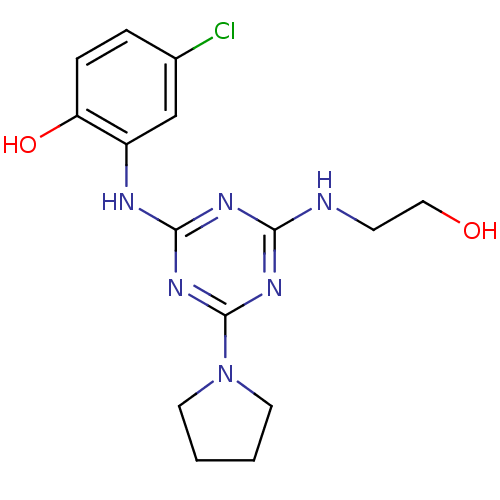
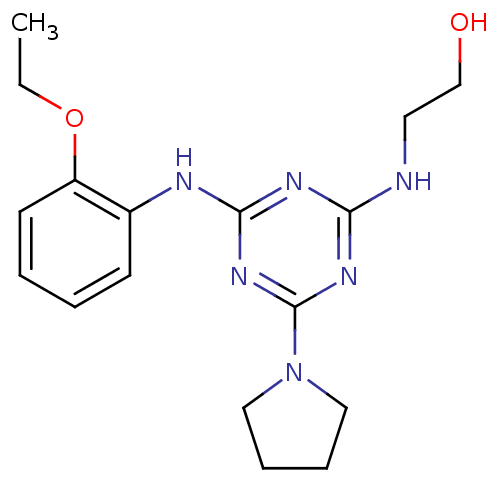
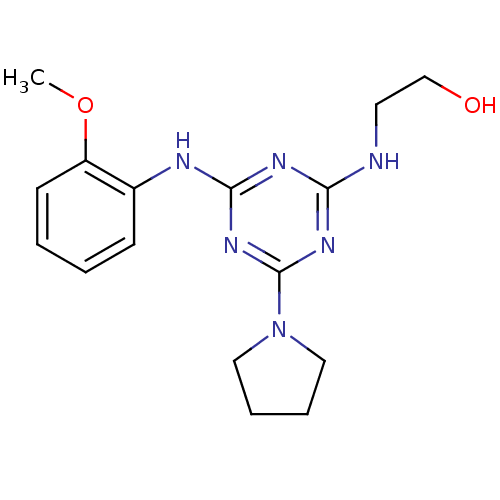
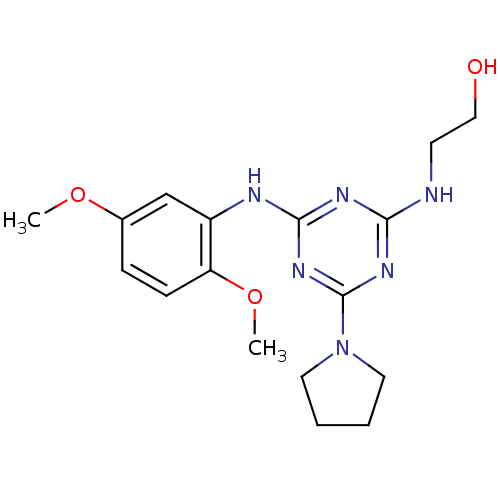
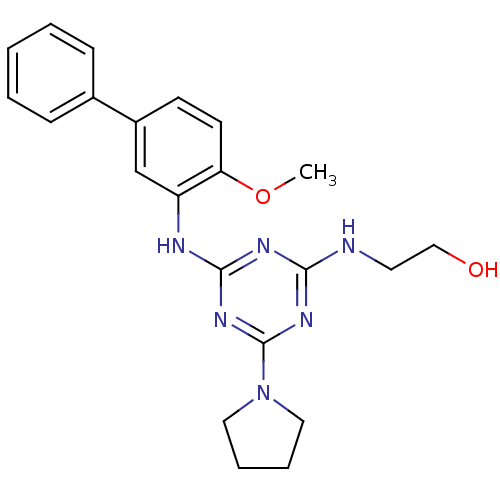
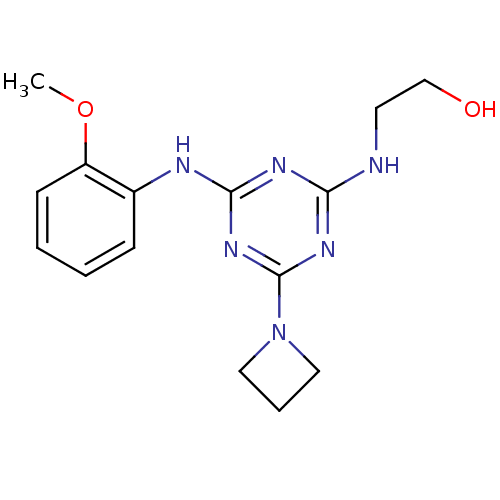
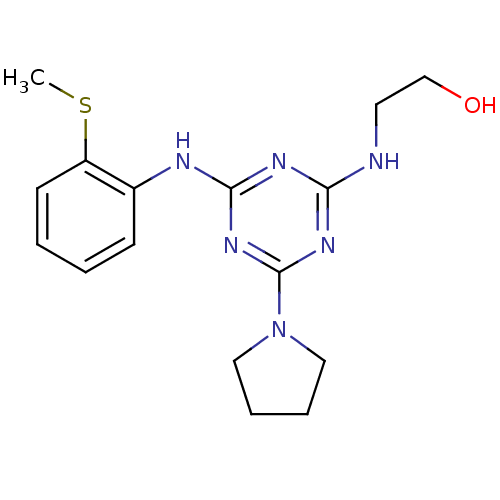
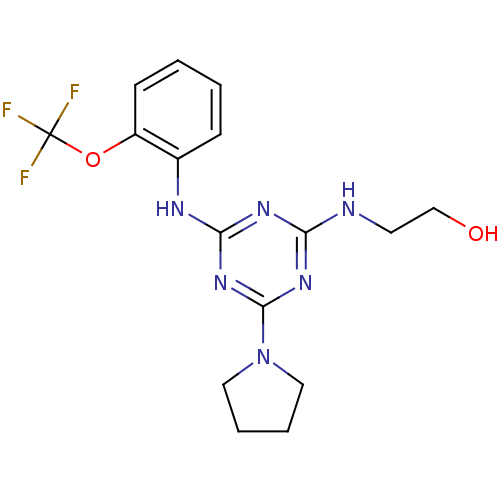
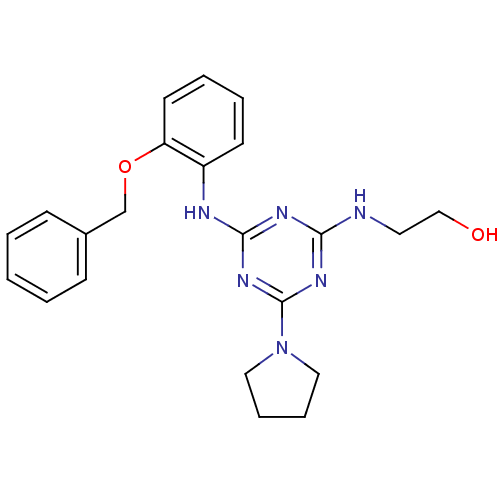
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 2241
Affinity DataIC50: 330nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 730nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 770nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.31E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.39E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.65E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 3.34E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 3.94E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 6.19E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 6.34E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 6.69E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 7.75E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 7.77E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 8.25E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+3nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.65E+4nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 1.88E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.07E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+4nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 3.11E+4nMpH: 5.9 T: 2°CAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair
Affinity DataIC50: 3.14E+4nMAssay Description:Enzyme activity was determined as the production of fluorescent resorufin from the substrate. The fluorescence was measured (excitation 570 nm, emiss...More data for this Ligand-Target Pair