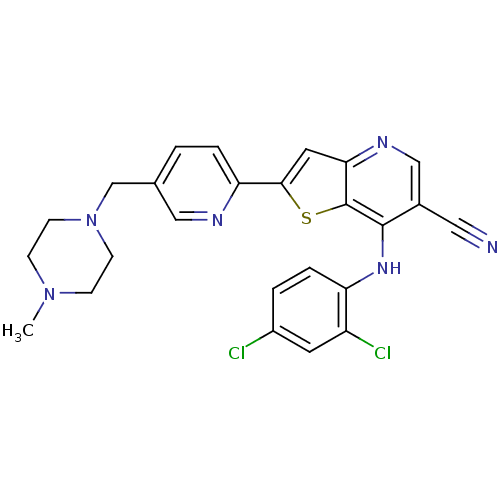
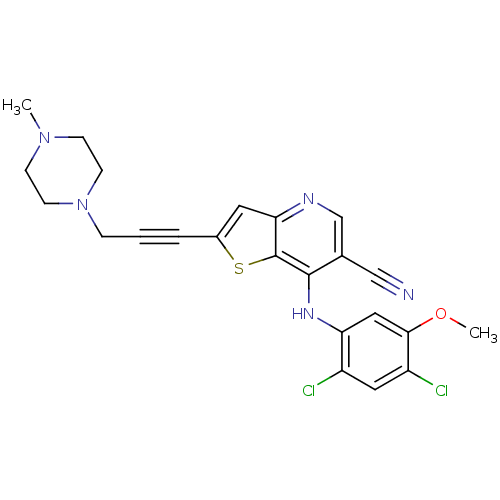
Report error Found 12 Enz. Inhib. hit(s) with all data for entry = 2
Affinity DataIC50: 7.20nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Kinase assays were performed using the europium/APC detection format (LANCE, Perkin Elmer). HTRF is based on the proximity of europium cryptate (dono...More data for this Ligand-Target Pair