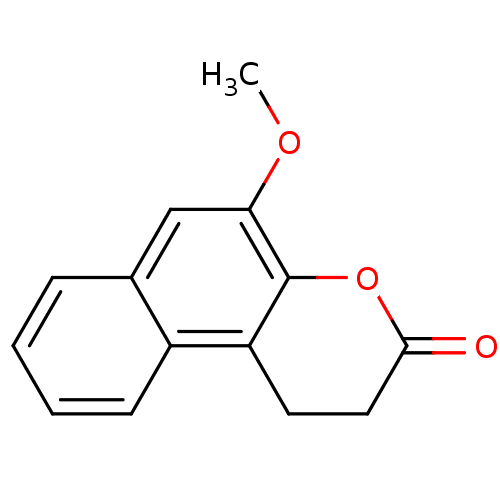
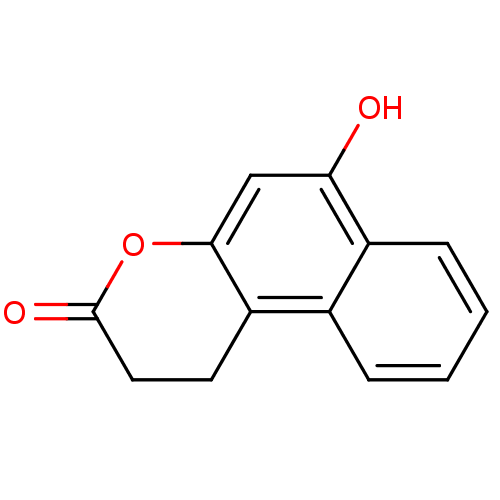
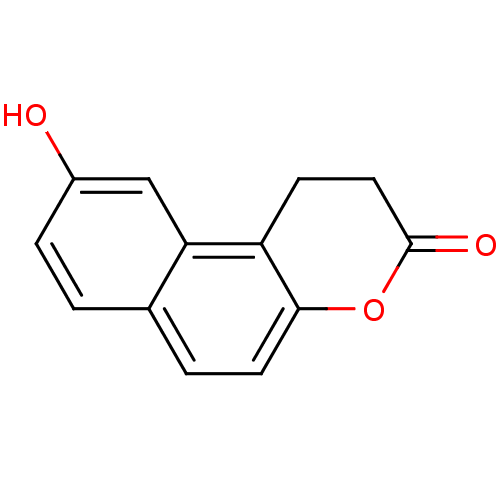
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50014678
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
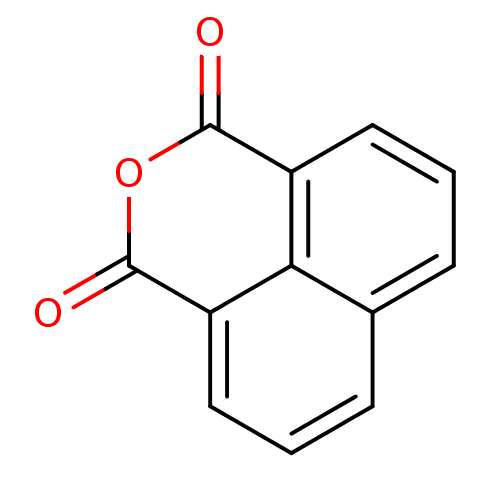
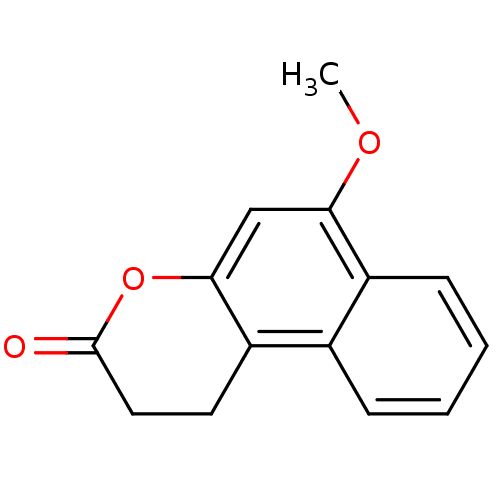
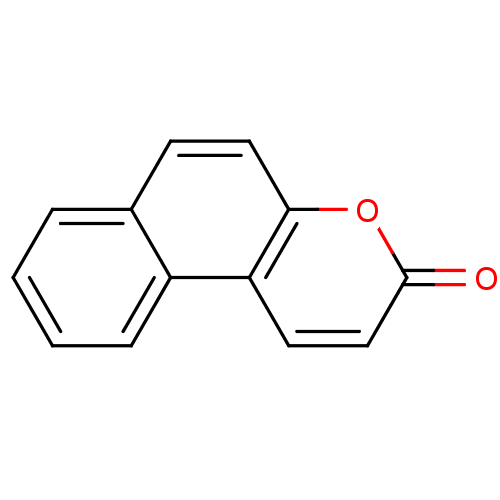
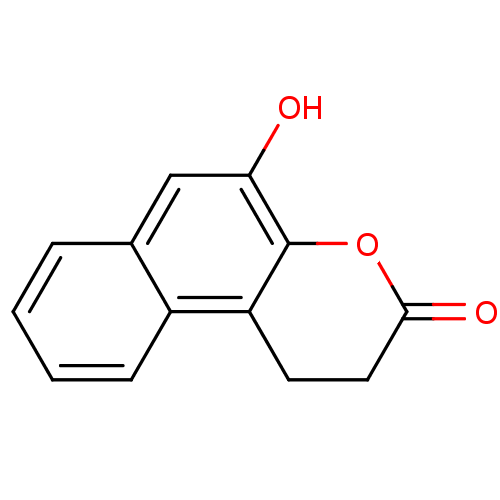
Affinity DataIC50: 8.80E+3nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
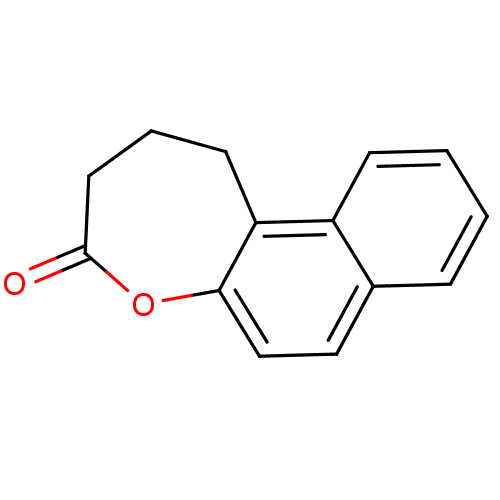
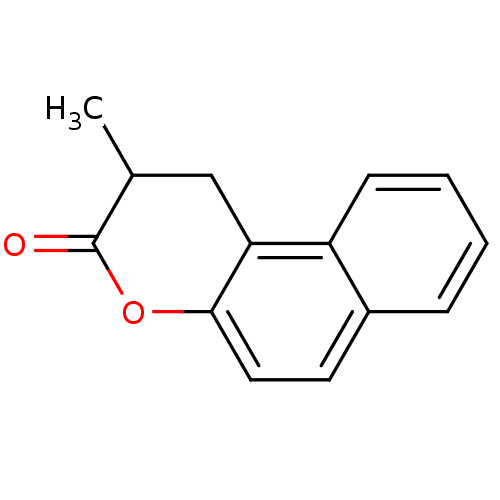
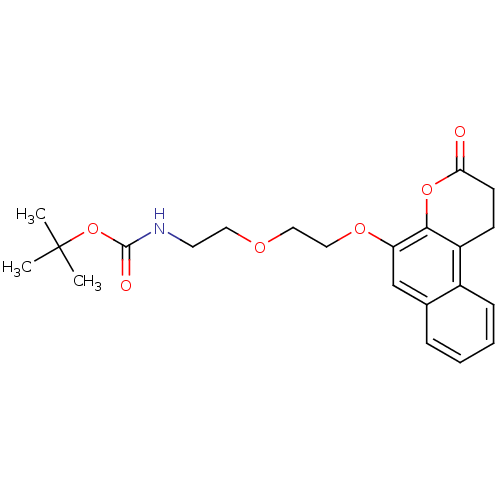
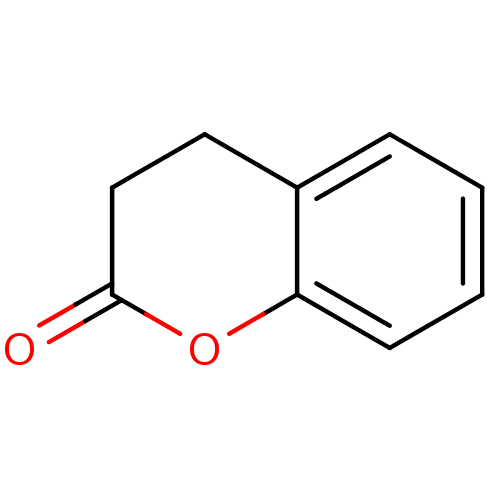
Affinity DataIC50: 1.20E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
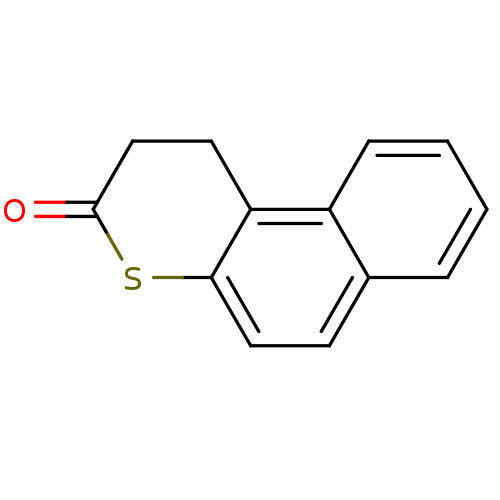
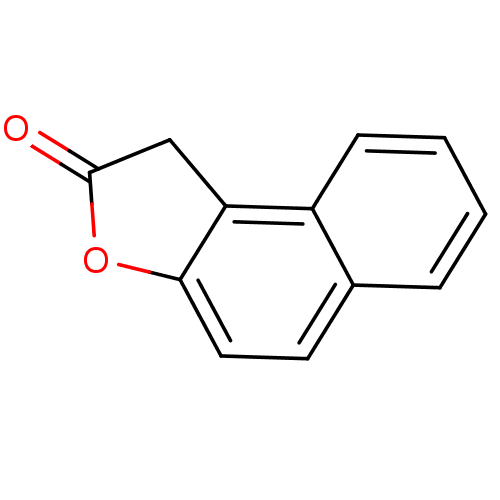
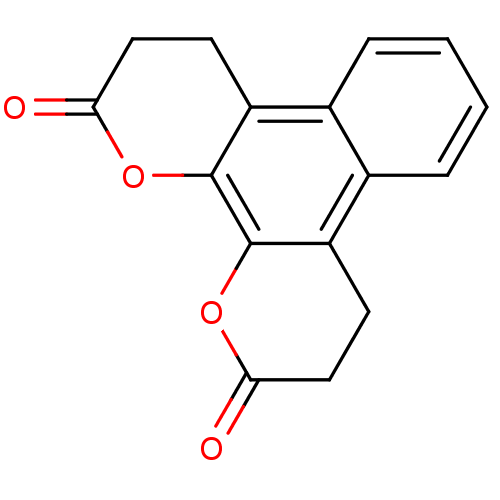
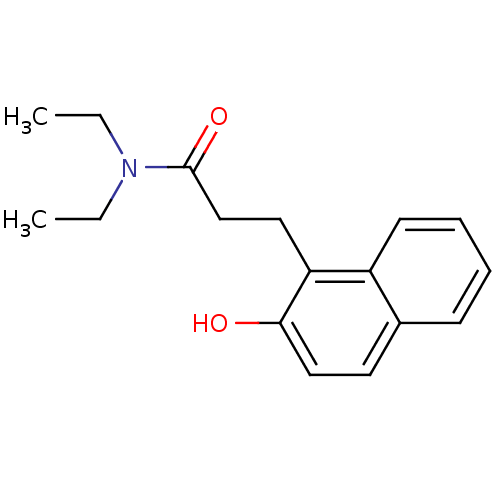
Affinity DataIC50: 2.50E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
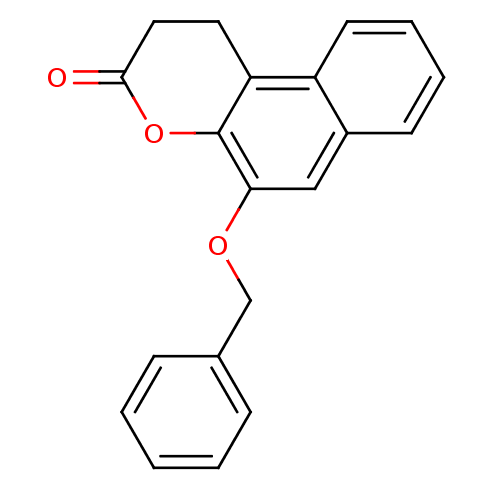
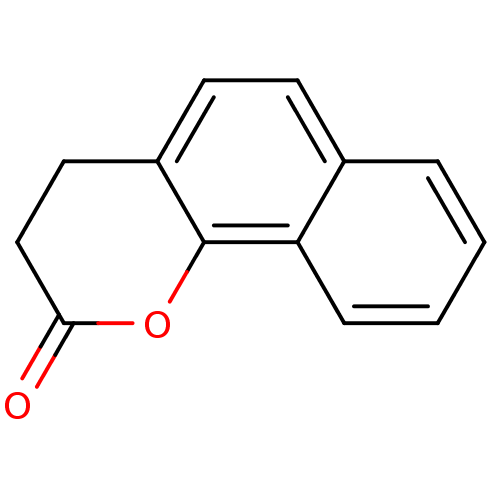
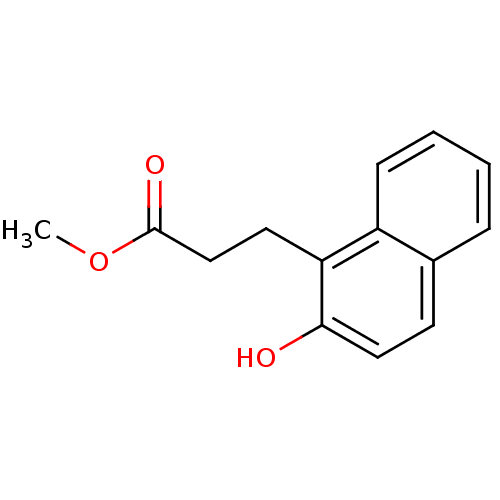
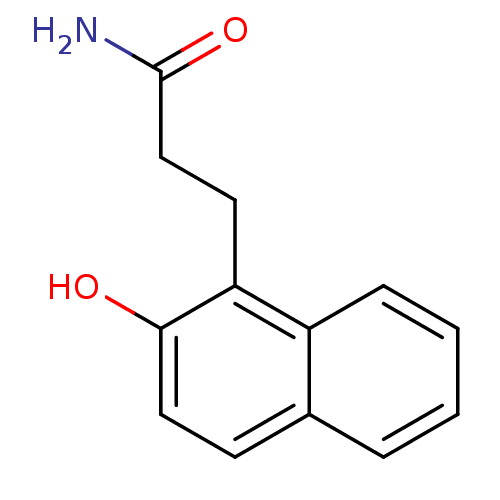
Affinity DataIC50: 2.70E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 3.90E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 4.30E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 4.40E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 5.20E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 5.90E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 6.50E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 7.00E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 7.30E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 7.40E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 8.10E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 8.60E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 8.60E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 8.90E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 9.70E+4nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.11E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.22E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.25E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.25E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.31E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair
TargetNAD-dependent histone deacetylase SIR2(Baker's yeast)
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Fred Hutchinson Cancer Research Center
Curated by ChEMBL
Affinity DataIC50: 1.34E+5nMAssay Description:In vitro inhibition of sirtuin 2 was evaluated using yeast whole cell lysatesMore data for this Ligand-Target Pair