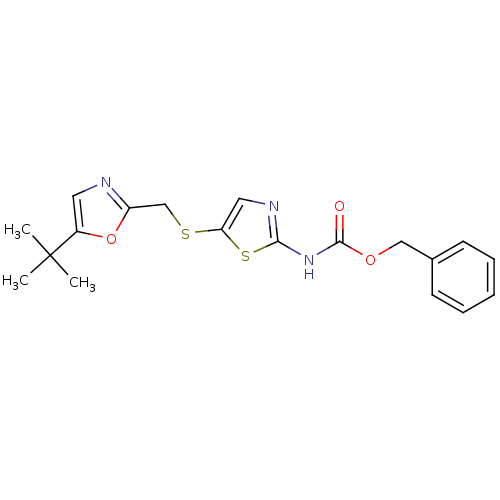
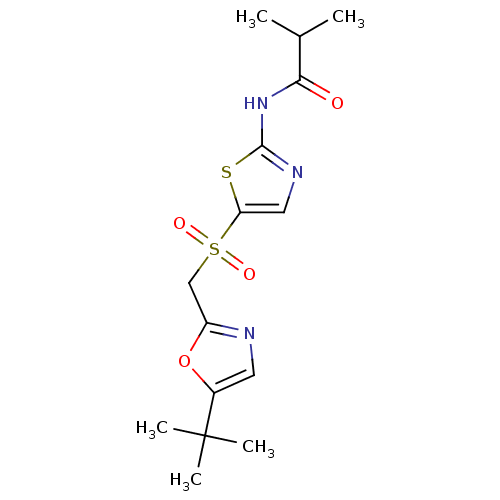
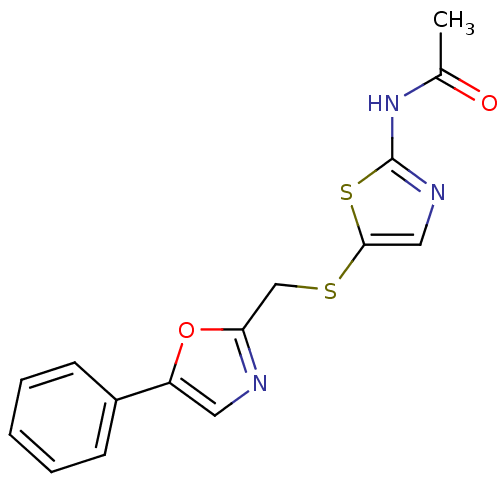
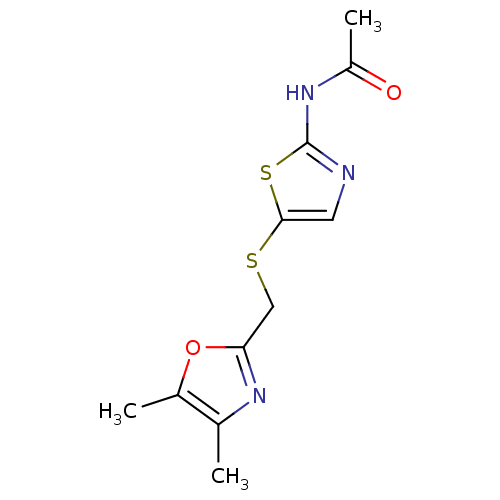
Report error Found 94 Enz. Inhib. hit(s) with all data for entry = 747
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 2nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 3nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 3nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 3nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 4nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 5nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 5nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 5nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 5nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 6nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 6nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 10nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 11nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 17nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 18nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 19nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 20nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 25nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 26nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 30nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 40nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 50nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 50nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 60nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 60nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 60nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 80nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 90nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 90nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 100nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 140nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 140nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 150nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 170nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 200nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 250nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 250nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 300nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 340nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 350nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 360nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 390nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase/G2/mitotic-specific cyclin- 1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 440nMpH: 8.0 T: 2°CAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 640nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Human)
Bristol-Myers Squibb Pharmaceutical Research Institute
Bristol-Myers Squibb Pharmaceutical Research Institute
Affinity DataIC50: 680nMAssay Description: The enzyme was assayed with substrate in the presence of 25 uM ATP/[gamma-33P] ATP and test compound. Dose response curves were generated to determ...More data for this Ligand-Target Pair
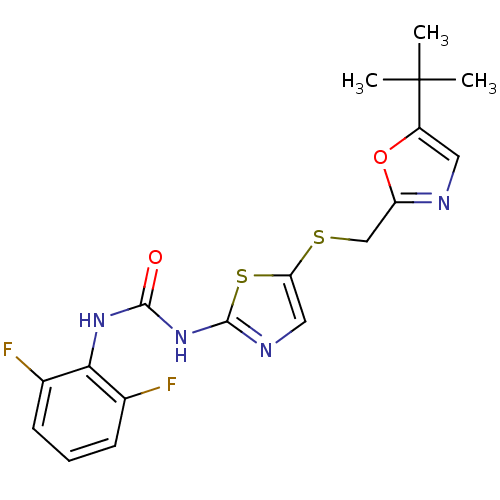
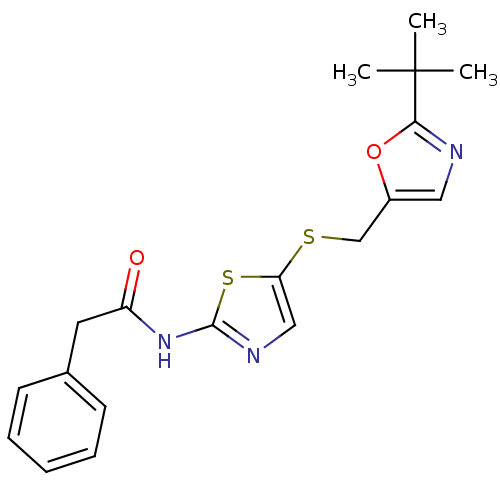
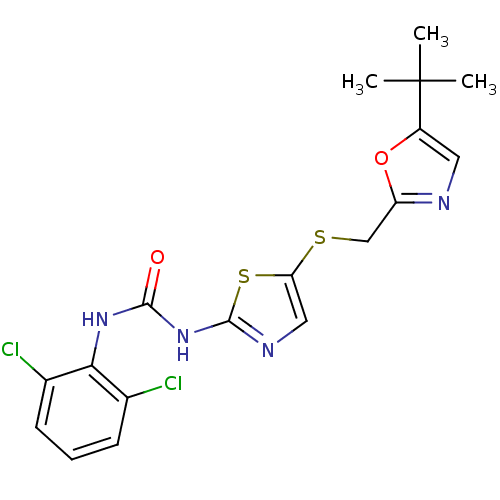


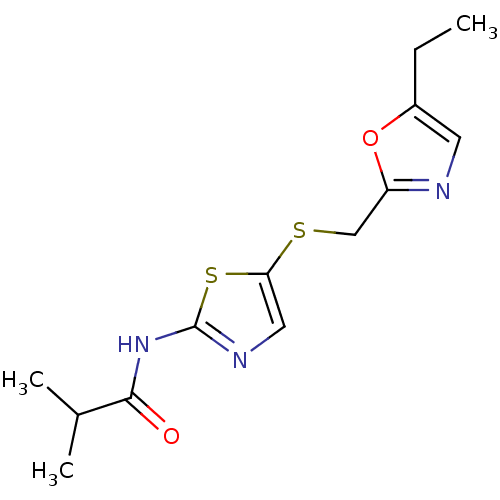
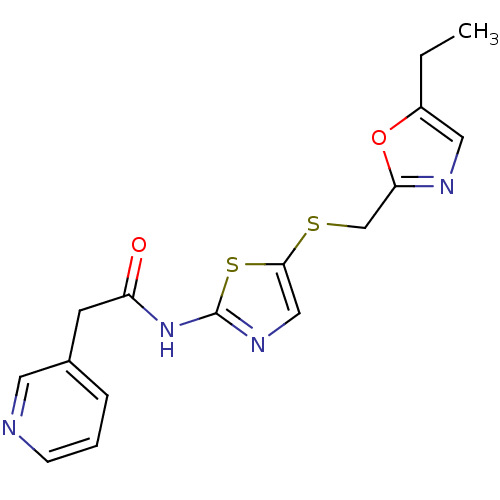

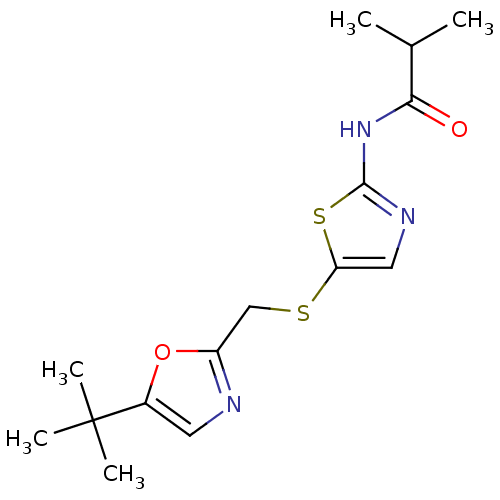
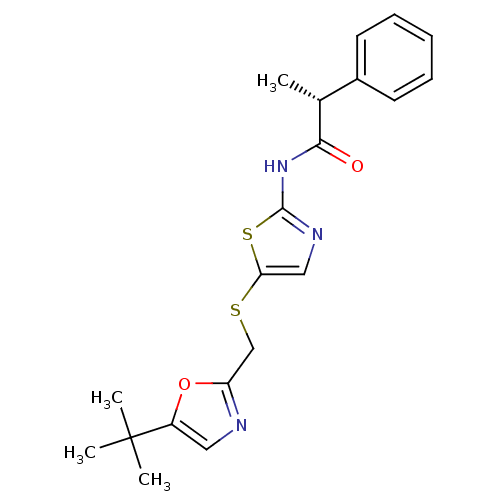






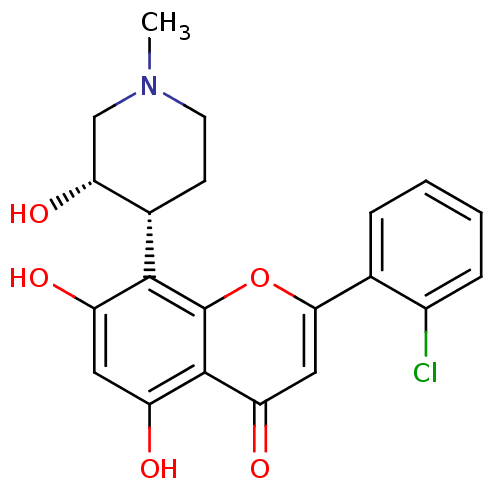
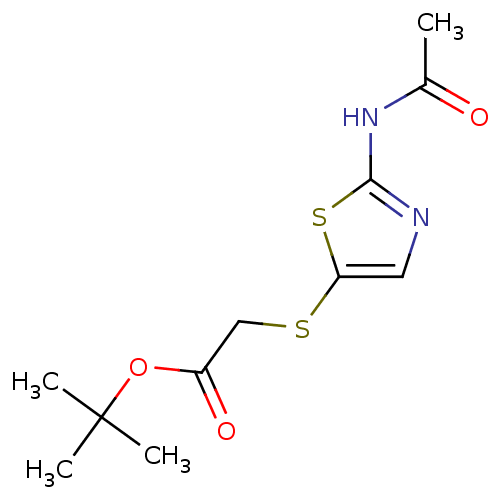
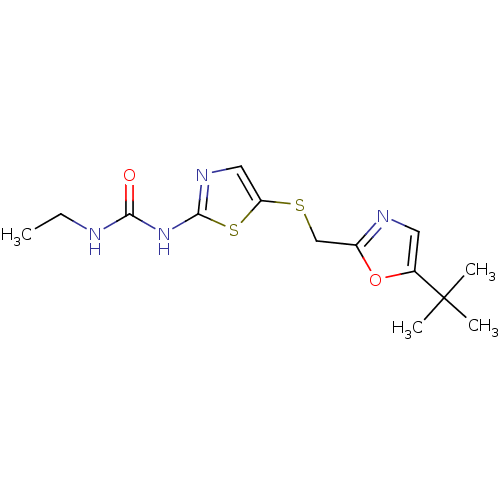

 3D Structure (crystal)
3D Structure (crystal)