Report error Found 77 Enz. Inhib. hit(s) with all data for entry = 50016491
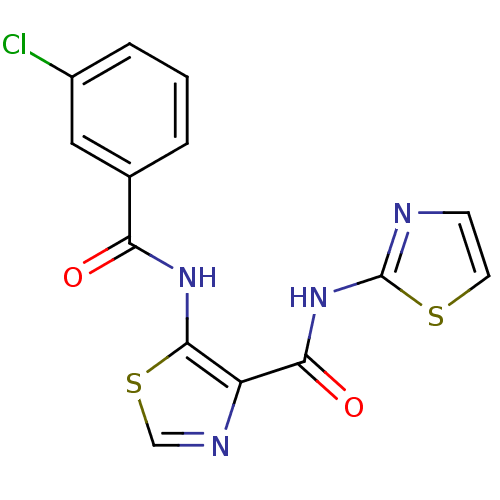
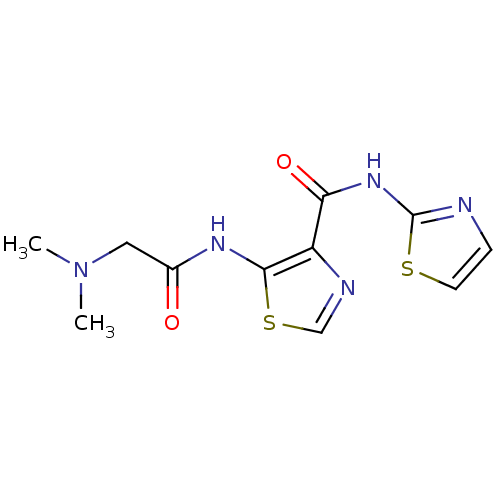
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
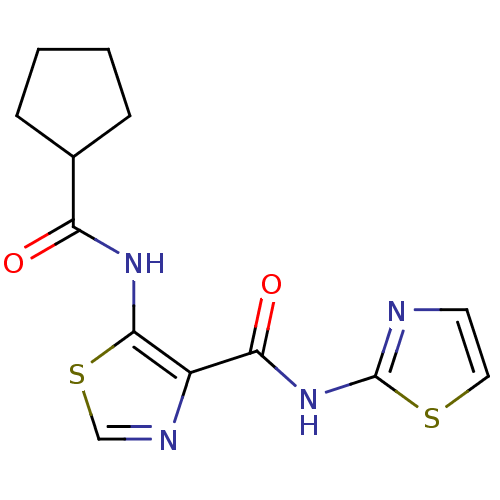
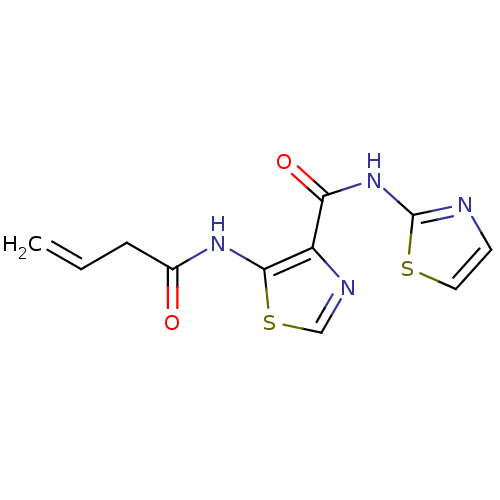
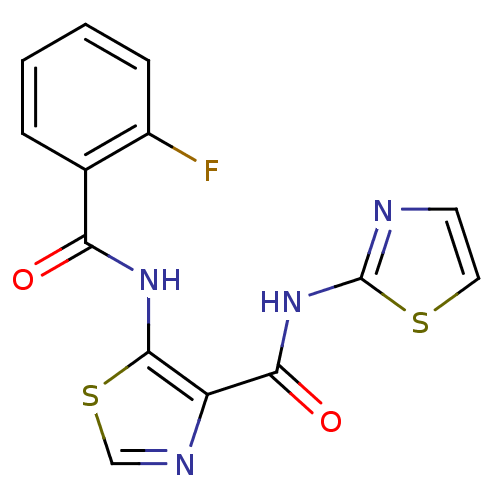
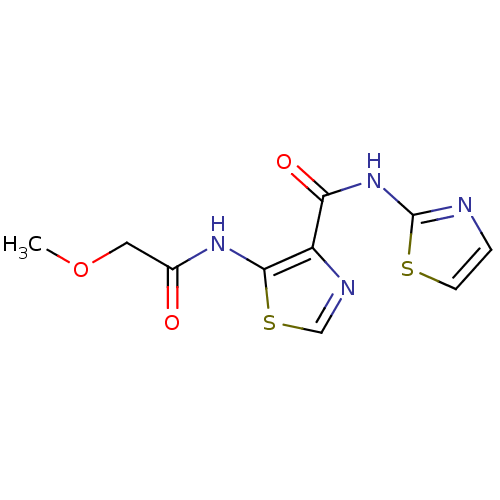
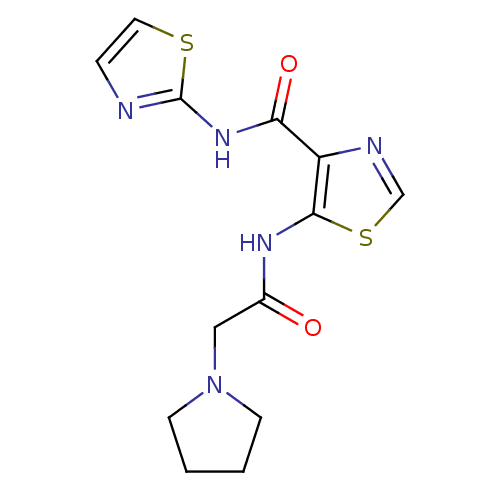
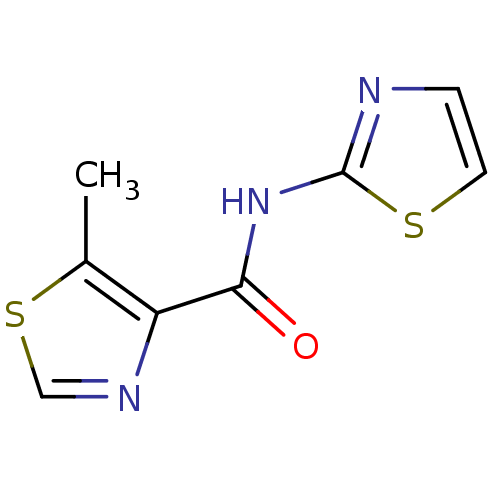
Affinity DataIC50: 24nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
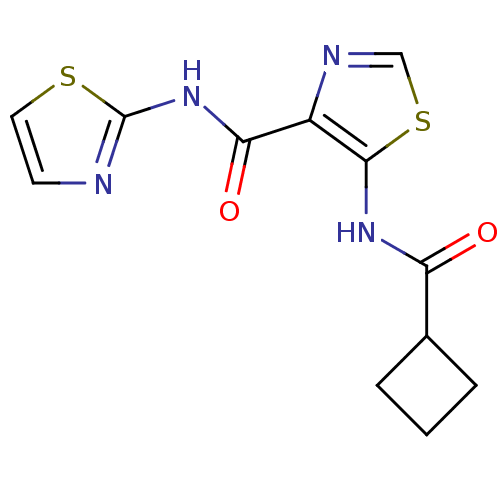
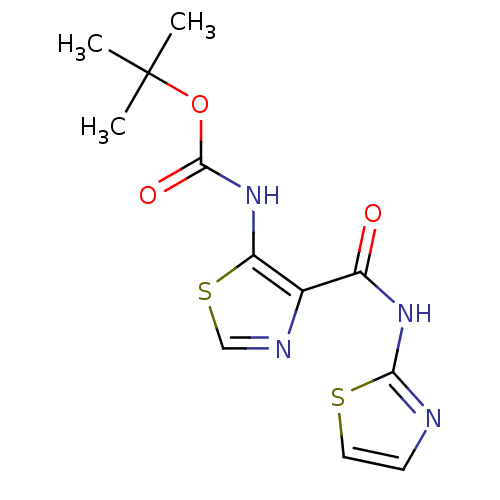
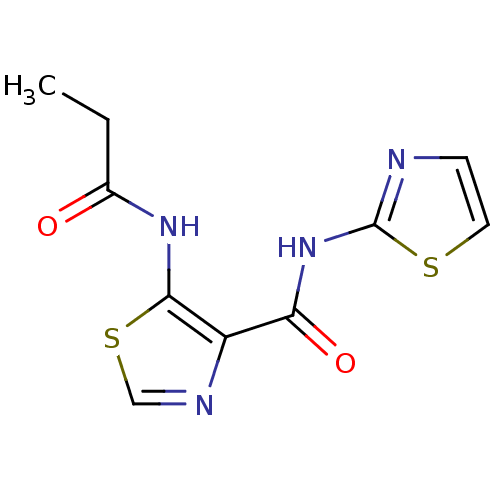
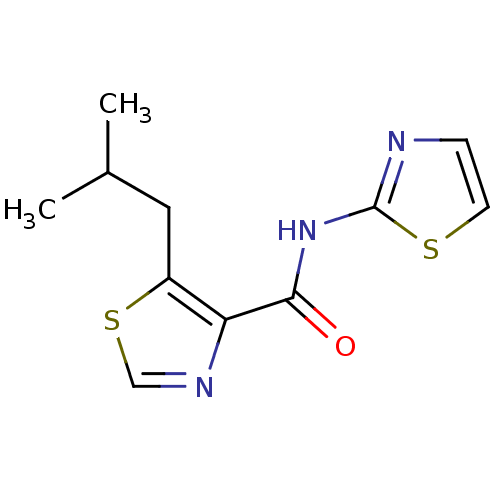
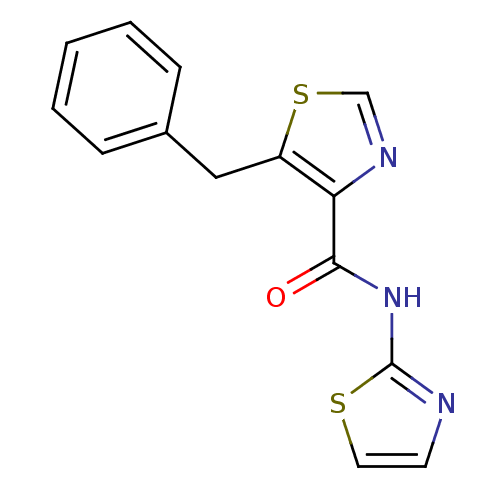
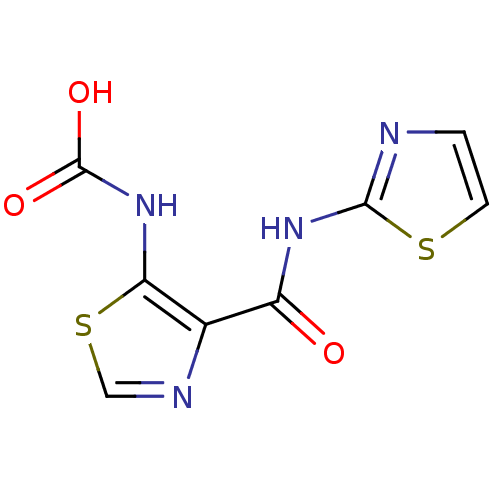
Affinity DataIC50: 29nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
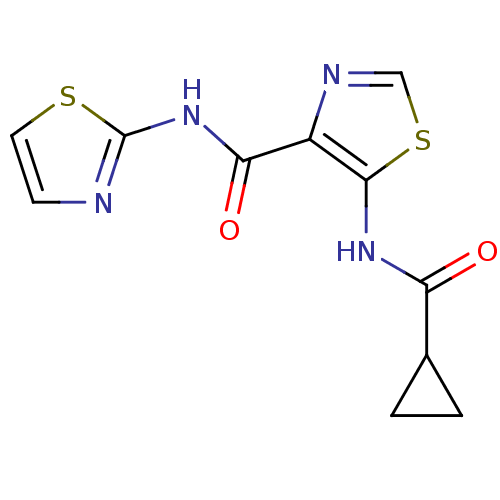
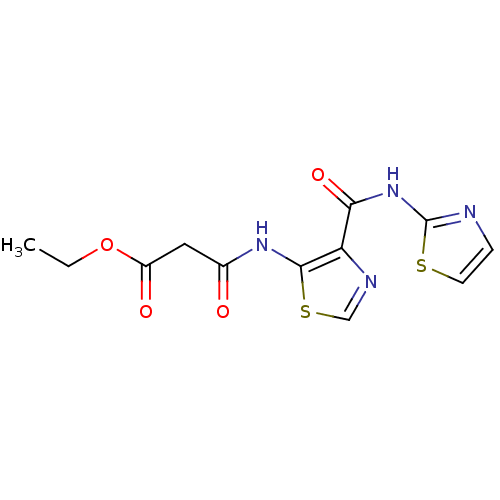
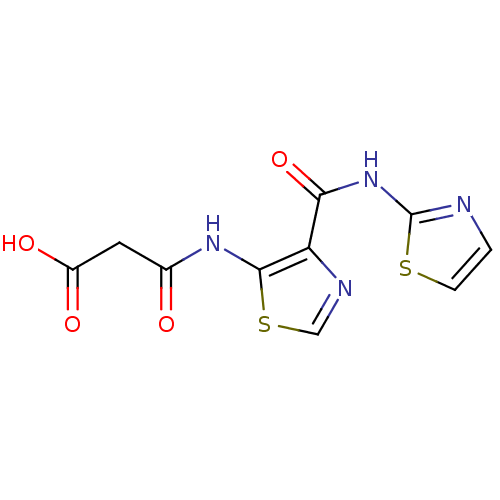
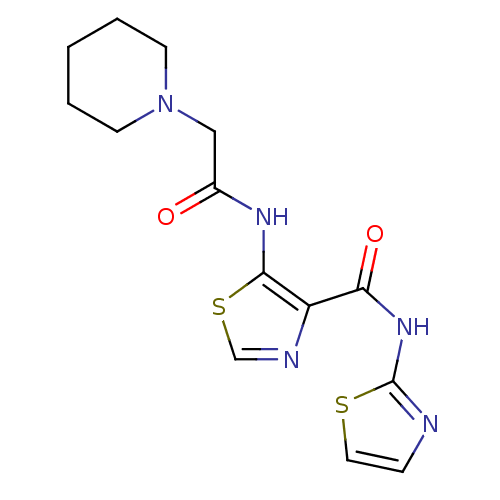
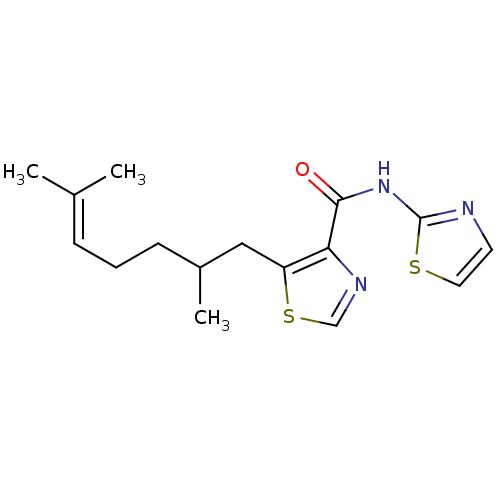
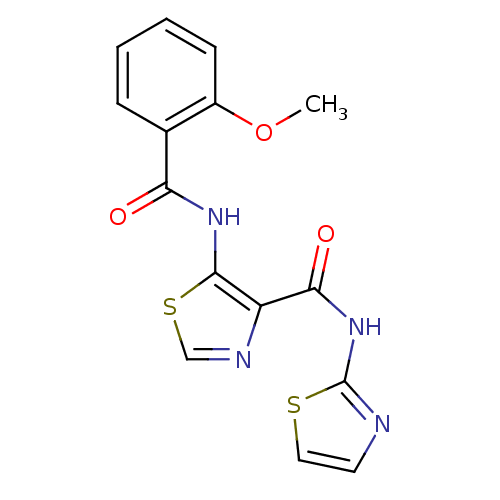
Affinity DataIC50: 40nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
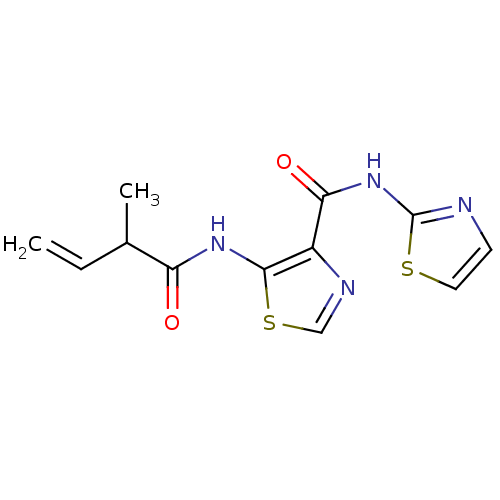
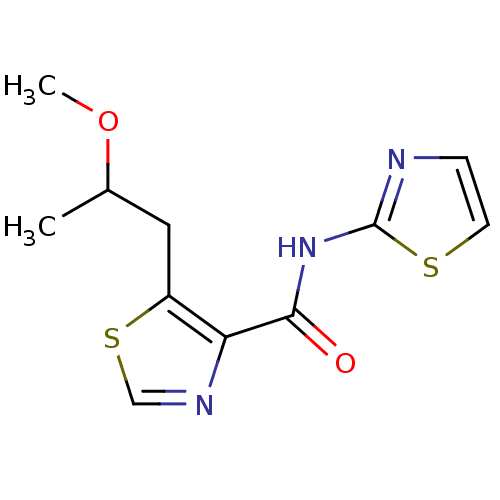
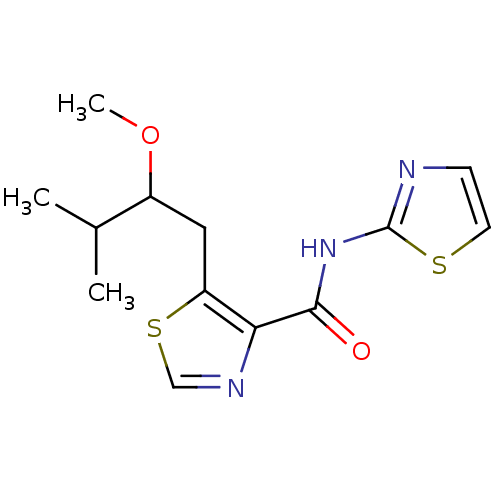
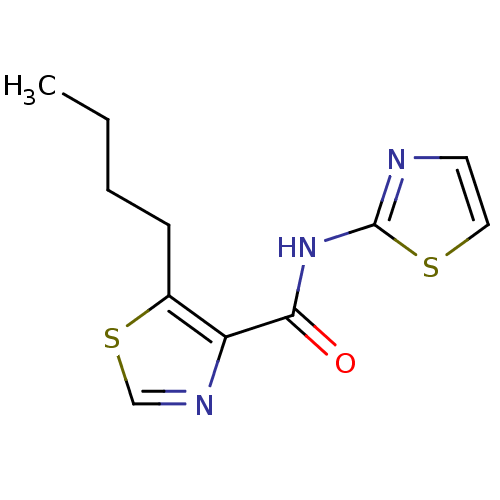
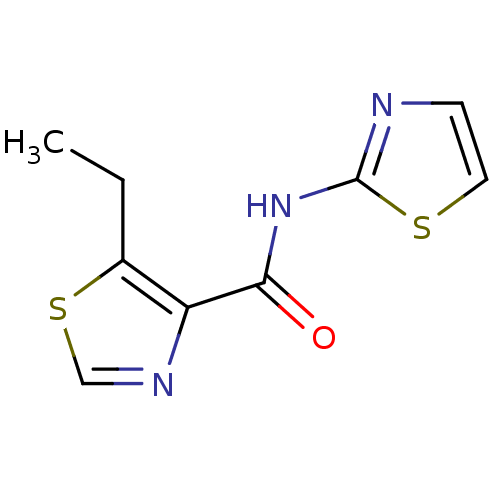
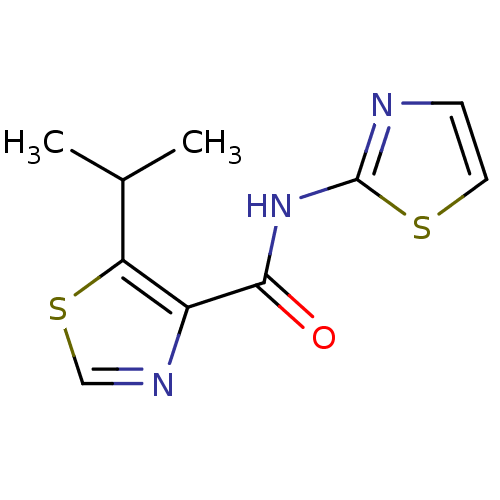
Affinity DataIC50: 42nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 54nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 60nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 65nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 66nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 74nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 76nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 76nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 81nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 82nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 88nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 89nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 92nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 160nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 160nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 180nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 180nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 190nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 190nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 210nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 270nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibitory concentration required against Escherichia coli MetAP1 (150 nM)More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 370nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 480nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 660nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 670nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 860nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 880nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 900nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 980nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 990nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 990nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.12E+3nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair
TargetMethionine aminopeptidase 1(Baker's yeast)
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of The Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.45E+3nMAssay Description:Inhibitory concentration required against Saccharomyces cerevisiae MetAP1 (330 nM) More data for this Ligand-Target Pair