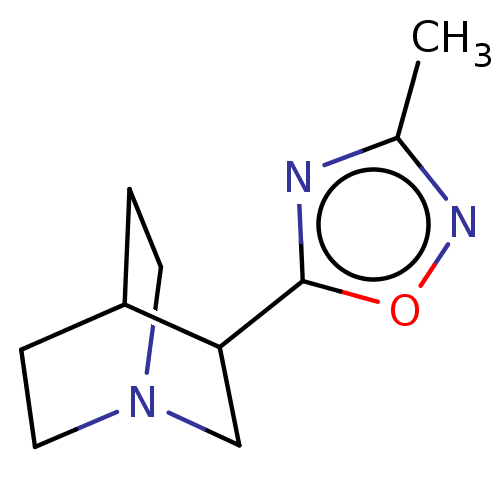
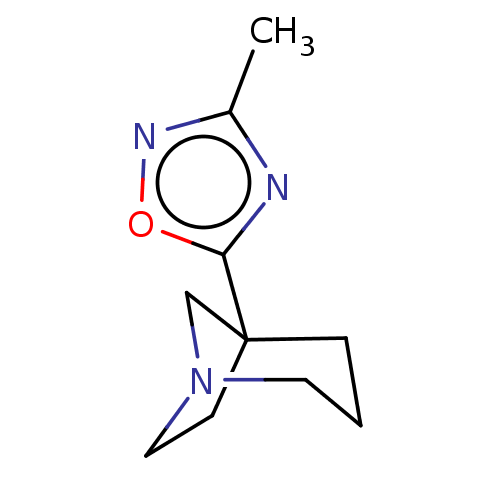
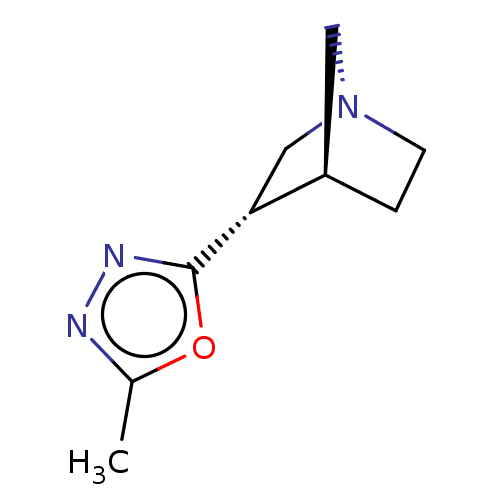
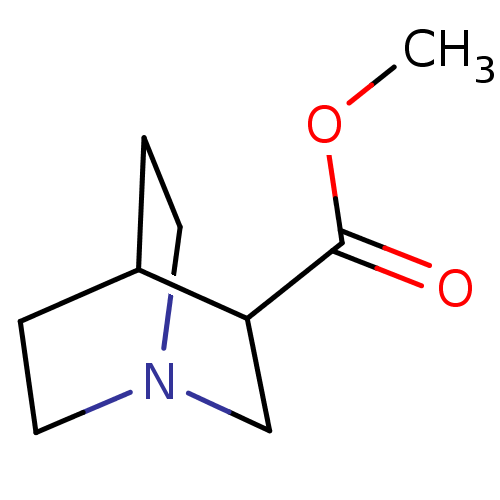
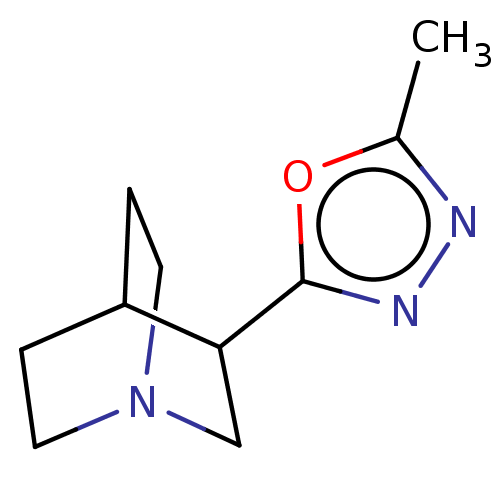
Report error Found 44 Enz. Inhib. hit(s) with all data for entry = 50048866
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.80nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:The compound was tested in vitro for central Muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 28nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 42nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 48nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 77nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 92nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 180nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 230nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 260nMAssay Description:The compound was tested for muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Value ranges from 250...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 260nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 520nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 980nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]oxotremorine-M from rat cerebral cortex; Valu...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.73E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3.50E+3nMAssay Description:The compound was tested in vitro for central Muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 4.20E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 4.80E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 6.40E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 9.40E+3nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 1.90E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.30E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.90E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3.10E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 5.10E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 6.20E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 7.00E+4nMAssay Description:The compound was tested in vitro for central muscarinic acetylcholine receptor affinity to displace [3H]quinuclidinyl benzilate from rat cerebral cor...More data for this Ligand-Target Pair

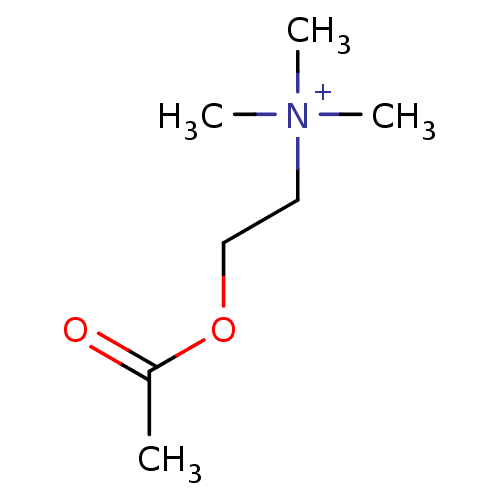
 3D Structure (crystal)
3D Structure (crystal)