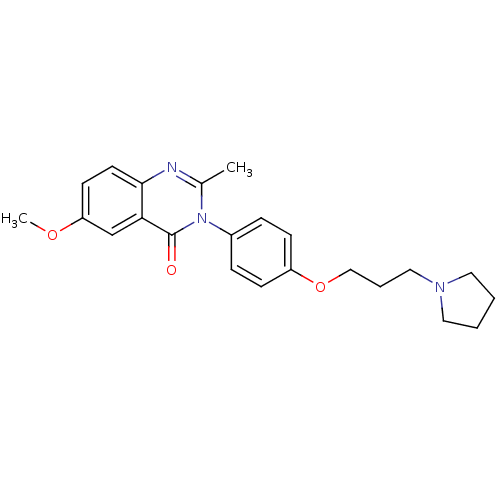
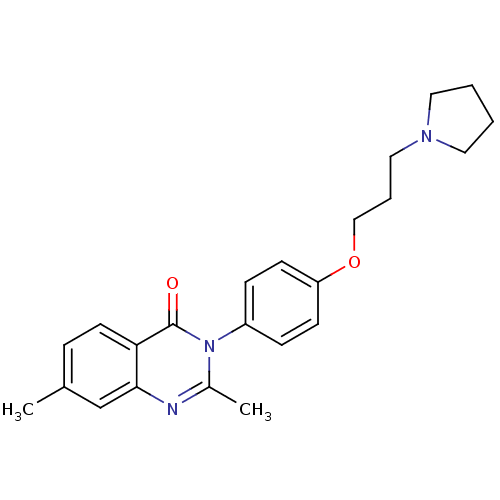
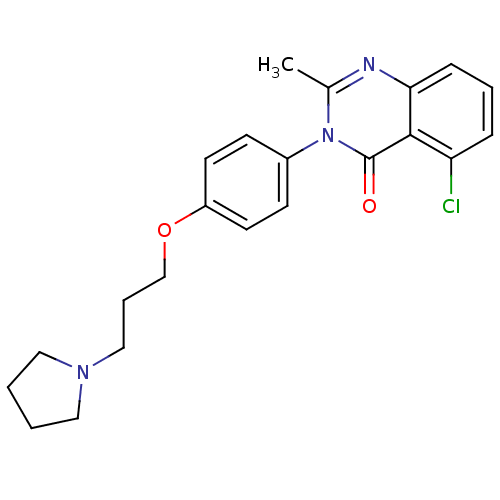
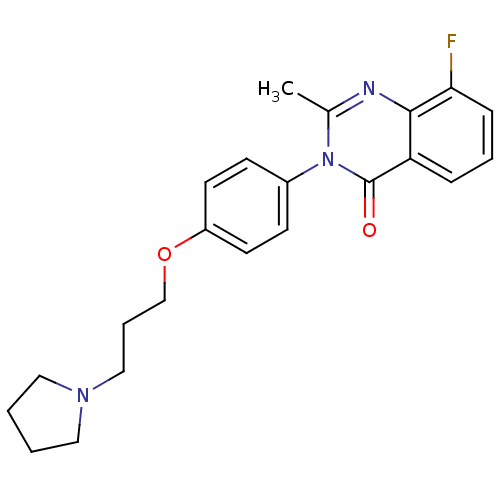
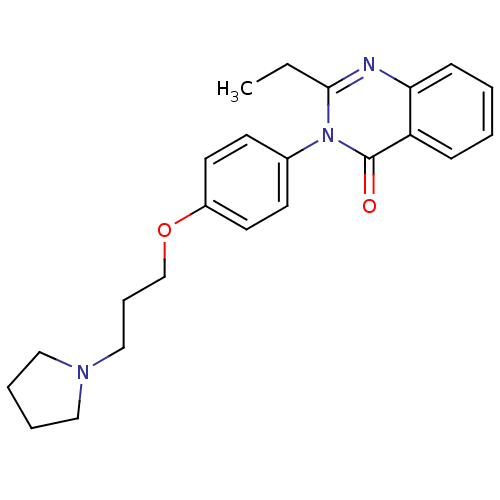
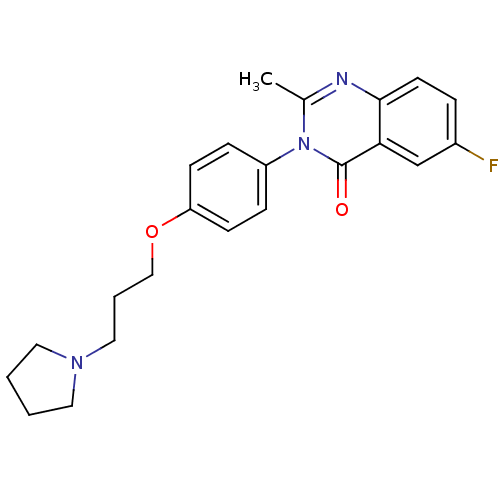
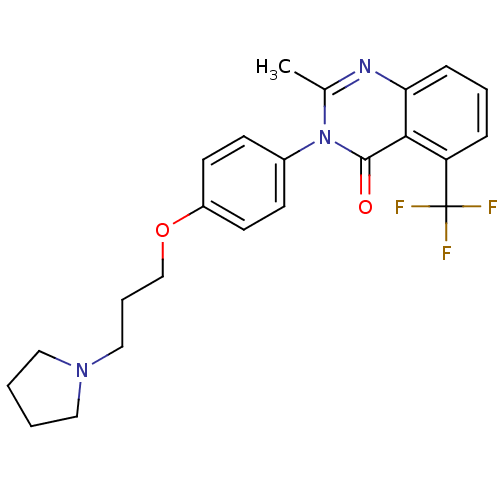
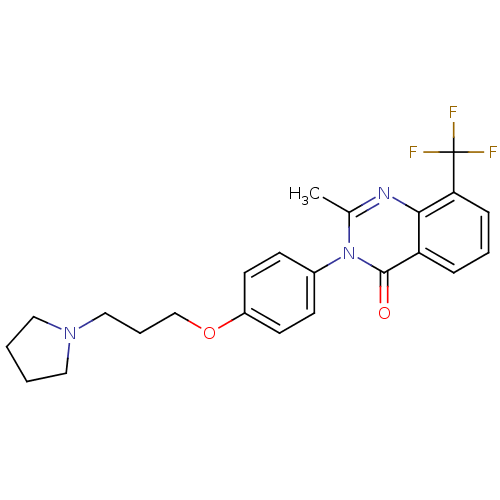
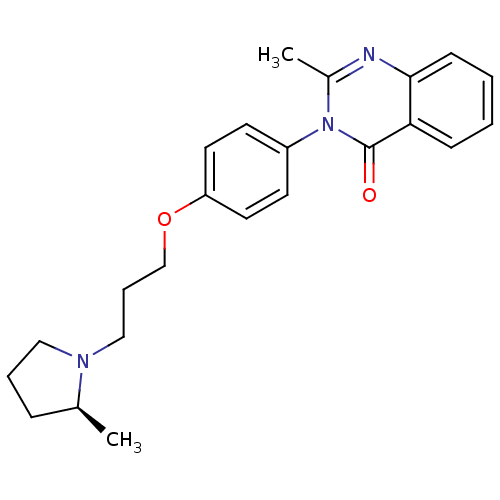
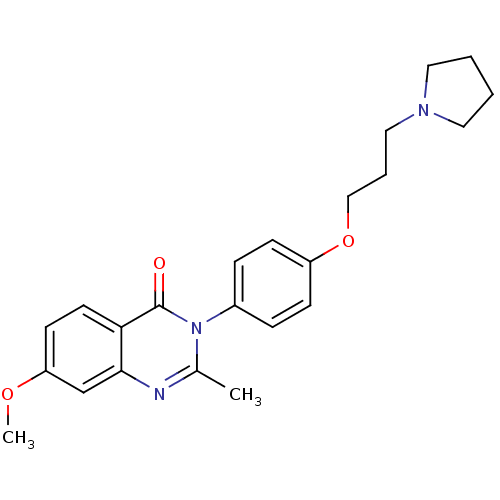
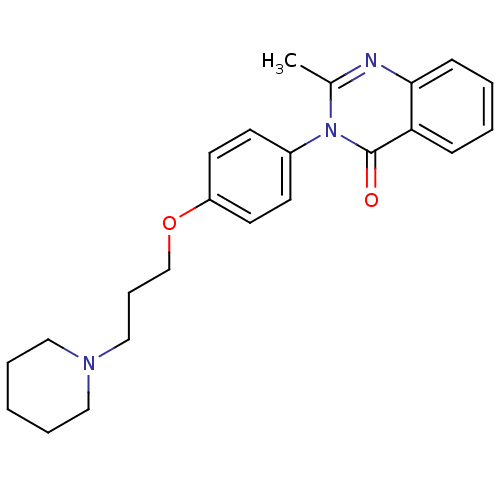
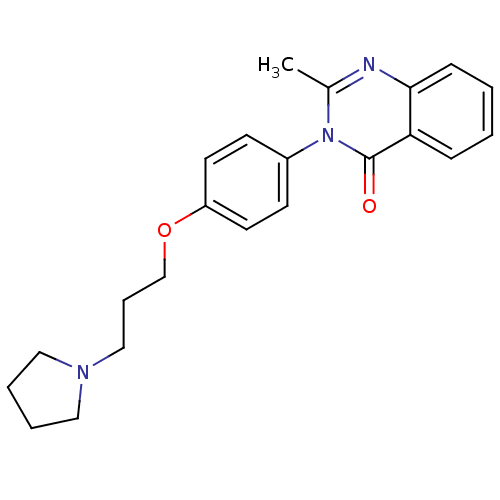
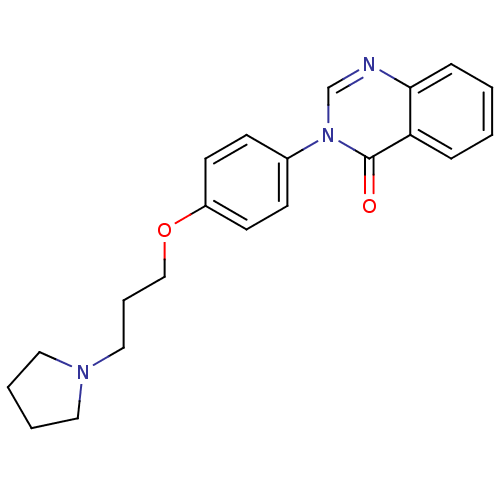
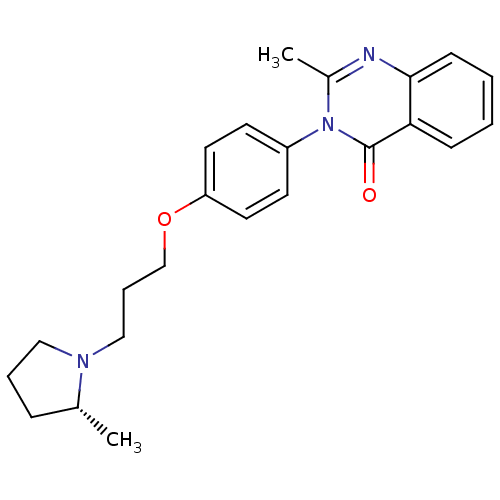
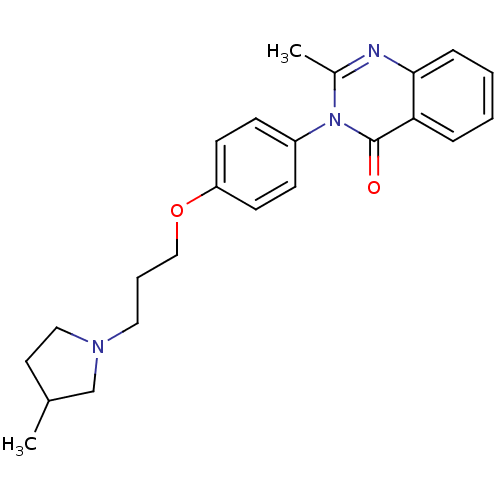
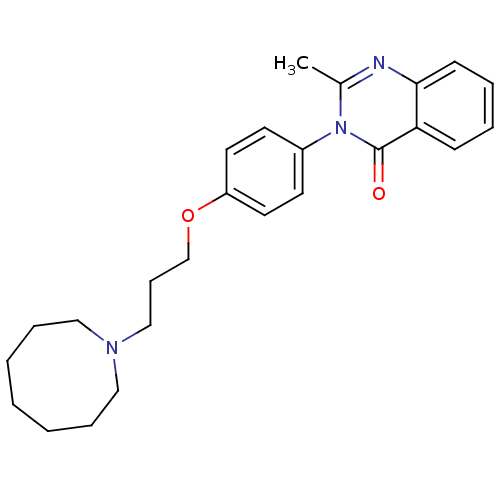
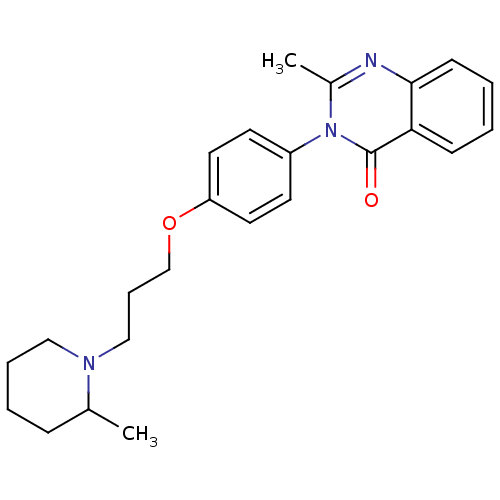
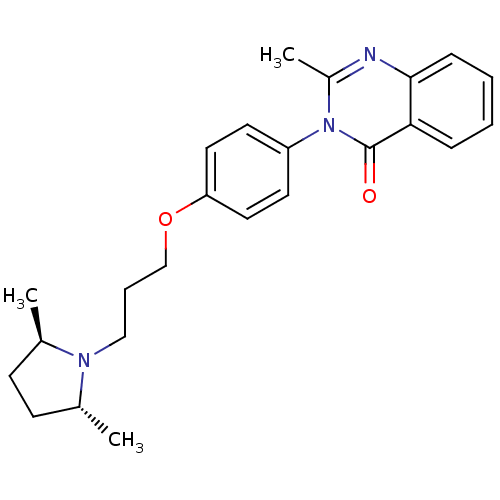
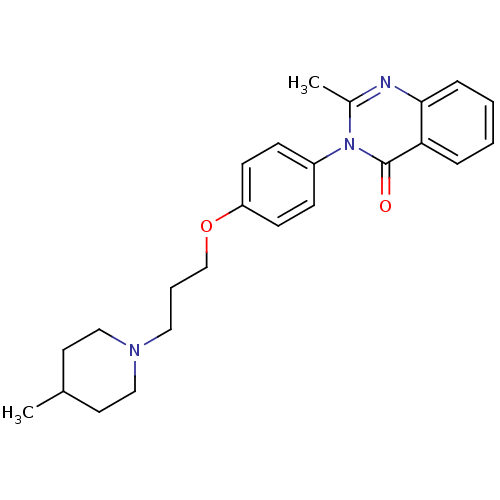
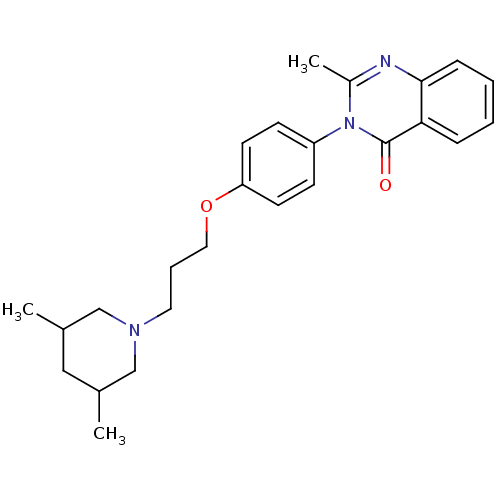
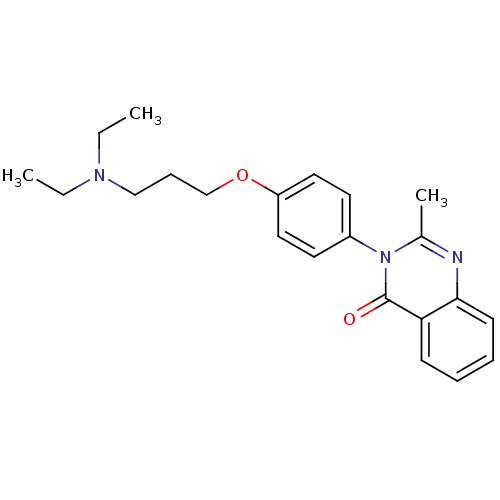
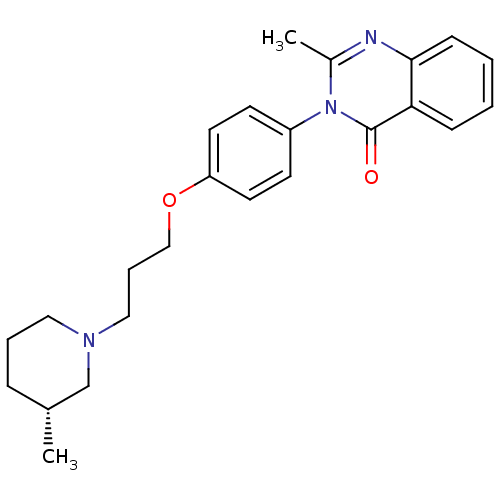
Report error Found 75 Enz. Inhib. hit(s) with all data for entry = 50026555
Affinity DataIC50: 0.660nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 0.690nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 0.720nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 0.810nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 0.960nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 0.980nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rhesus monkey histamine H3 receptor expressed in HEK293T cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat histamine H3 receptor expressed in HEK293 cells coexpressed with CRE-beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 350nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 710nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.80E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.90E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 5.30E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.00E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.00E+3nMAssay Description:Displacement of [35S]N-[(4R)-1'-[(2R)-6-cyano-1,2,3,4-tetrahydro-2-naphthalenyl]-3,4-dihydro-4-hydroxyspiro[2H-1-benzopyran-2,4'-piperidin]-6-yl]meth...More data for this Ligand-Target Pair