Report error Found 156 Enz. Inhib. hit(s) with all data for entry = 50016930
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
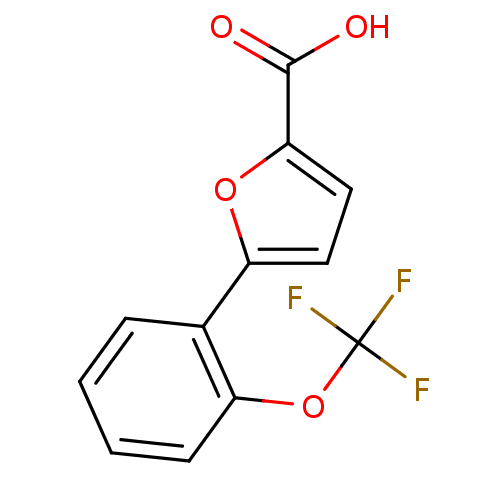
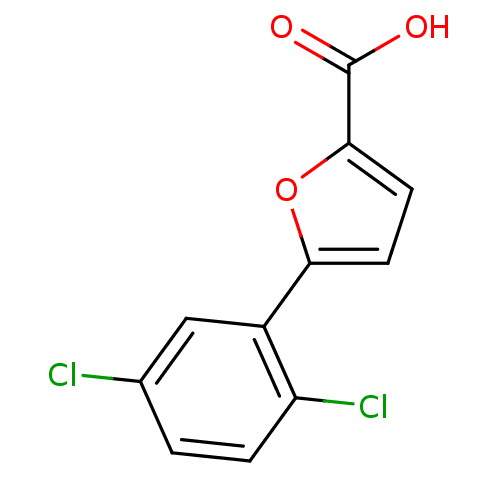
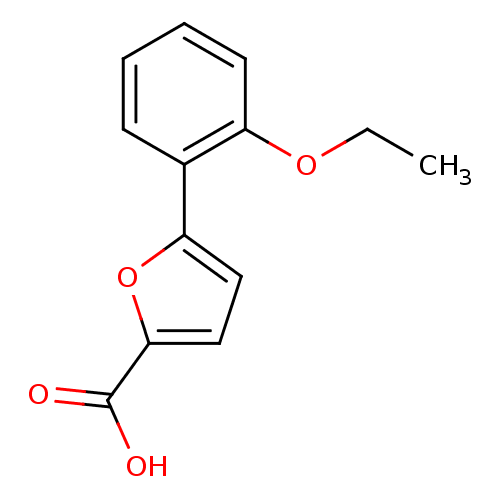
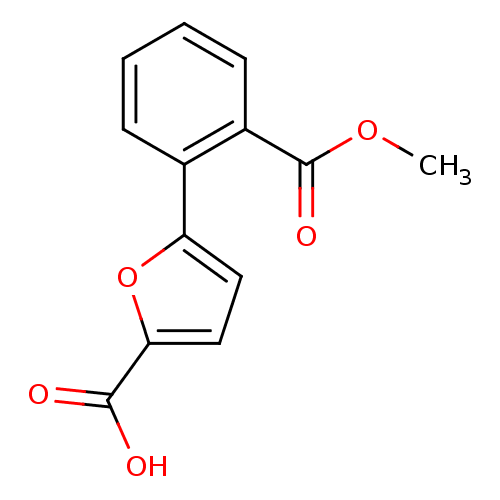
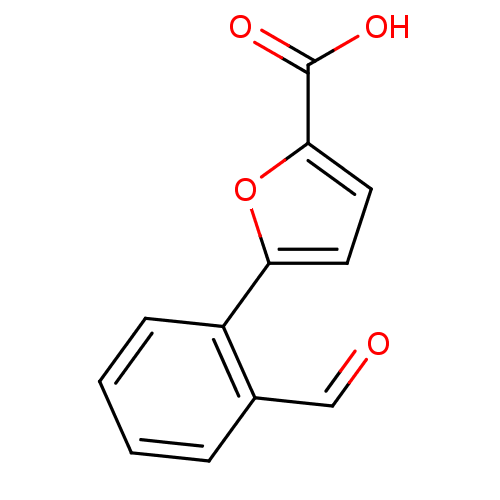
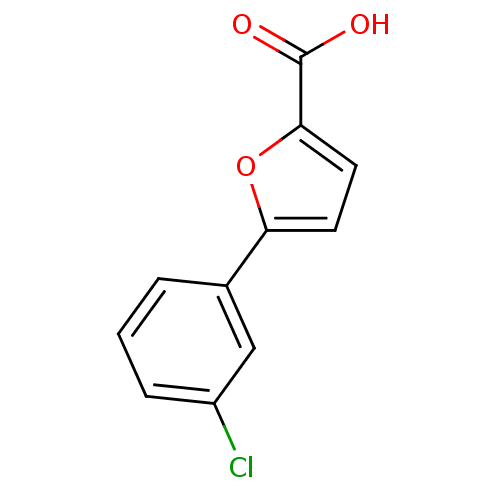
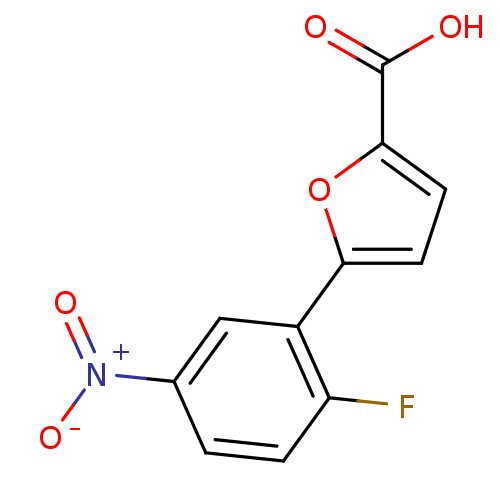
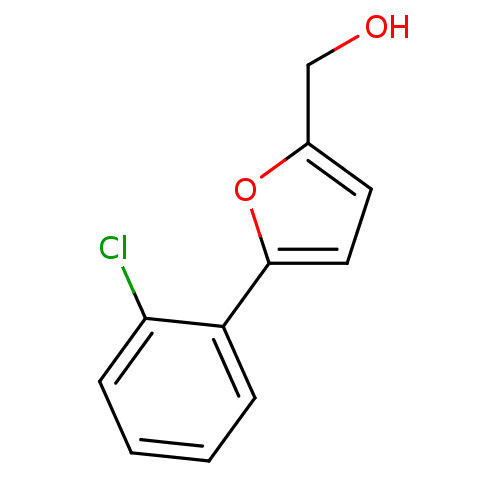
Affinity DataIC50: 290nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
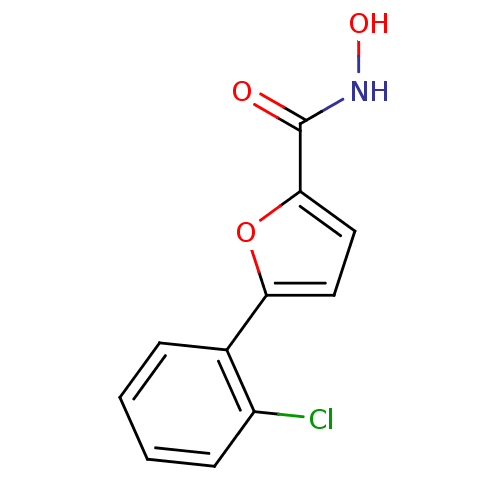
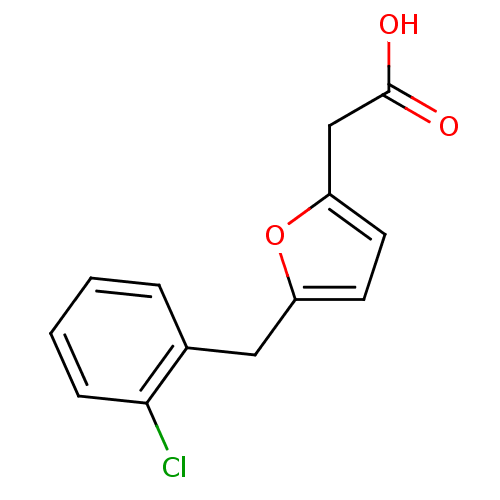
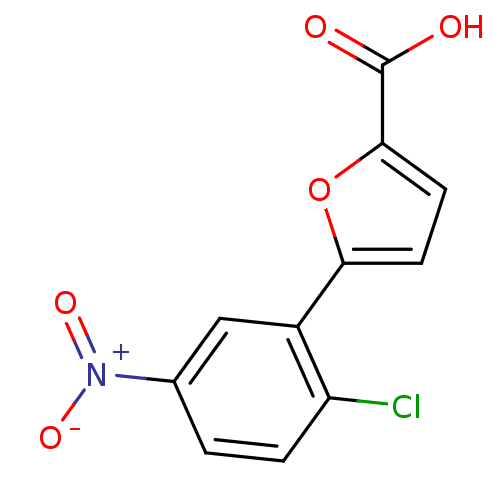
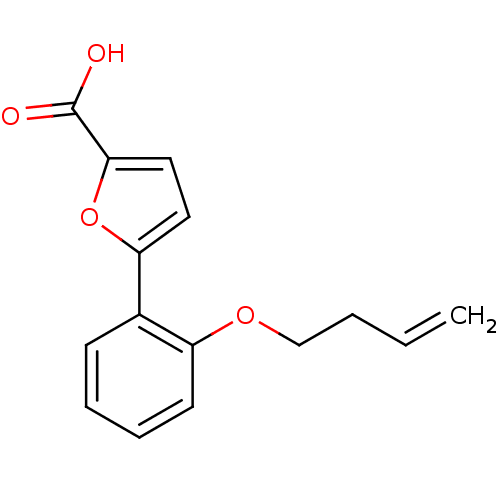
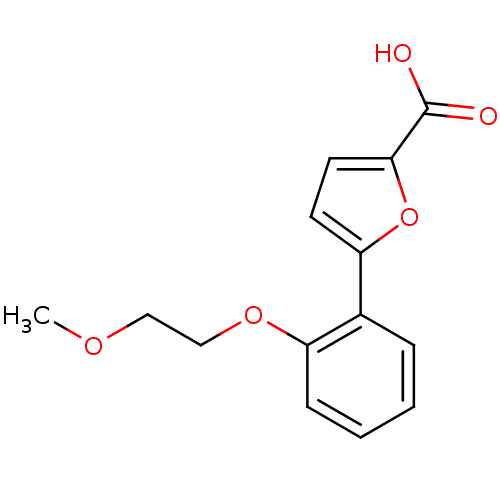
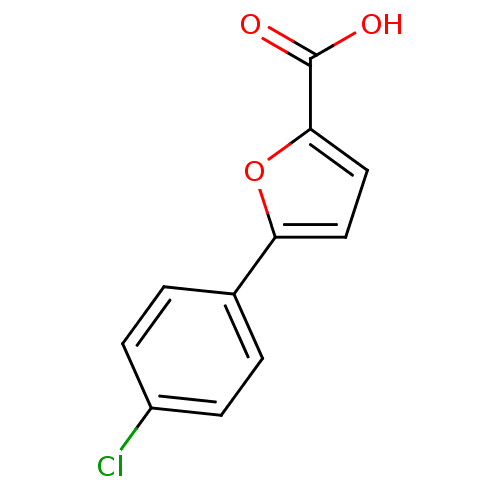
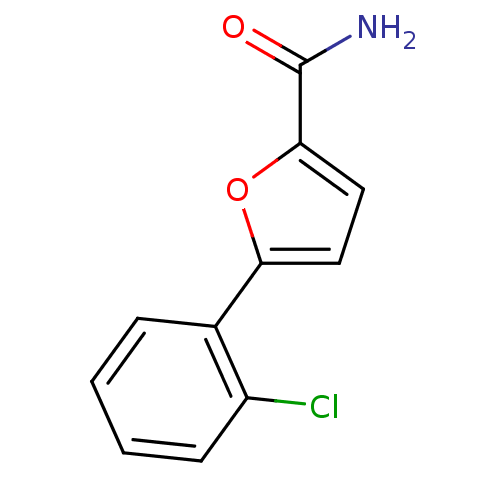
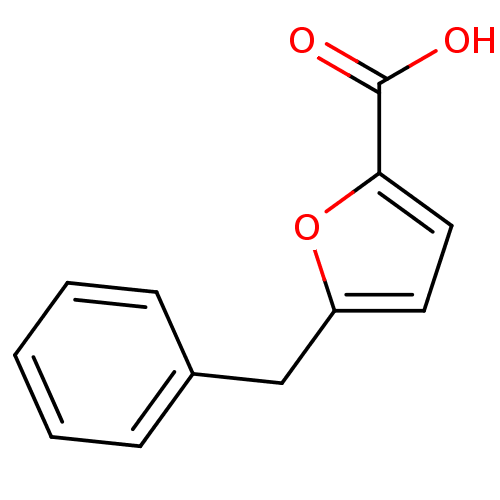
Affinity DataIC50: 368nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
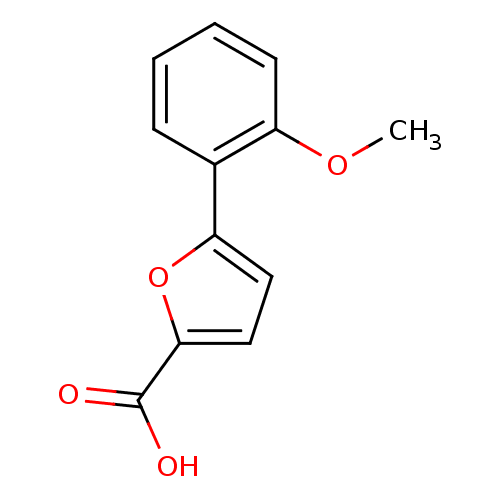
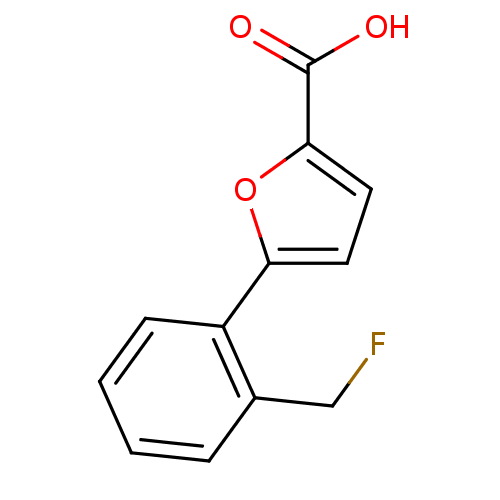
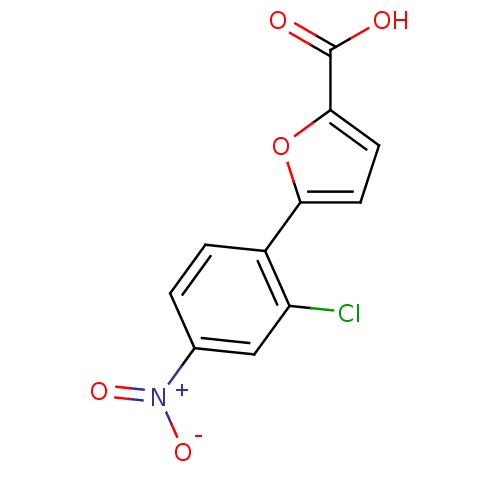
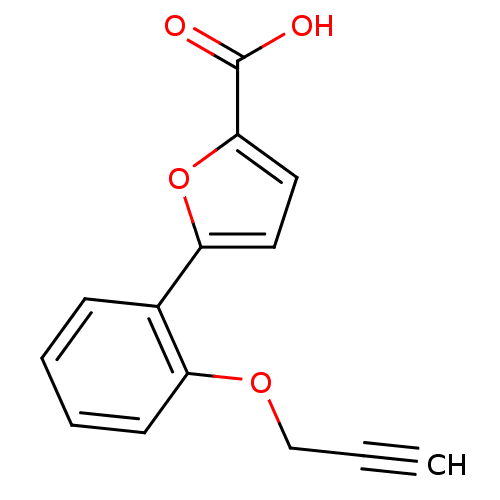
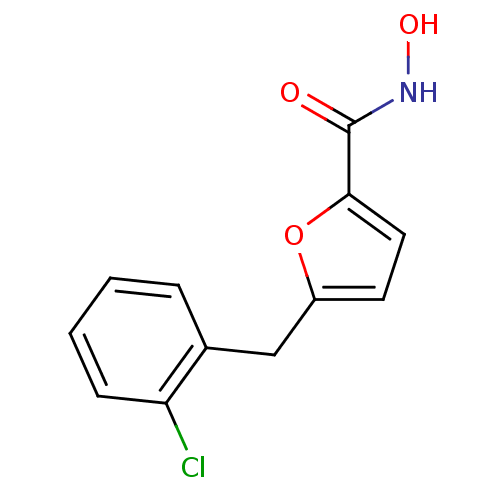
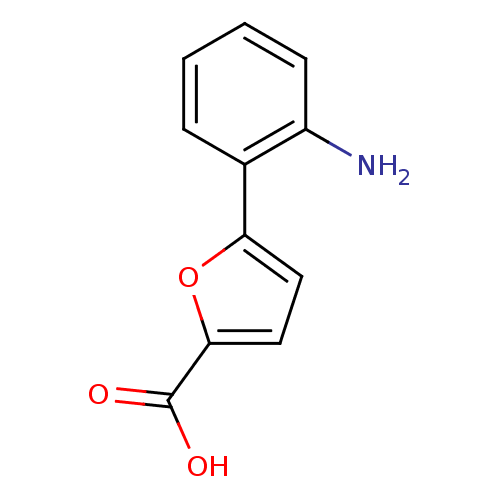
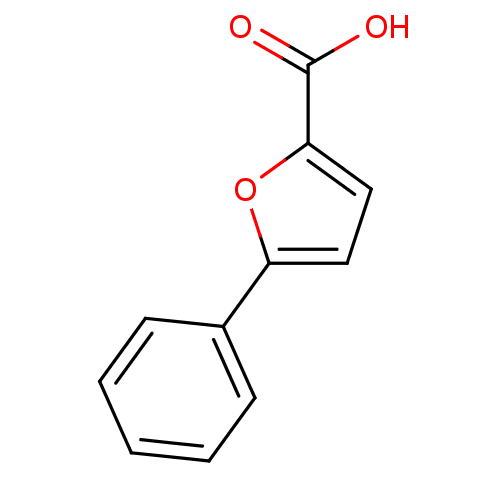
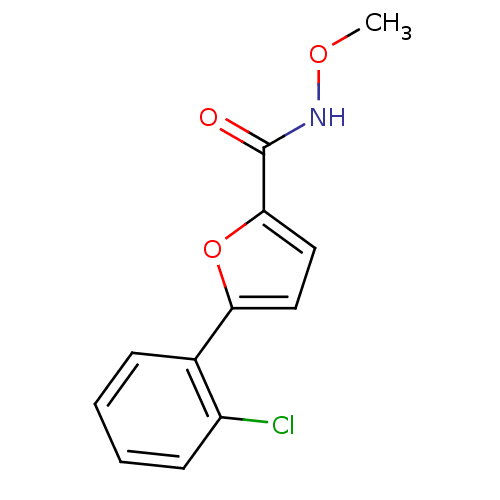
Affinity DataIC50: 469nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
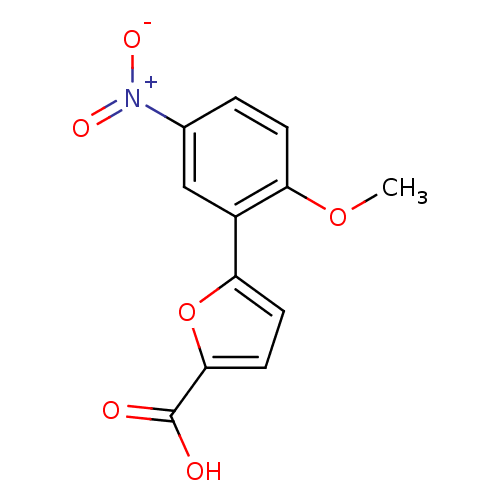
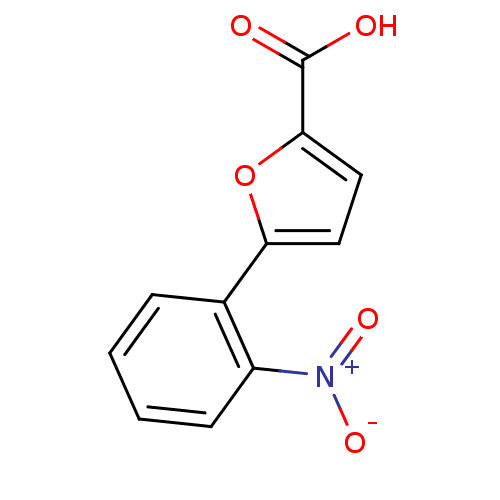
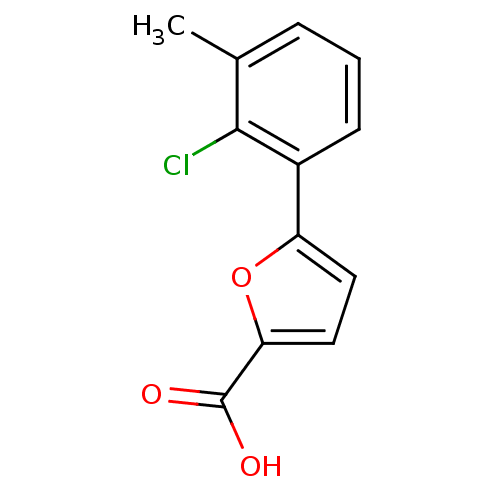
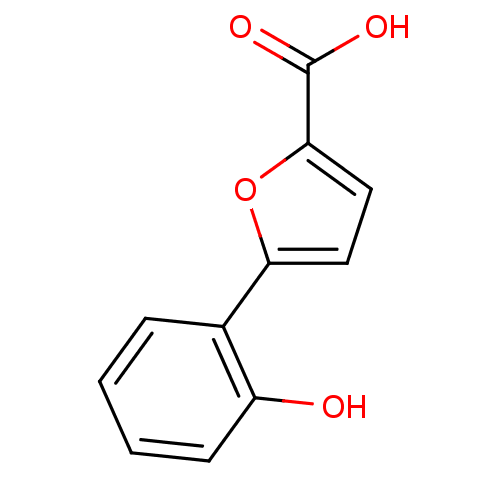
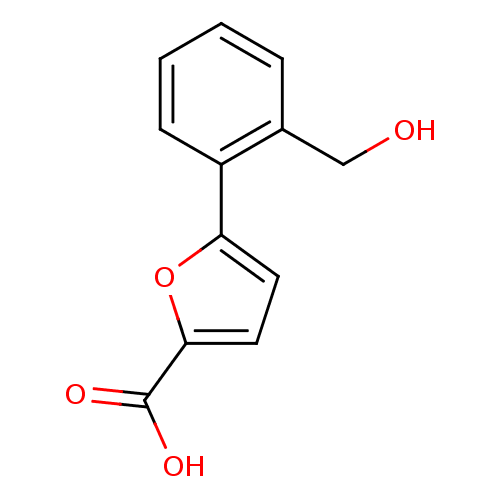
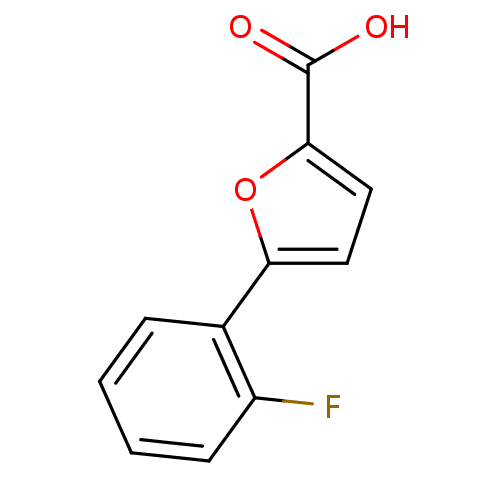
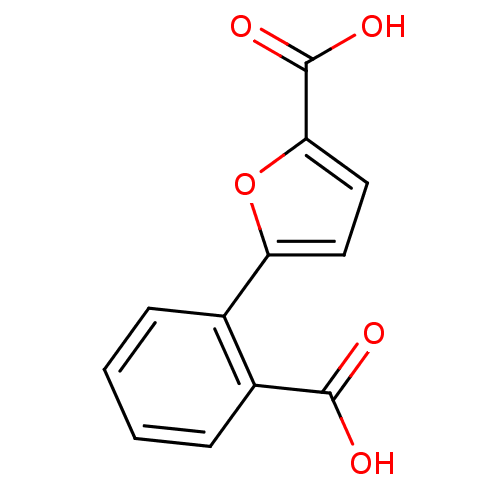
Affinity DataIC50: 511nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 558nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 642nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 693nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 784nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 793nMAssay Description:Inhibitory activity against Fe(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 893nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.08E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.11E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.13E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.22E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.32E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.56E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.71E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.47E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.65E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.61E+3nMAssay Description:Inhibitory activity against Co(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.71E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 5.78E+3nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 5.90E+3nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 6.56E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 6.92E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 7.17E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 9.45E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 9.84E+3nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.17E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.21E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.36E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.49E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.59E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.63E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.35E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.41E+4nMAssay Description:Inhibitory activity against Co(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.81E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.85E+4nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.11E+4nMAssay Description:Inhibitory activity against Fe(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.35E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.37E+4nMAssay Description:Inhibitory activity against Fe(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.45E+4nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.62E+4nMAssay Description:Inhibitory activity against Fe(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.83E+4nMAssay Description:Inhibitory activity against Co(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.84E+4nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibitory activity against Mn(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.25E+4nMAssay Description:Inhibitory activity against Co(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair
TargetMethionine aminopeptidase(Escherichia coli (strain K12))
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Graduate School of Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.29E+4nMAssay Description:Inhibitory activity against Ni(II) form of Escherichia coli MetAPMore data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)