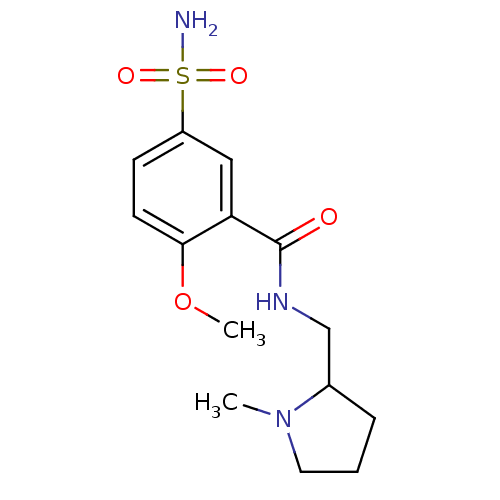
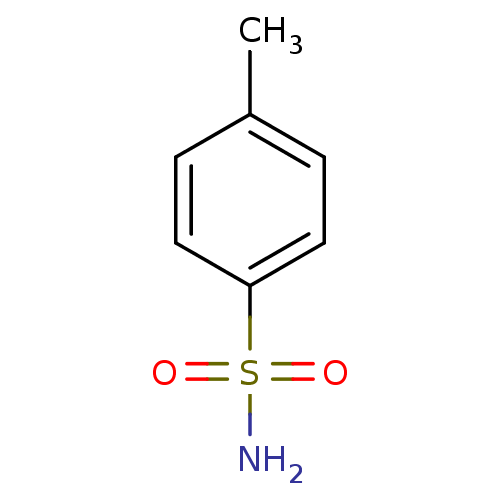
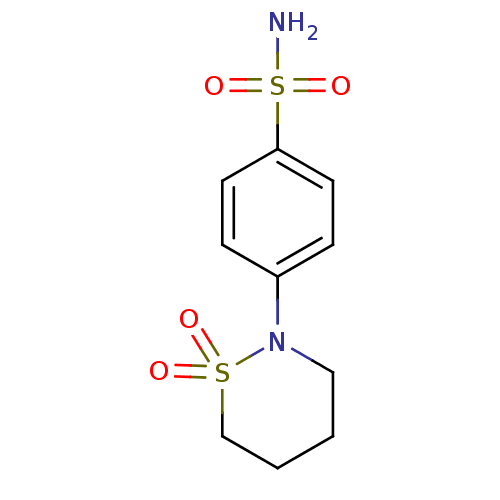
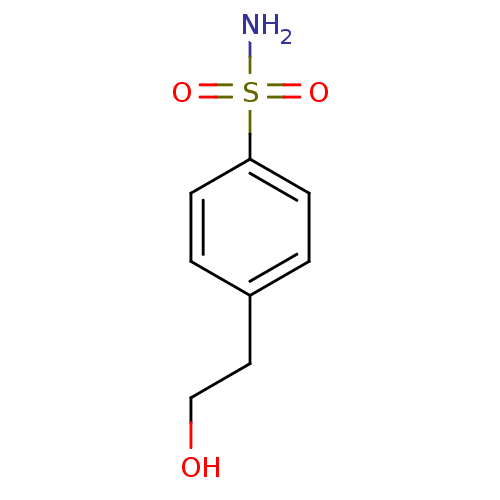
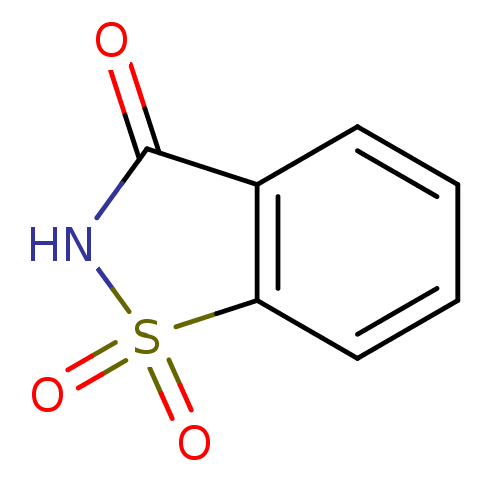
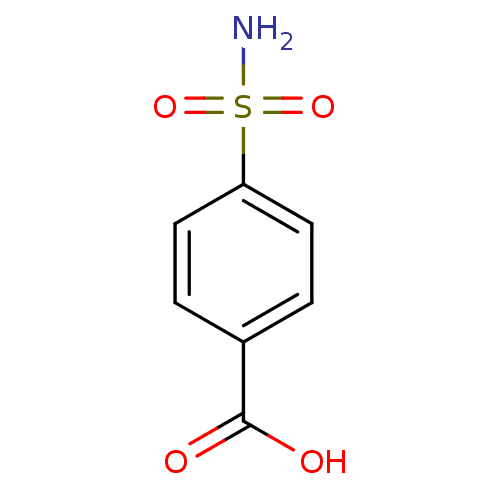
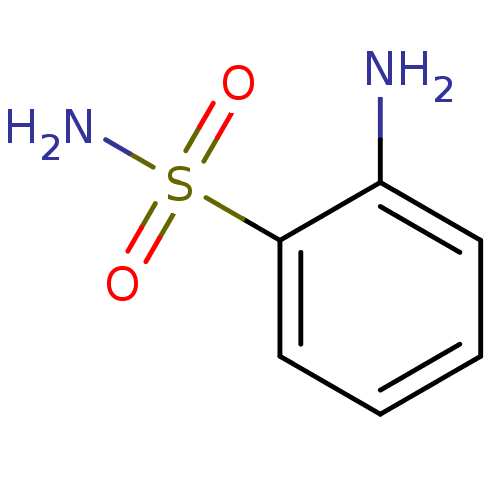
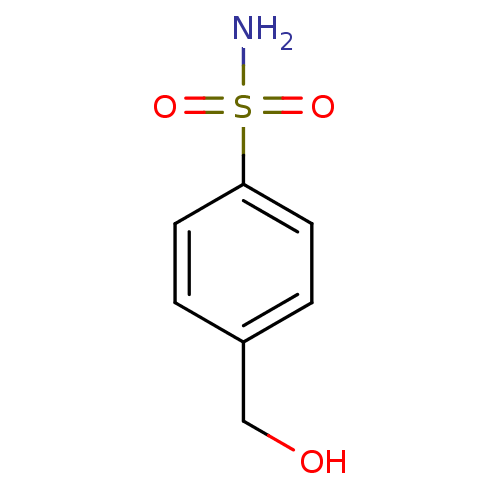
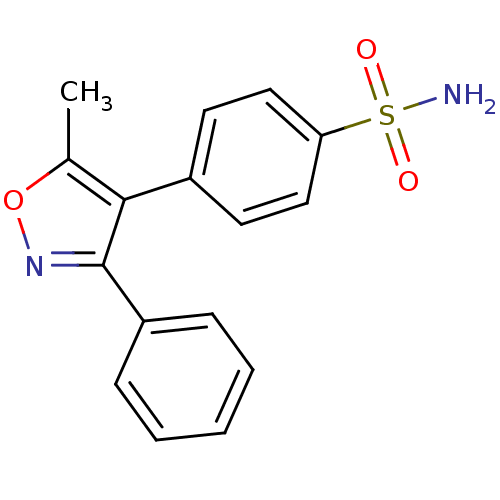
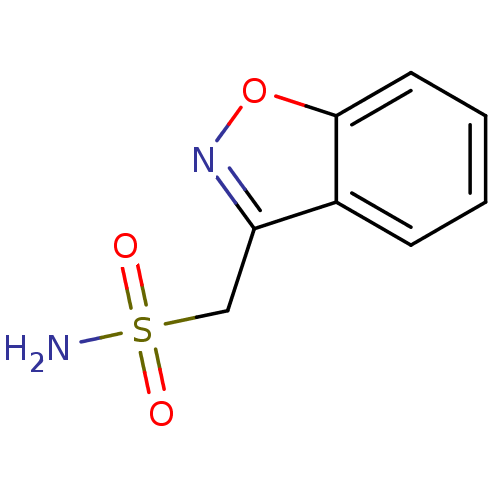
Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 3203
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
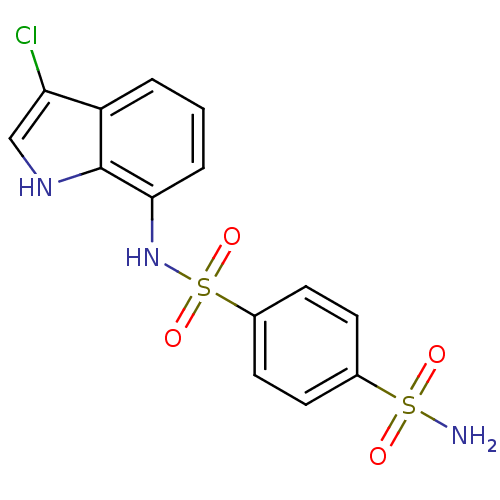
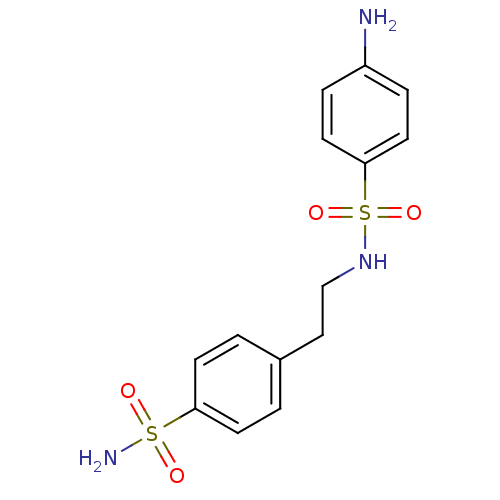
Affinity DataKi: 97nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
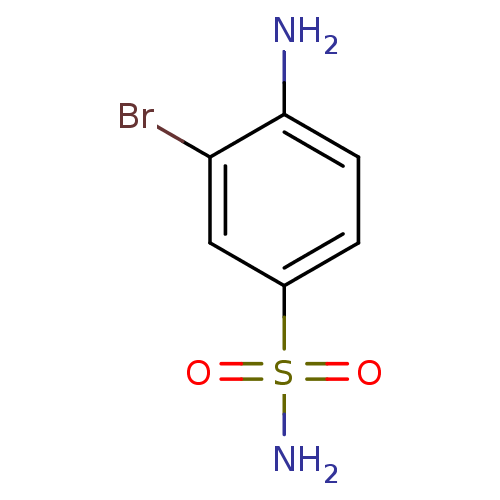
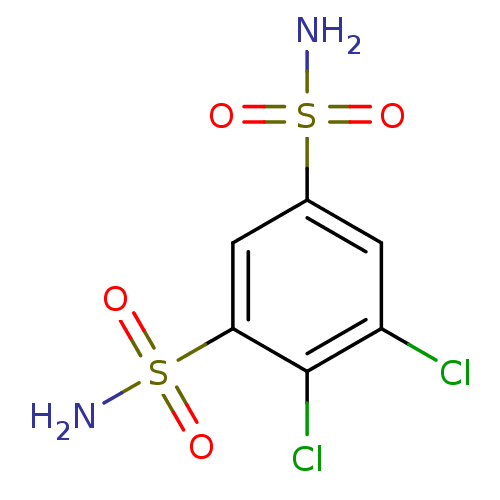
Affinity DataKi: 186nM ΔG°: -37.8kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
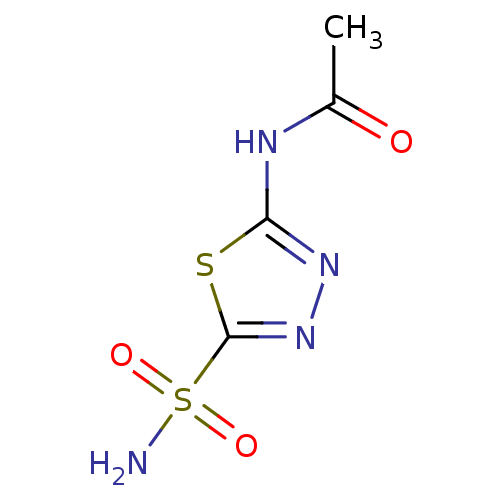
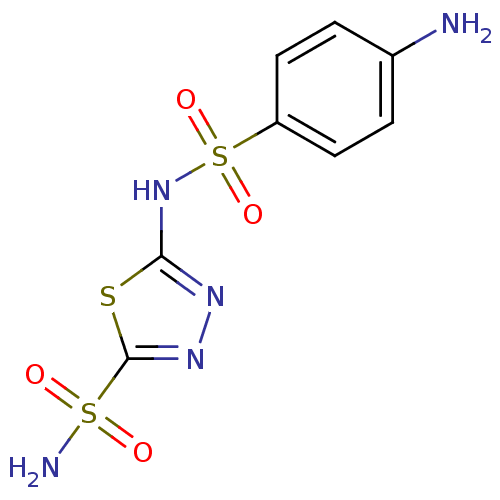
Affinity DataKi: 481nM ΔG°: -35.5kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
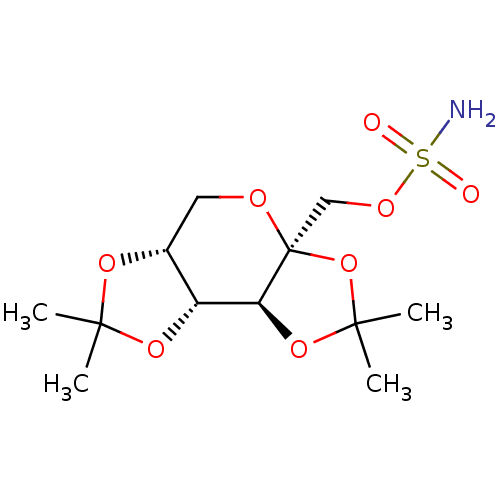
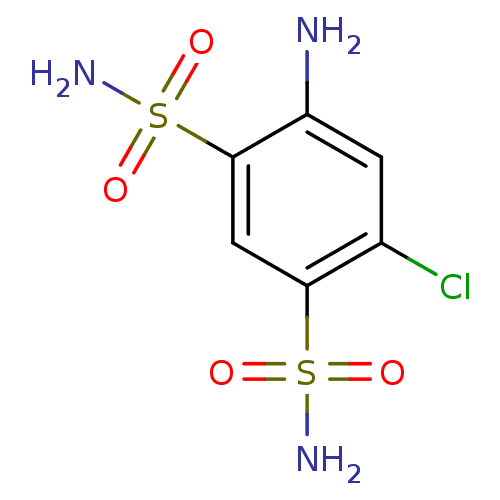
Affinity DataKi: 612nM ΔG°: -34.9kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 612nM ΔG°: -34.9kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 744nM ΔG°: -34.4kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 750nM ΔG°: -34.4kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 781nM ΔG°: -34.3kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 810nM ΔG°: -34.2kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 839nM ΔG°: -34.1kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 853nM ΔG°: -34.1kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 872nM ΔG°: -34.0kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 905nM ΔG°: -33.9kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.03E+3nM ΔG°: -33.6kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.72E+3nM ΔG°: -32.4kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 2.30E+3nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 4.92E+3nM ΔG°: -29.8kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 5.16E+3nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 5.51E+3nM ΔG°: -29.5kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 7.48E+3nM ΔG°: -28.8kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 7.52E+3nM ΔG°: -28.8kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 7.71E+3nM ΔG°: -28.7kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 7.93E+3nM ΔG°: -28.6kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 7.96E+3nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 8.10E+3nM ΔG°: -28.6kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 8.21E+3nM ΔG°: -28.5kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 8.69E+3nM ΔG°: -28.4kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 8.74E+3nM ΔG°: -28.4kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 9.23E+3nM ΔG°: -28.3kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 9.56E+3nM ΔG°: -28.2kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 9.56E+3nM ΔG°: -28.2kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 9.84E+3nM ΔG°: -28.1kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.04E+4nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.15E+4nM ΔG°: -27.7kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.27E+4nM ΔG°: -27.5kJ/molepH: 8.3 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 1.30E+4nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetBeta-carbonic anhydrase 1(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
Kochi Medical School
Kochi Medical School
Affinity DataKi: 2.87E+4nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair