Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 50048850
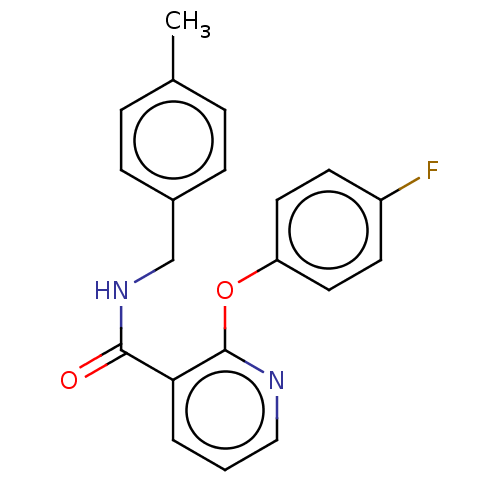
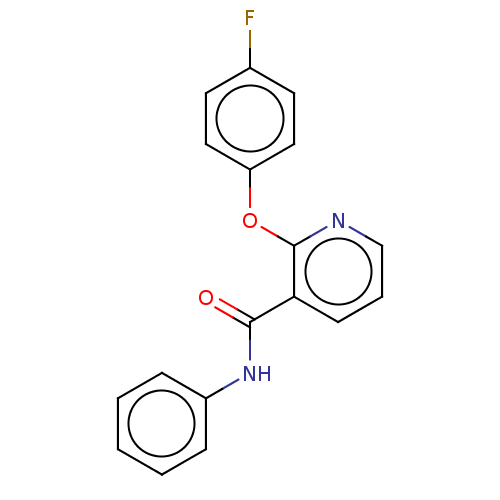
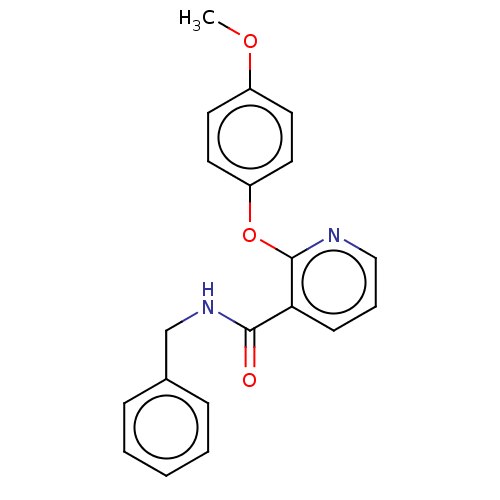
Affinity DataIC50: 220nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
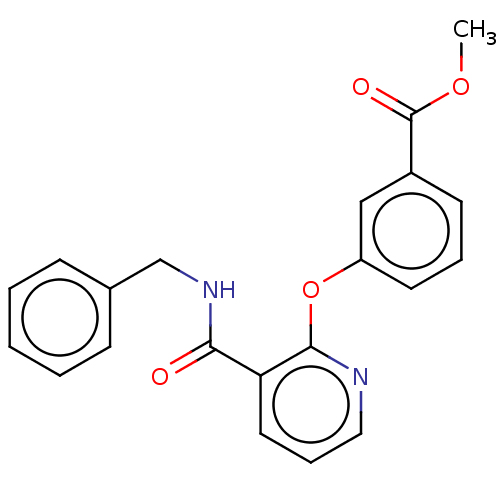
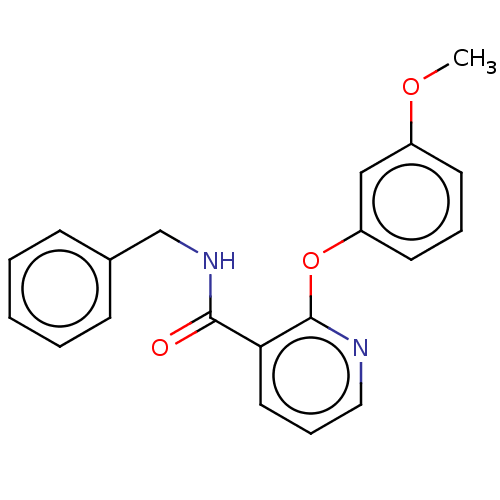
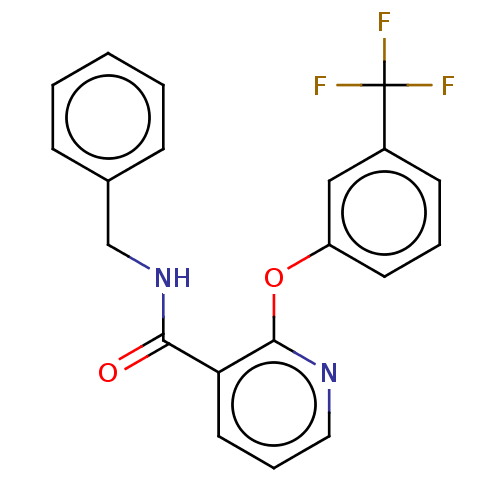
Affinity DataIC50: 540nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
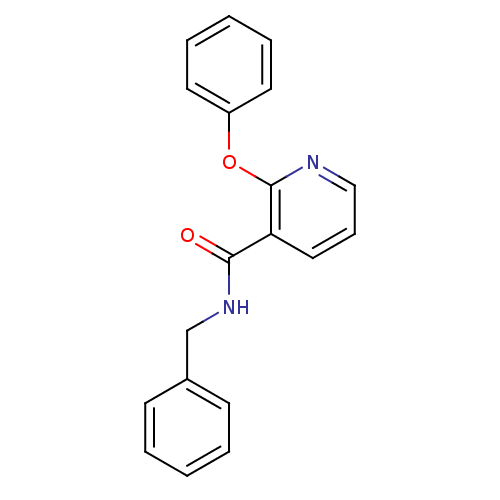
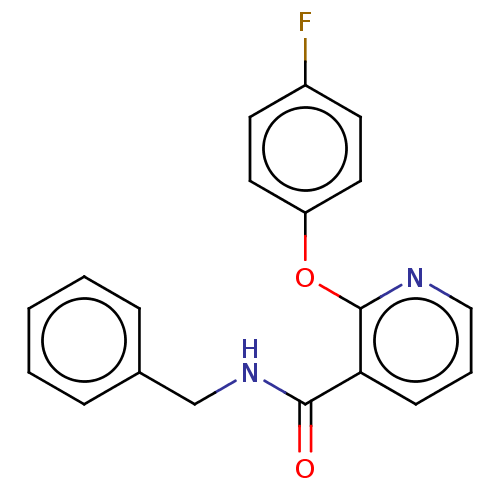
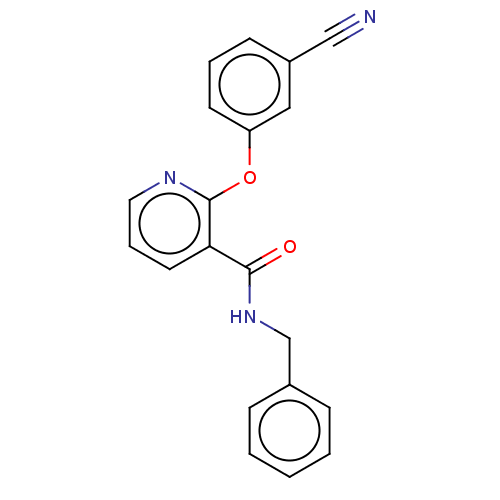
Affinity DataIC50: 730nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
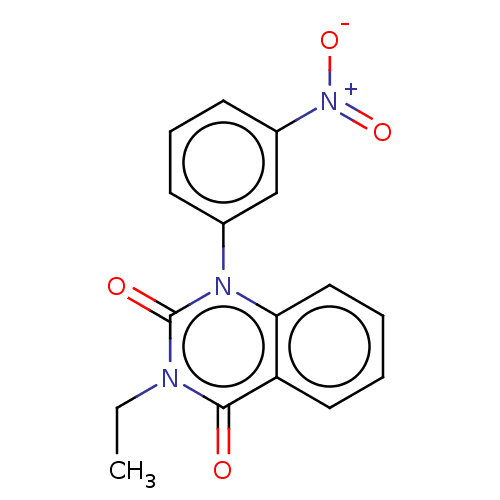
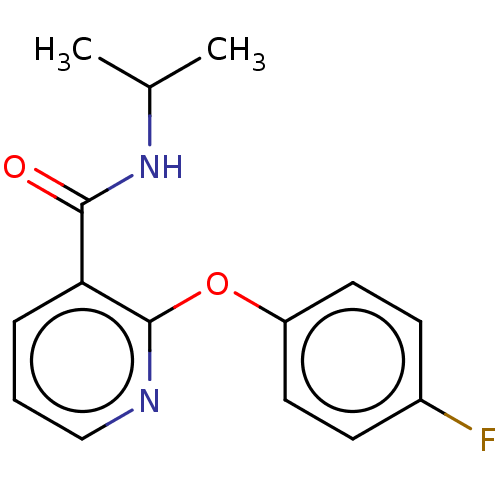
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 9.80E+3nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of [3H]cAMP (NEN NET-275) binding to Calcium-Independent Phosphodiesterase from rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of [3H]rolipram binding to rat brain membranes.More data for this Ligand-Target Pair