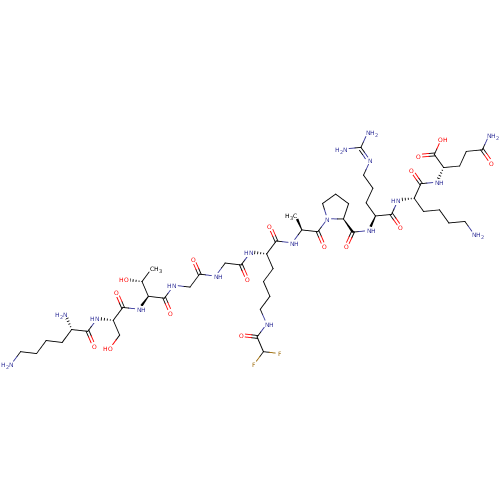
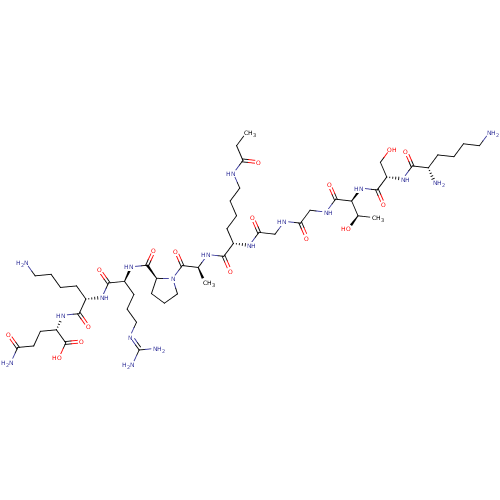
Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 50038598
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 8.60E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 1.60E+4nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase HST2(Baker's yeast)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 2.10E+4nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair