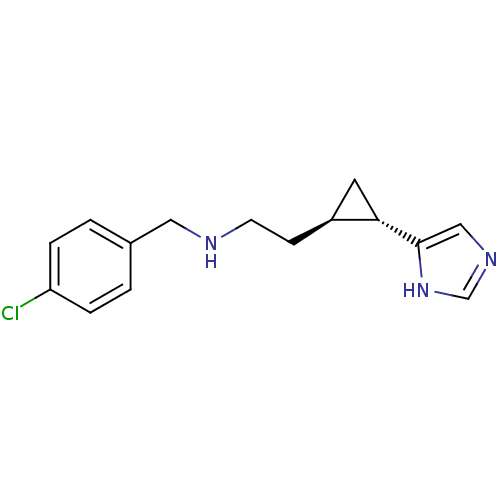
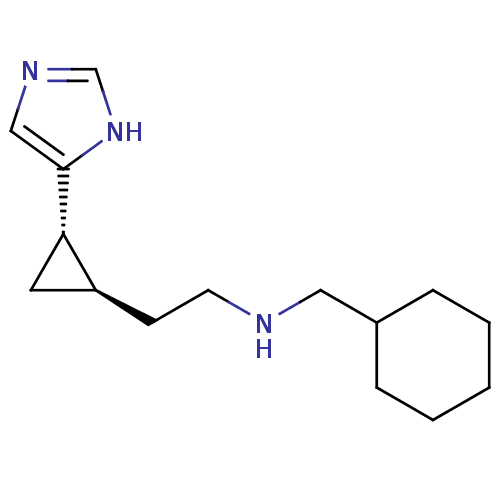
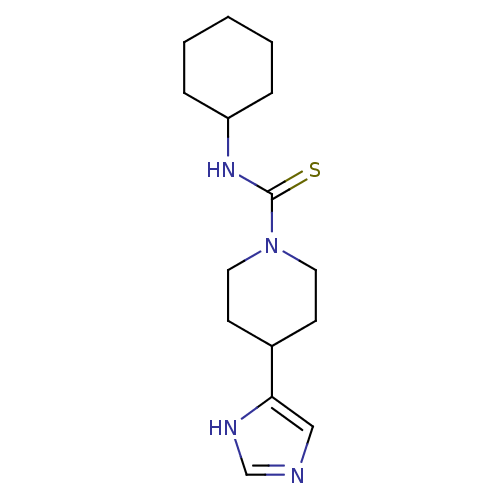
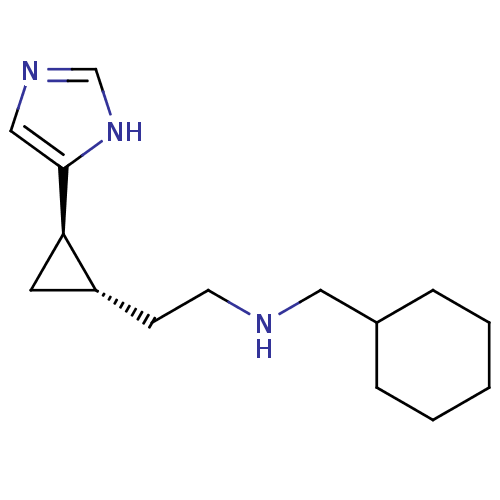
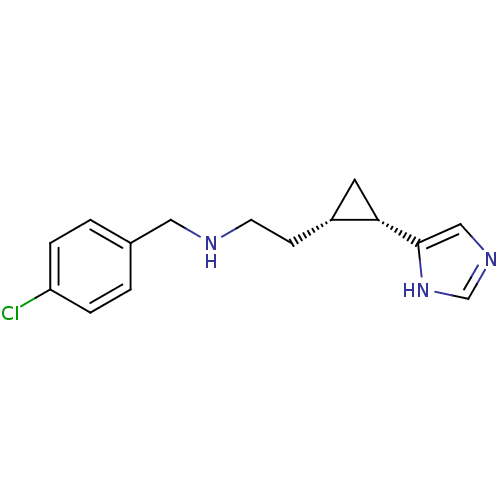
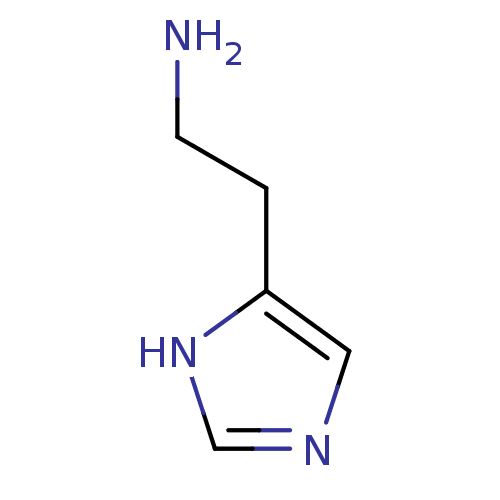
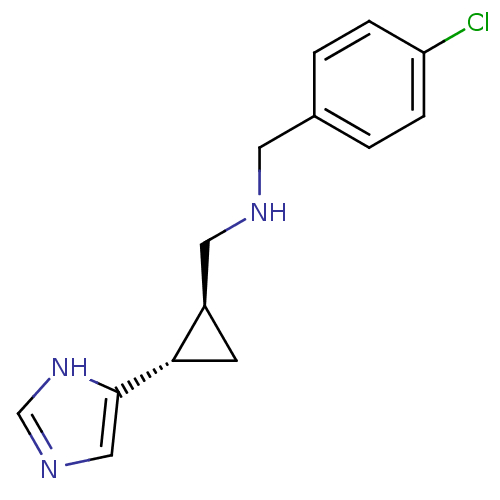
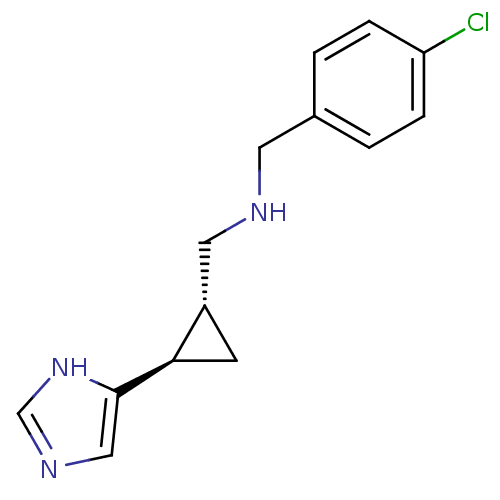
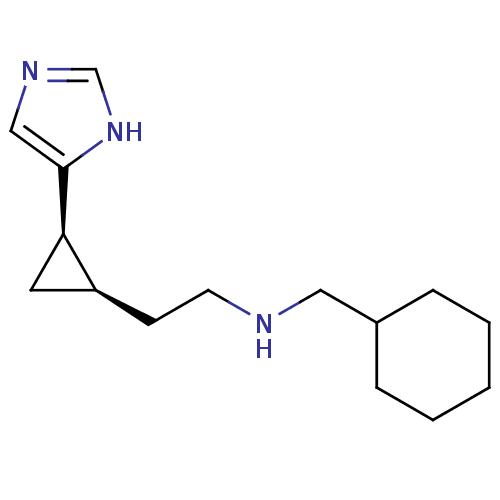
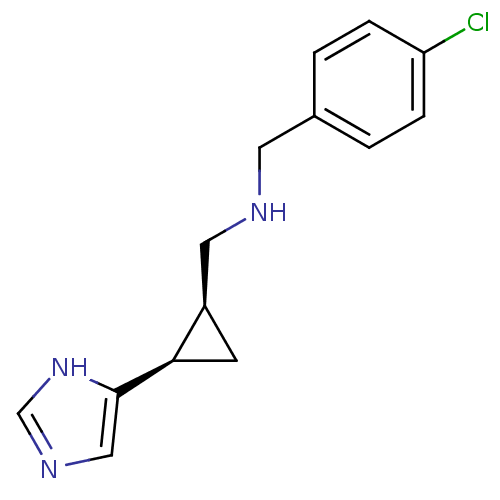
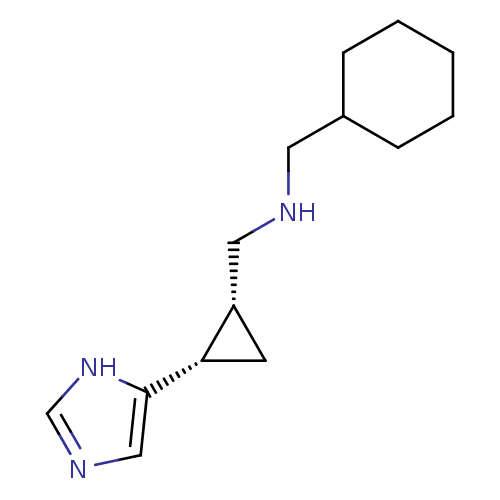
Report error Found 64 Enz. Inhib. hit(s) with all data for entry = 50018304
Affinity DataKi: 3.60nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMAssay Description:Antagonist activity at human H3 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataKi: 13.9nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Antagonist activity at human H3 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Antagonist activity at human H3 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataKi: 20.5nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 31.2nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 34.4nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Antagonist activity at human H3 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataKi: 37.2nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 43.7nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 45nMAssay Description:Agonist activity at human H1 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 51.1nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 88nMAssay Description:Agonist activity at human H3 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 88nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 90.4nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 103nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Antagonist activity at human H4 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataKi: 115nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 118nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 124nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 143nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 177nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 203nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 289nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 650nMAssay Description:Agonist activity at human H2 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:Antagonist activity at human H4 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataIC50: 860nMAssay Description:Antagonist activity at human H4 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]histamine form human H4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]Nalpha-methylhistamine form human H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Antagonist activity at human H4 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataEC50: 1.80E+3nMAssay Description:Agonist activity at human H4 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+3nMAssay Description:Antagonist activity at human H1 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataIC50: 8.50E+3nMAssay Description:Antagonist activity at human H1 receptor expressed in 293-EBNA cells assessed as inhibition of histamine agonist activity by luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Agonist activity at human H4 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Agonist activity at human H1 receptor expressed in 293-EBNA cells by luciferase reporter gene assayMore data for this Ligand-Target Pair