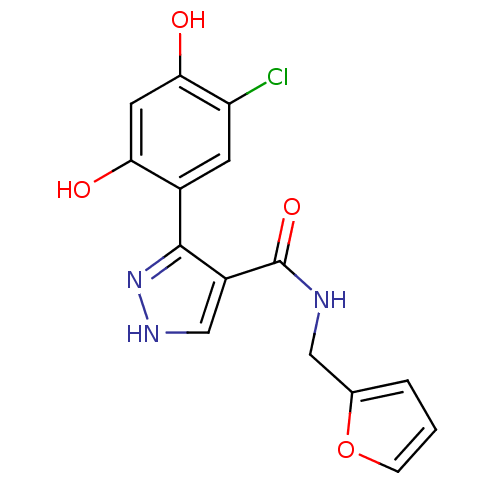
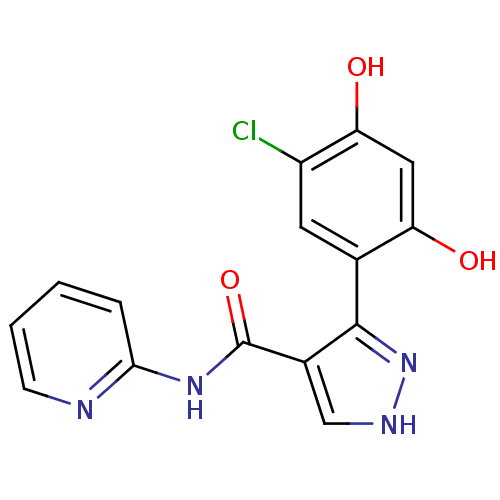
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 1949
Affinity DataIC50: 258nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 461nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 5.35E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 1.07E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 1.47E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 3.15E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 3.56E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair