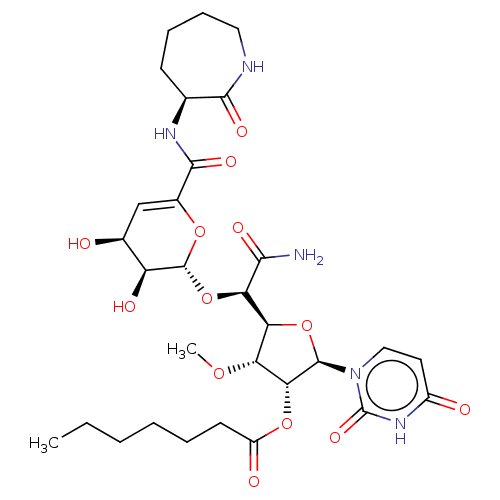
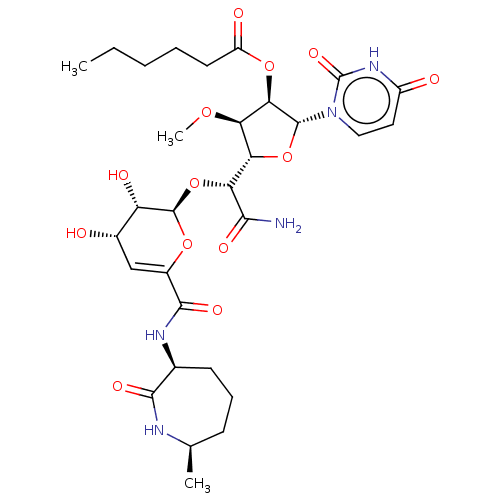
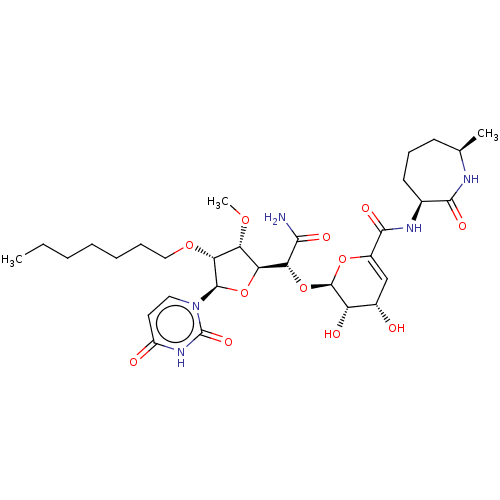
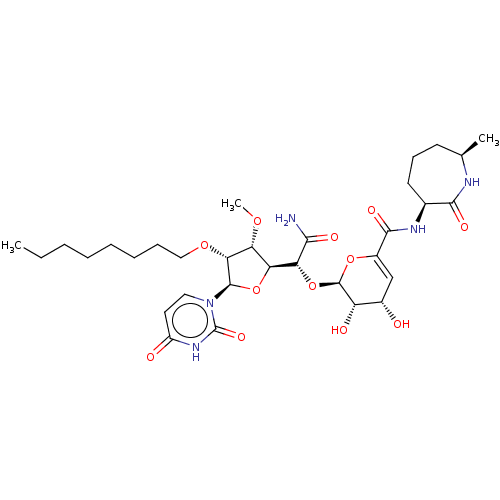
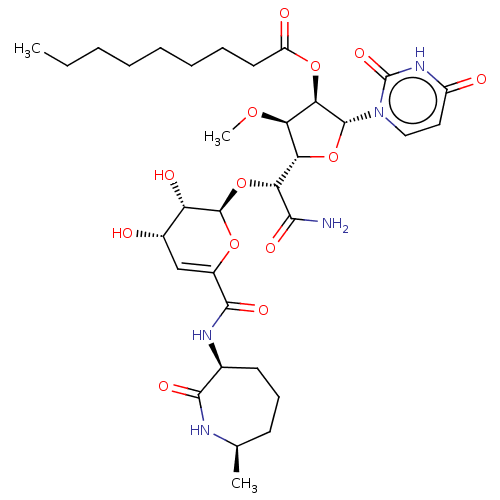
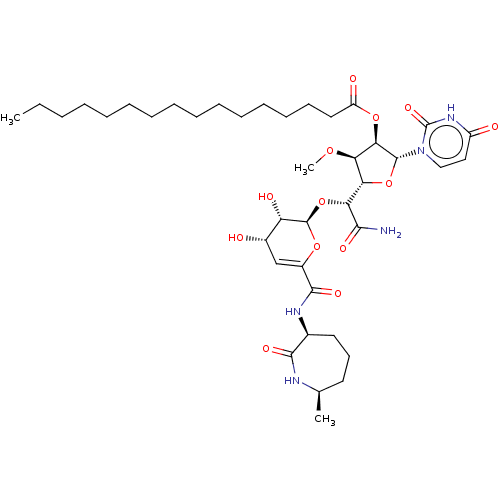
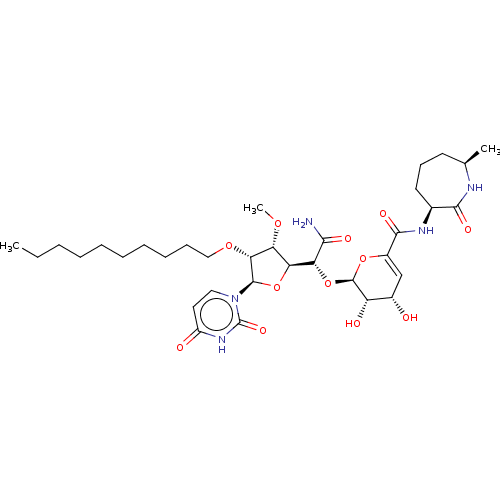
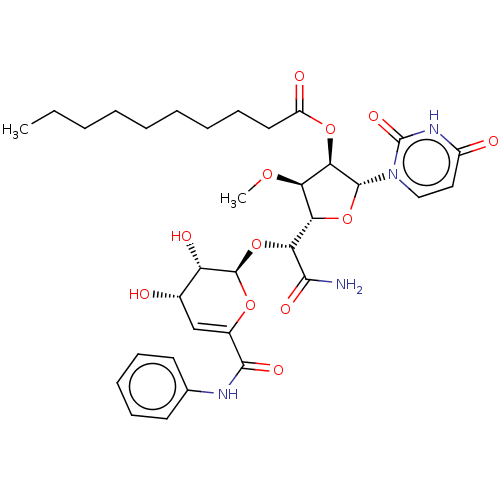
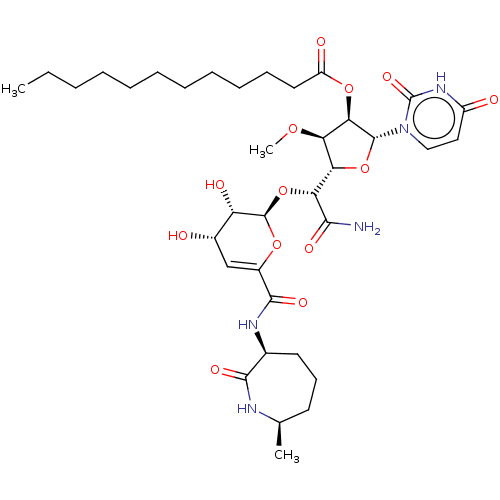
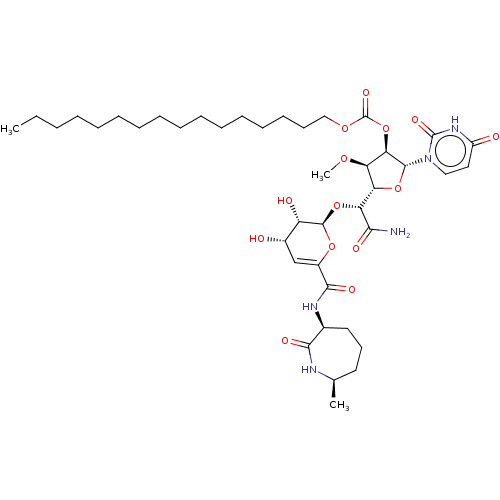
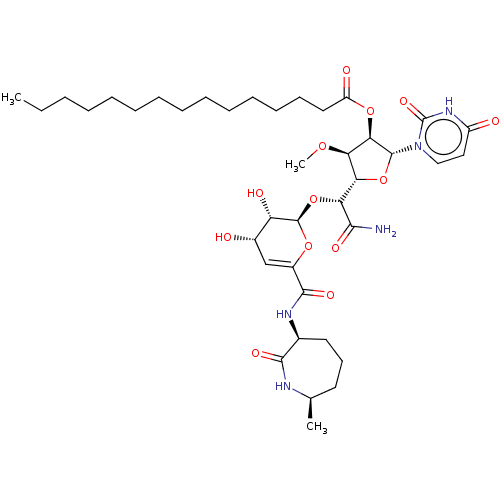
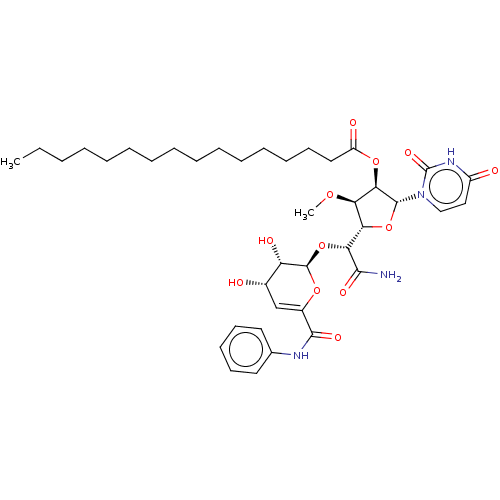
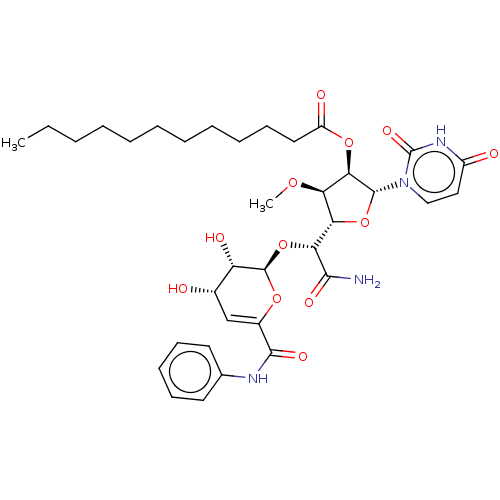
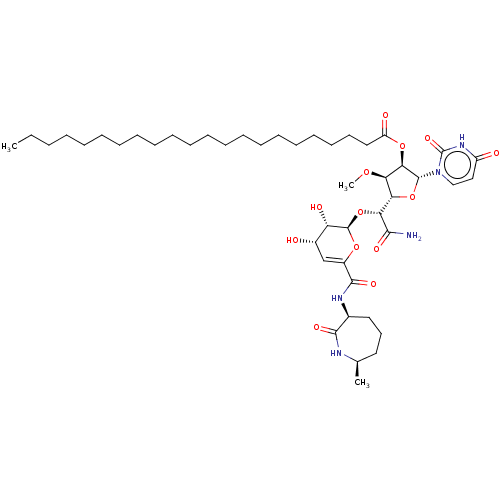
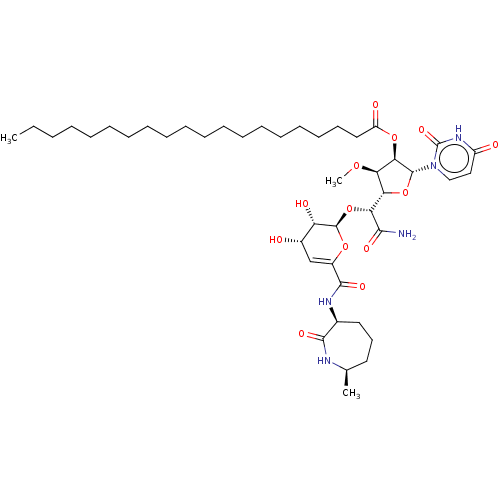
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 50048557
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 12.2nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
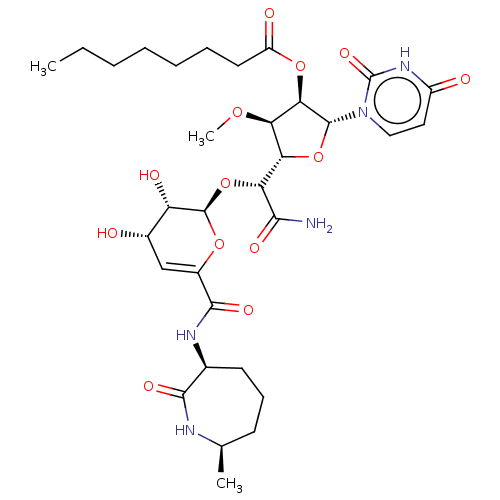
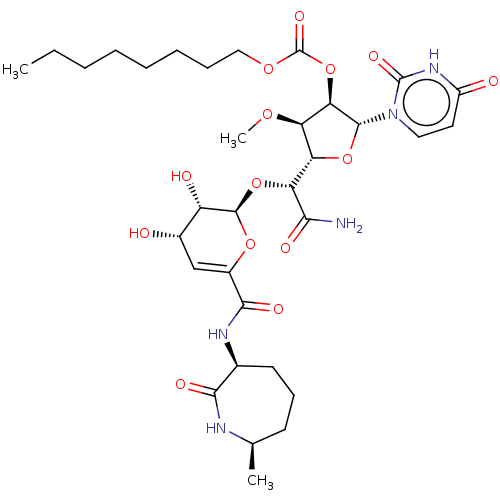
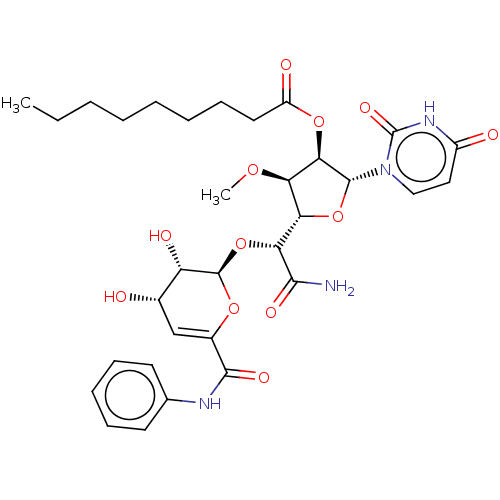
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 17.1nMAssay Description:Inhibitory concentration against translocase-IMore data for this Ligand-Target Pair
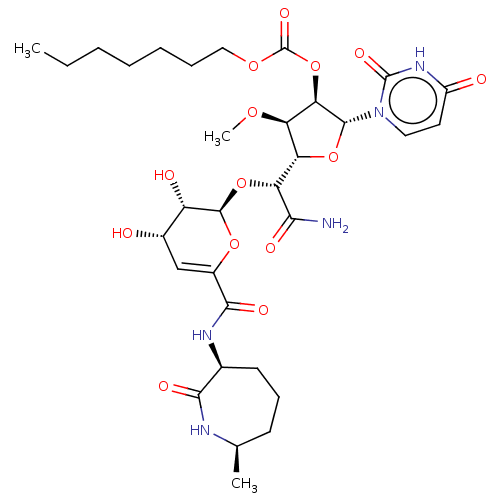
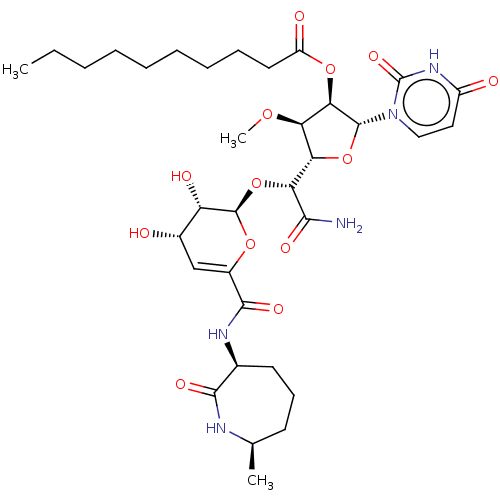
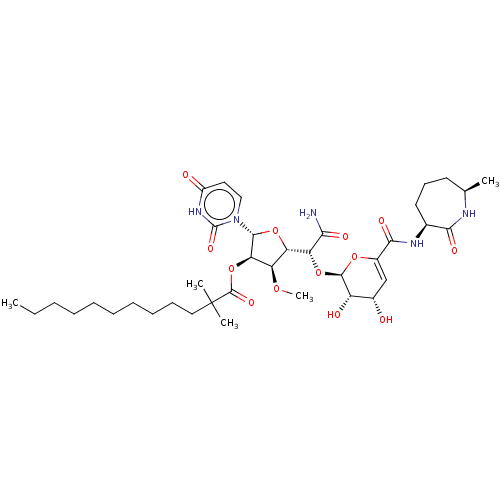
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 17.6nMAssay Description:Inhibitory concentration against translocase-IMore data for this Ligand-Target Pair
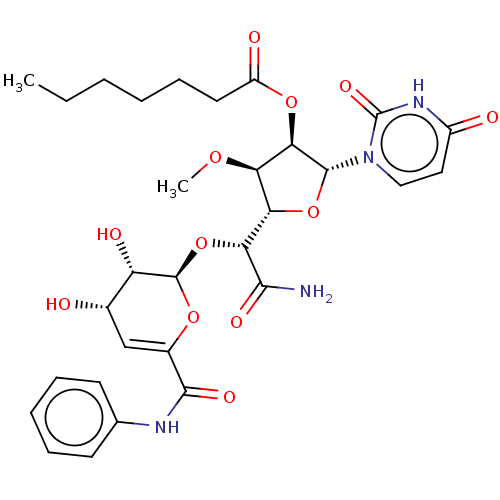
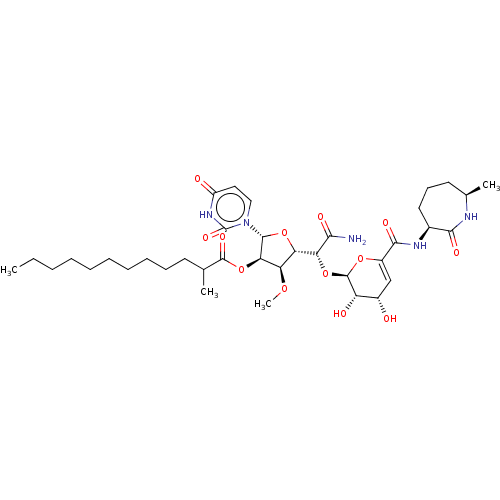
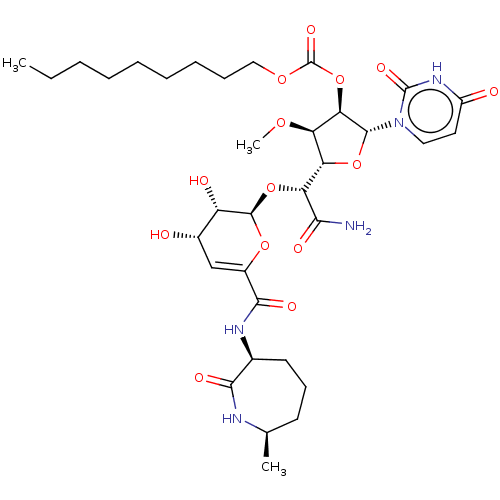
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 35.9nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 58.7nMAssay Description:Inhibitory concentration against translocase-IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 59.9nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 63.1nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 64.7nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 76.5nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 80.7nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 182nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 215nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 234nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 253nMAssay Description:Inhibitory concentration against translocase-IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 267nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 275nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 371nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 414nMAssay Description:Inhibitory concentration against translocase-IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 523nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 675nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 745nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 769nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 884nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 1.03E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 1.25E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 1.85E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 2.75E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 2.87E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 2.93E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 2.97E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 3.23E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 3.48E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 5.66E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 9.41E+3nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Sankyo
Curated by ChEMBL
Sankyo
Curated by ChEMBL
Affinity DataIC50: 3.41E+4nMAssay Description:Inhibitory concentration required against Translocase IMore data for this Ligand-Target Pair