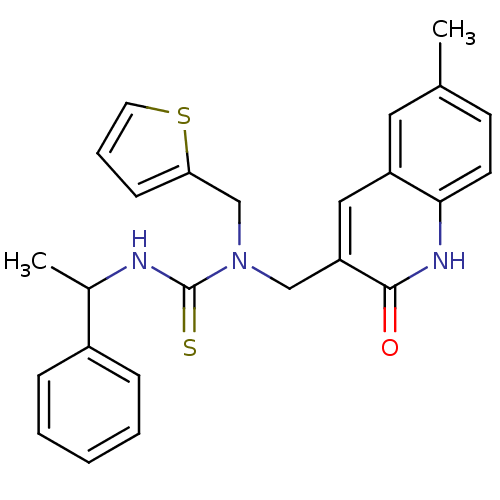
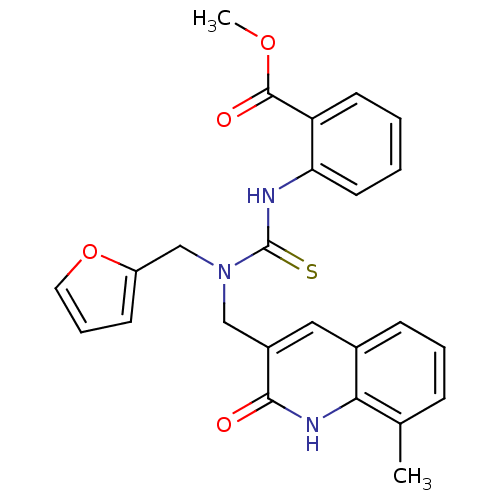
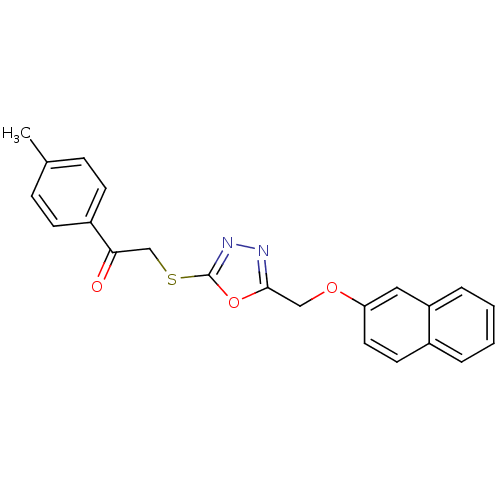
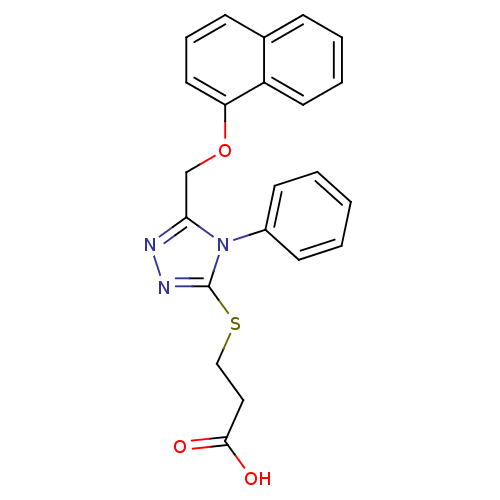
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 4727
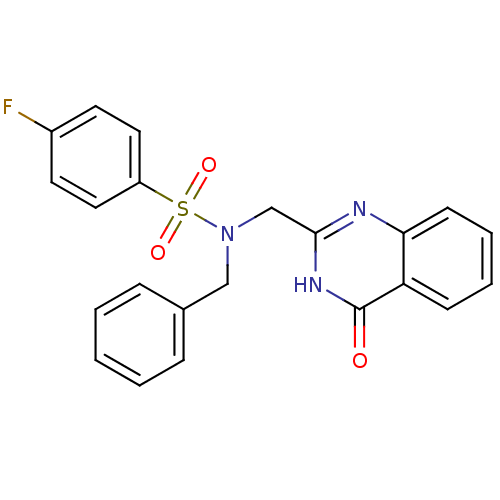
Affinity DataKi: 580nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
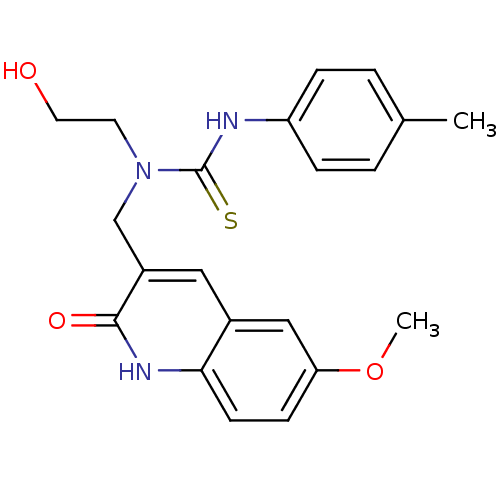
Affinity DataKi: 1.20E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
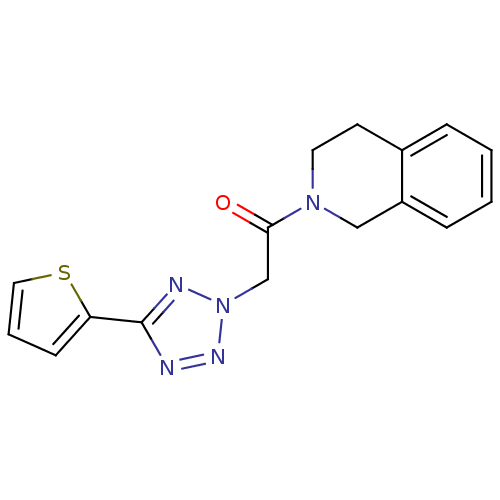
Affinity DataKi: 1.60E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
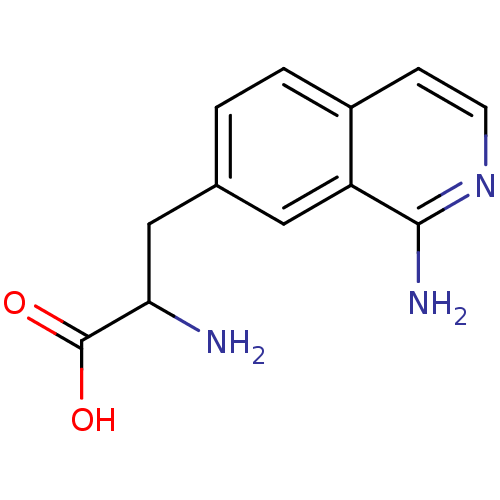
Affinity DataKi: 1.60E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 3.80E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 6.50E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 7.40E+3nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.04E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.33E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.43E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 2.05E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 2.25E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 3.50E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 3.70E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 5.00E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 5.80E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 6.20E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 8.20E+4nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.29E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 1.45E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 2.90E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+5nMAssay Description:The aromatic compounds screening were characterized by molecular docking as well as by in vitro inhibition assay against carboxypeptidase(CPs) of dif...More data for this Ligand-Target Pair