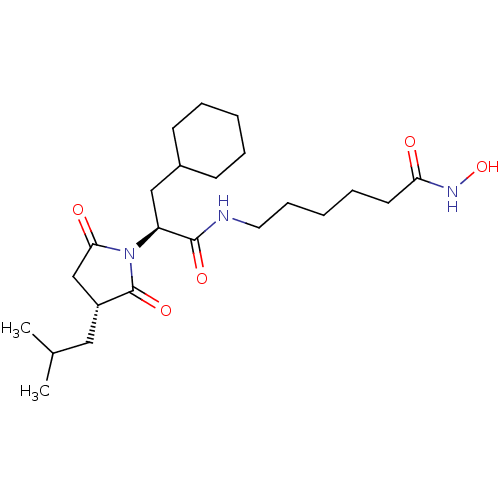
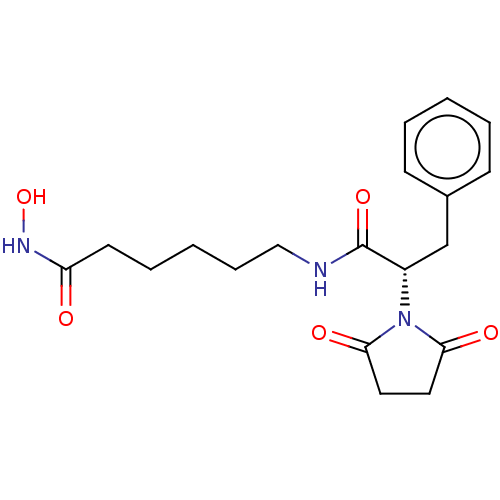
Report error Found 13 Enz. Inhib. hit(s) with all data for entry = 50048530
Affinity DataIC50: 1.40nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 99nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 640nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Concentration required to inhibit Histone Deacetylase (HDAC) from K562 erythroleukemia cellsMore data for this Ligand-Target Pair
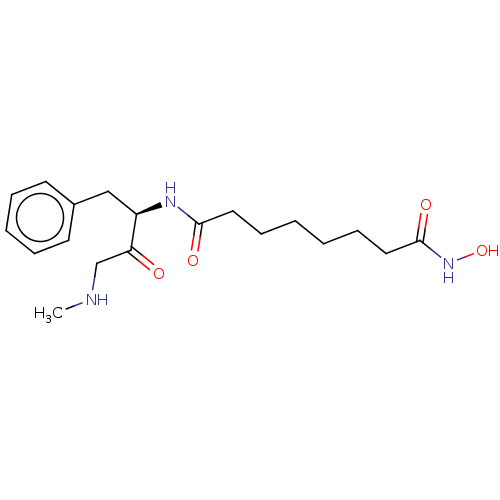
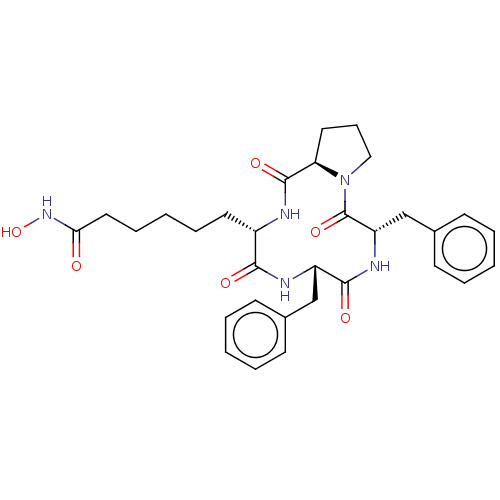

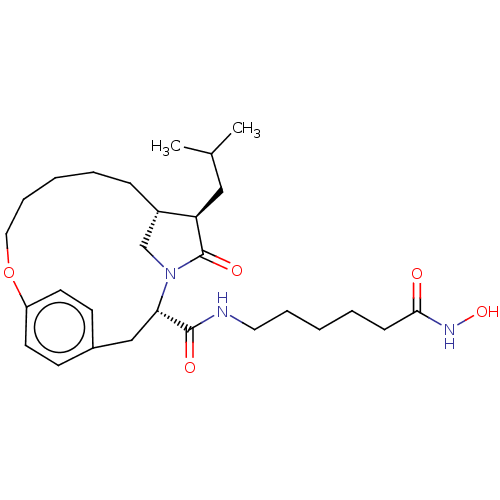

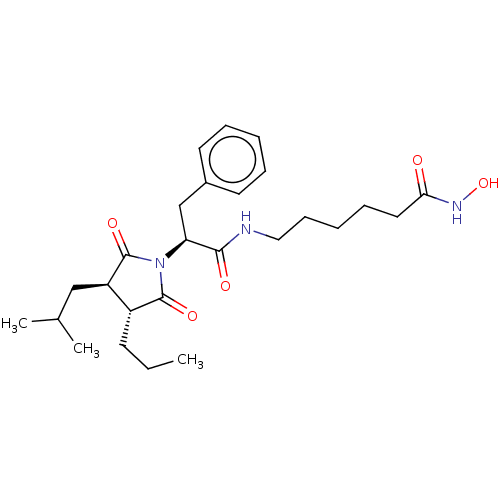
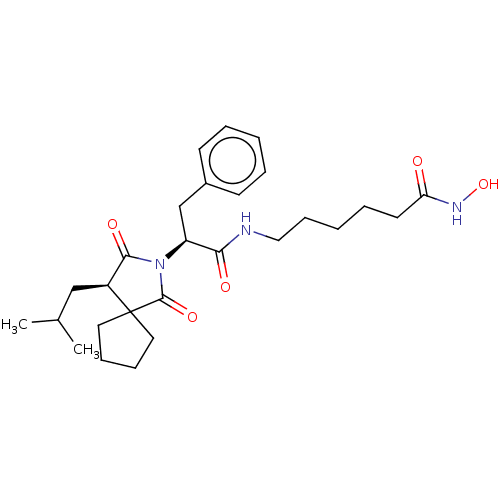
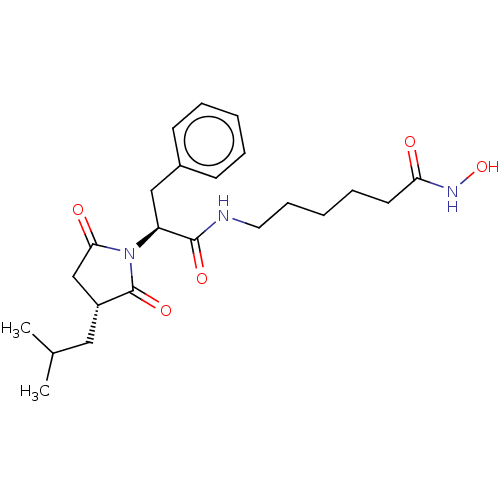
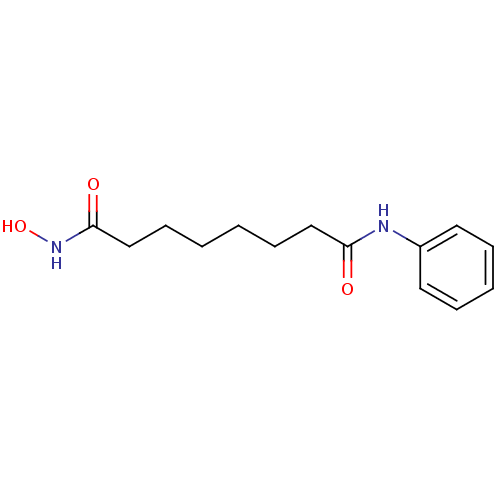
 3D Structure (crystal)
3D Structure (crystal)