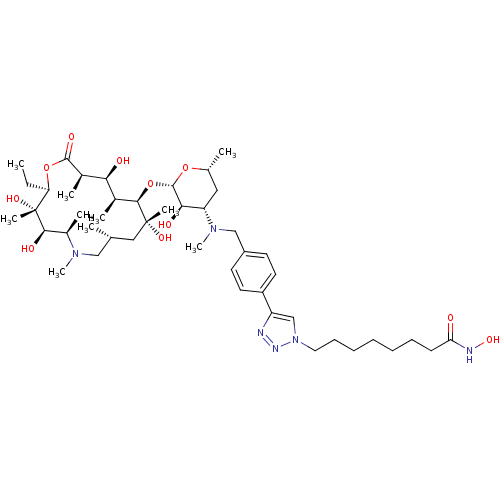
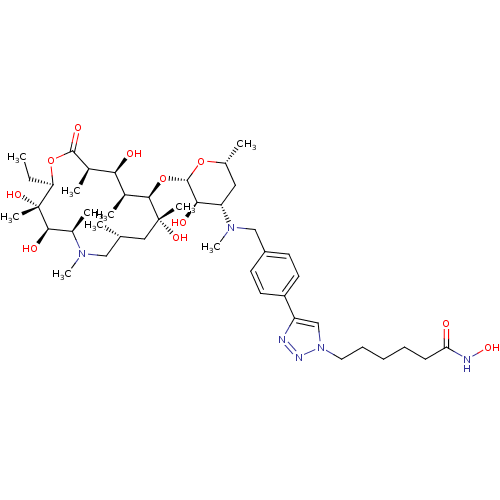
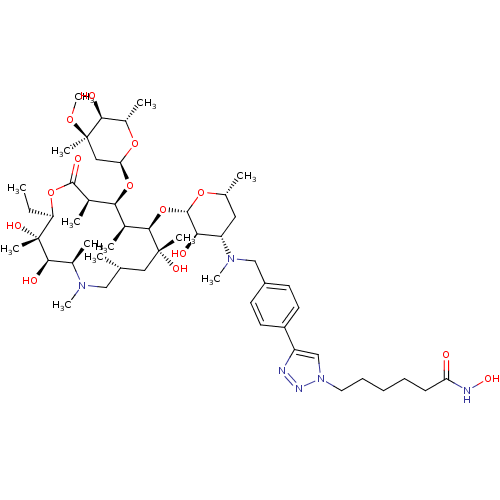
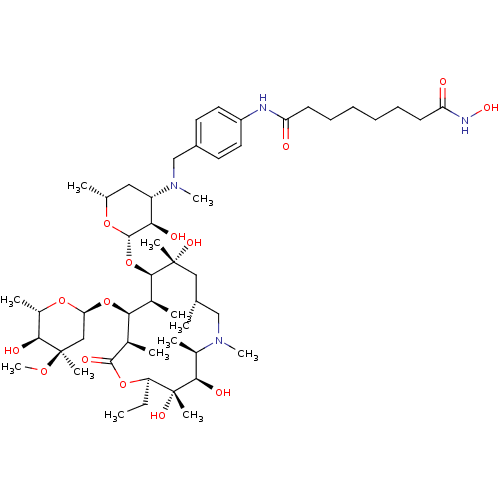
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 3041
Affinity DataIC50: 1.90nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 10.6nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 13.9nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 44.3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 55.6nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 58.9nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 72.4nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 88.8nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 91.6nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 107nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 123nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 146nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 223nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 227nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 994nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.39E+3nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.86E+3nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.89E+3nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.32E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 3.74E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 3.99E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.42E+3nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.73E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.75E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 5.88E+3nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 6.68E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 6.78E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 7.13E+3nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMT: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMpH: 8.0 T: 2°CAssay Description:The Fluor-de-Lys HDAC activity assay kit (Biomol) was used. Purified recombinant HDAC enzyme was incubated with Fluor-de-Lys substrate in the presenc...More data for this Ligand-Target Pair


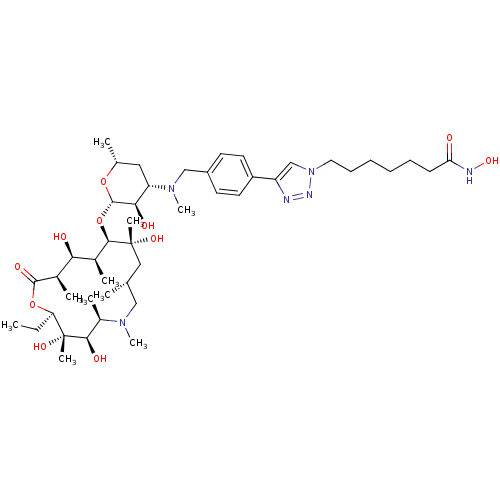

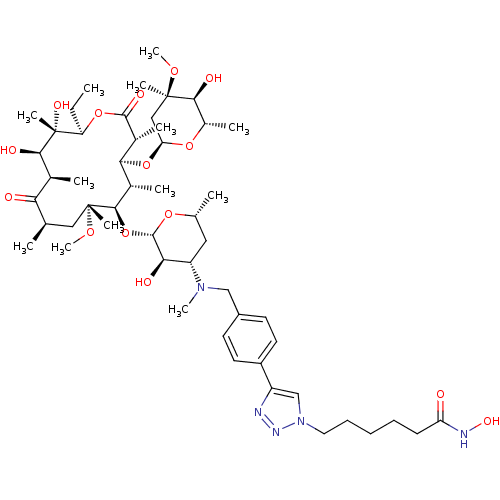
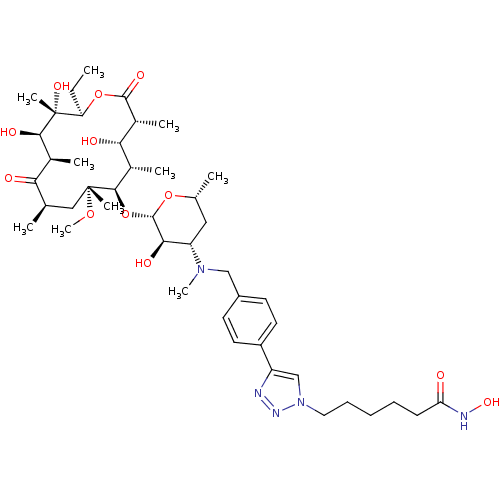
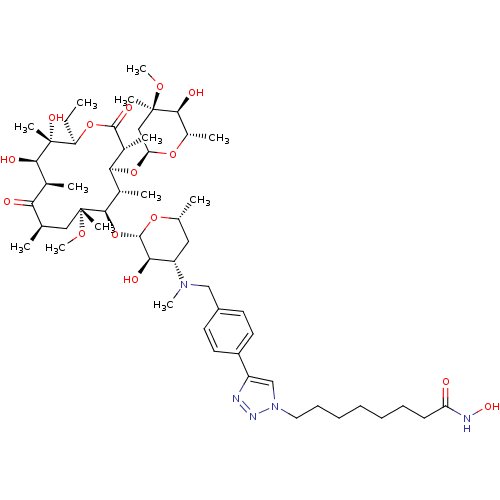

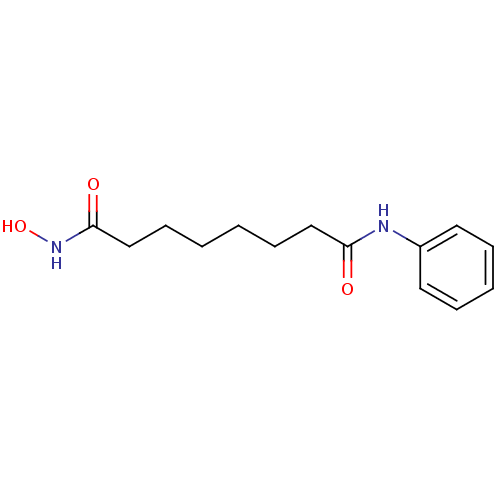
 3D Structure (crystal)
3D Structure (crystal)