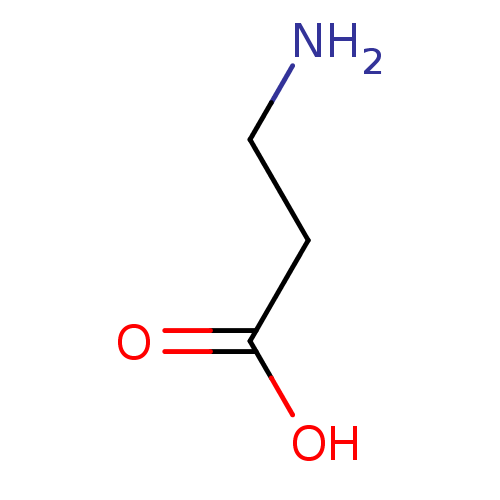
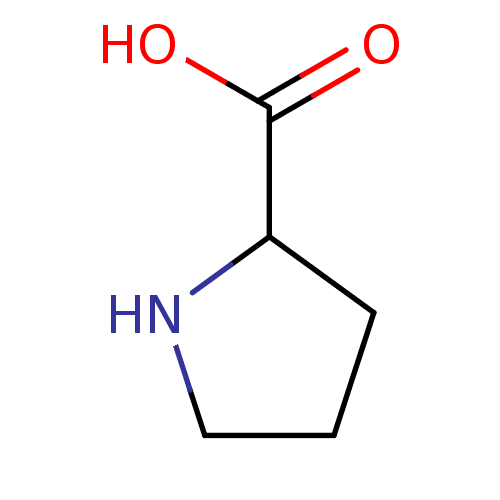
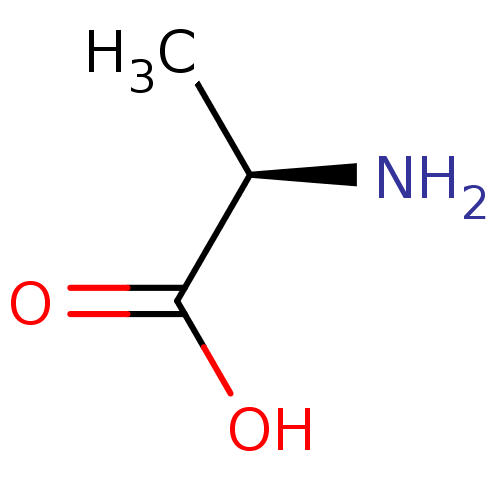
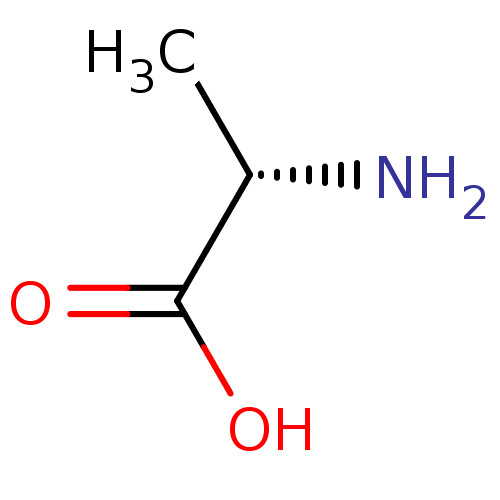
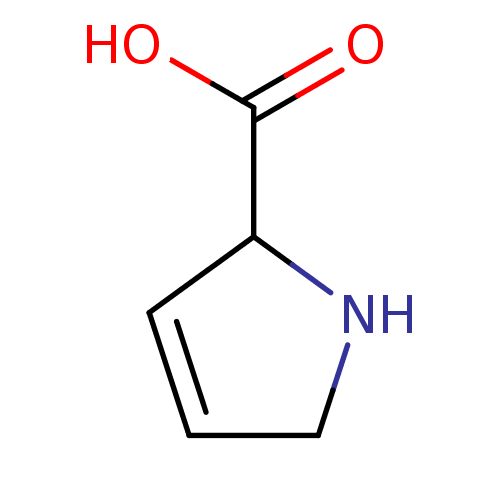
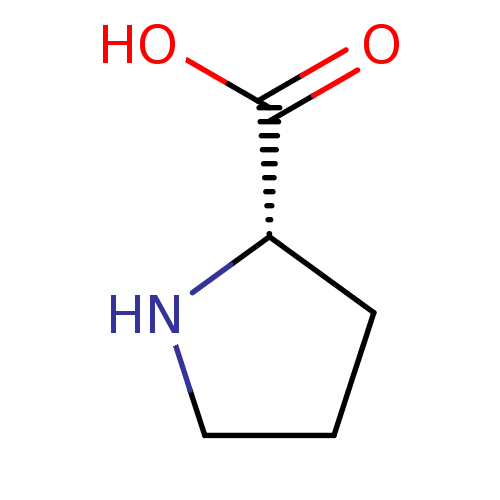
Report error Found 12 Enz. Inhib. hit(s) with all data for entry = 50000012
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 7.90E+3nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+4nMAssay Description:Inhibitory concentration of the compound required to inhibit [3H]-strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.72E+4nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.37E+4nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 9.09E+4nMAssay Description:Inhibitory concentration of the compound required to inhibit [3H]-strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.41E+4nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.98E+5nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.42E+5nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 3.99E+5nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 4.14E+5nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 6.94E+5nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
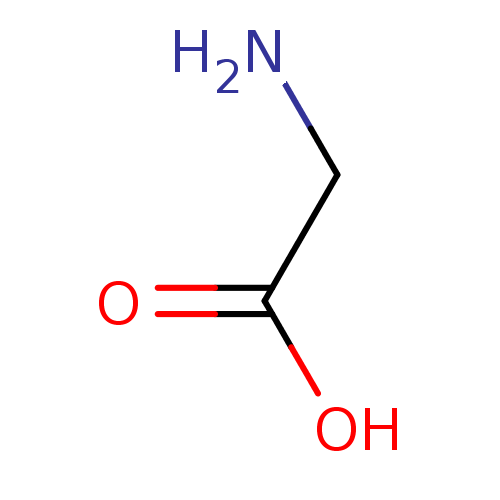
 3D Structure (crystal)
3D Structure (crystal)