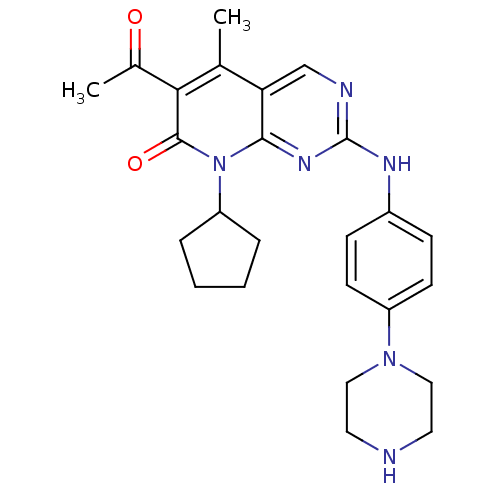
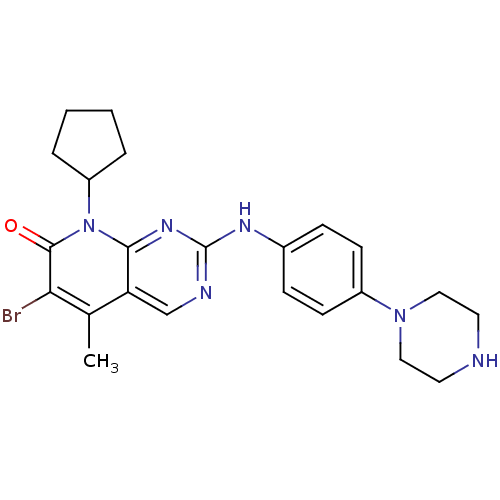
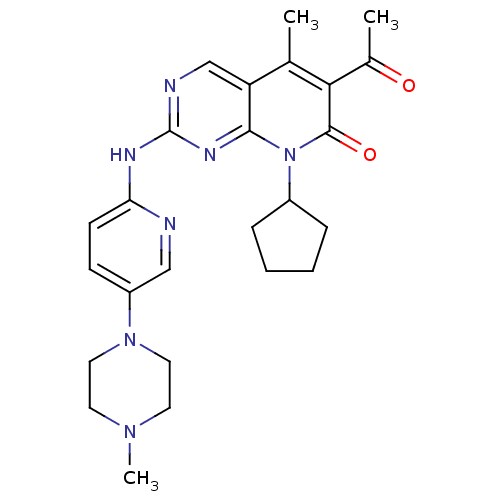
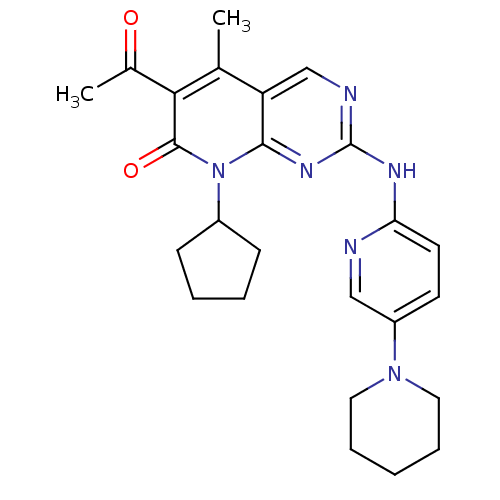
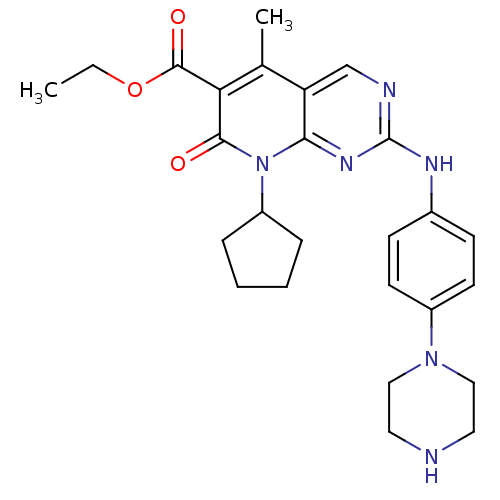
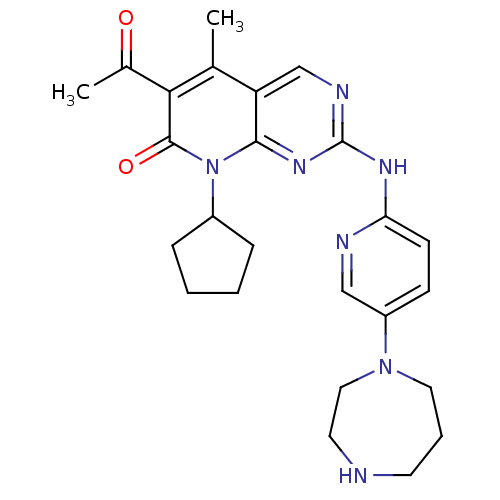
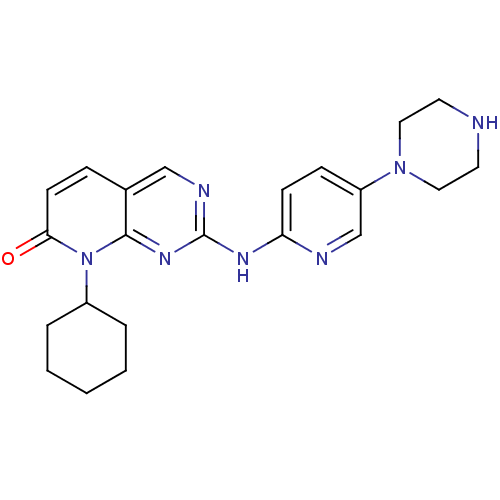
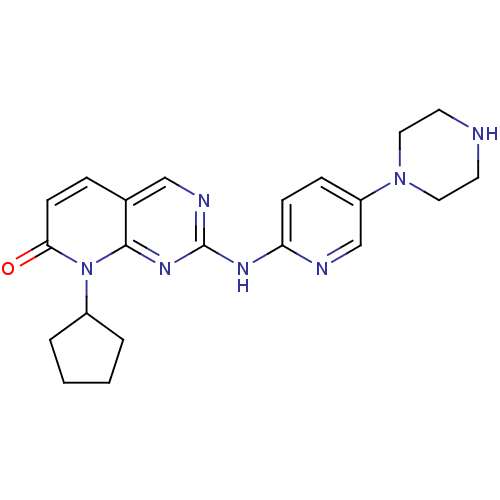
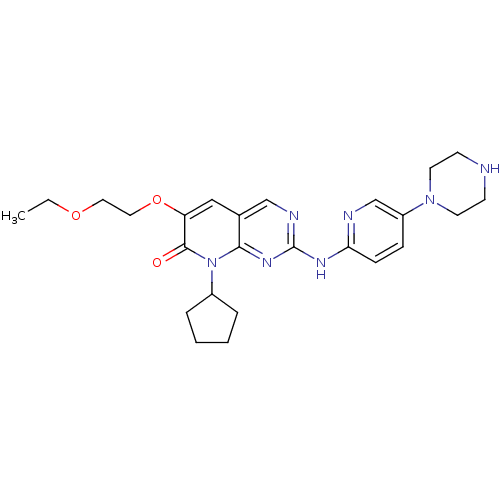
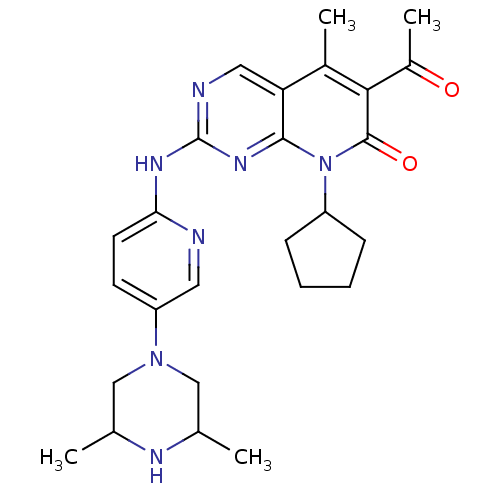
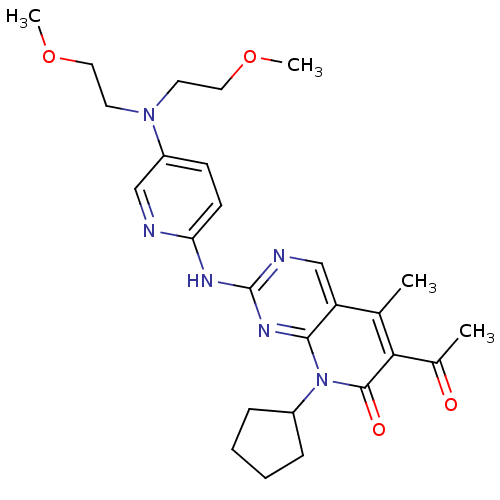
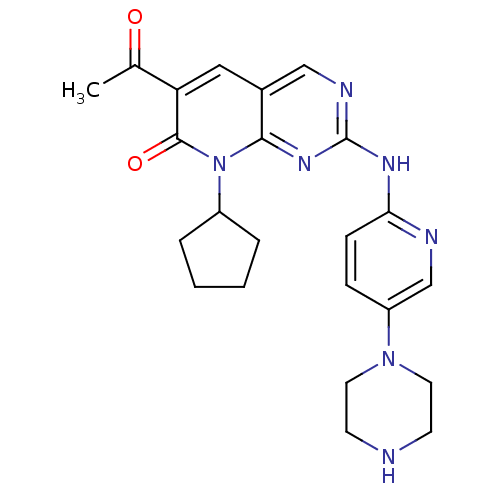
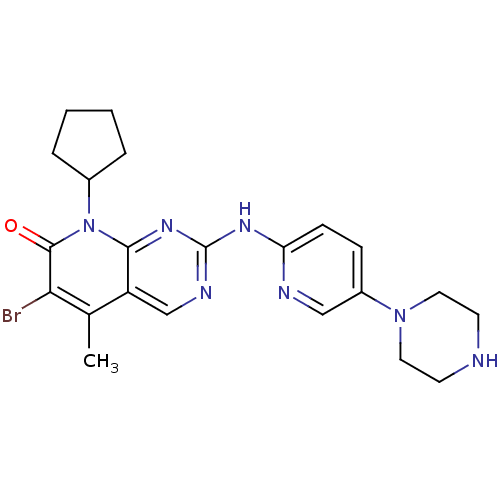
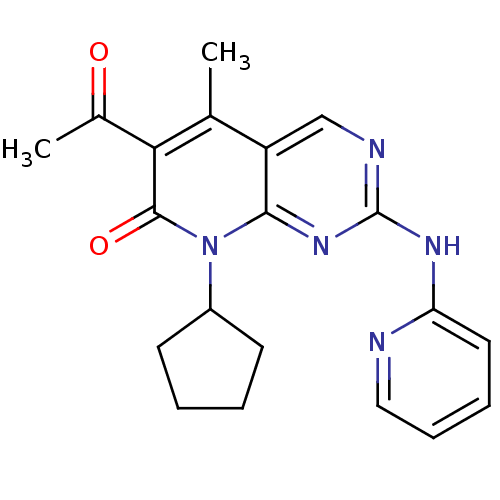
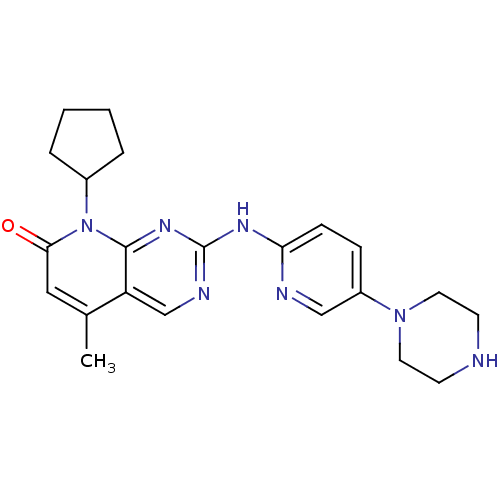
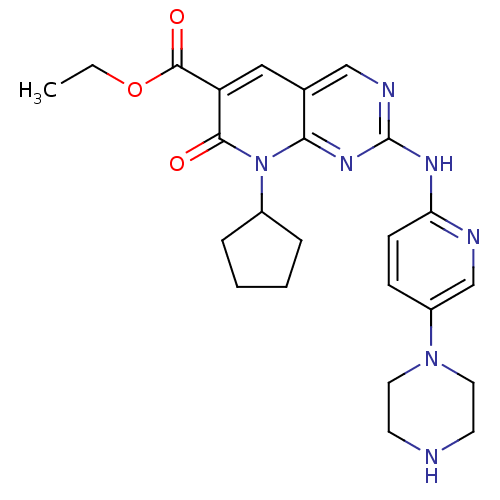
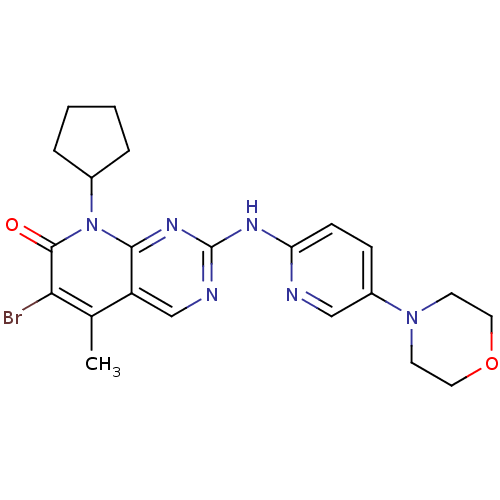
Report error Found 94 Enz. Inhib. hit(s) with all data for entry = 806
Affinity DataIC50: 2nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 123nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 124nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 136nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 145nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 439nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 595nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 835nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 920nMpH: 7.4 T: 2°CAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 950nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.76E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.95E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80E+3nMAssay Description:The enzyme was assayed with substrate GST- retinoblastoma in the presence of 25 uM ATP/[gamma-32P] ATP. IC50 is the inhibitor concentration, which in...More data for this Ligand-Target Pair