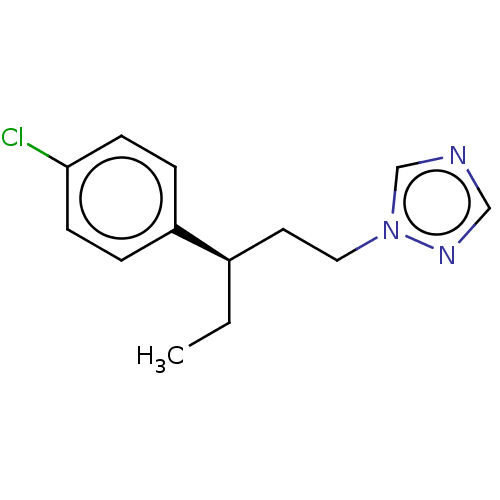
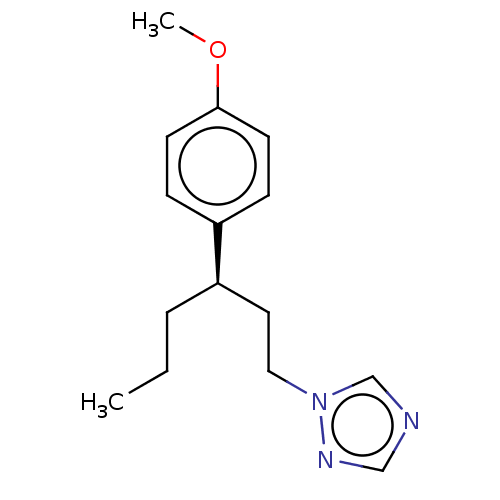
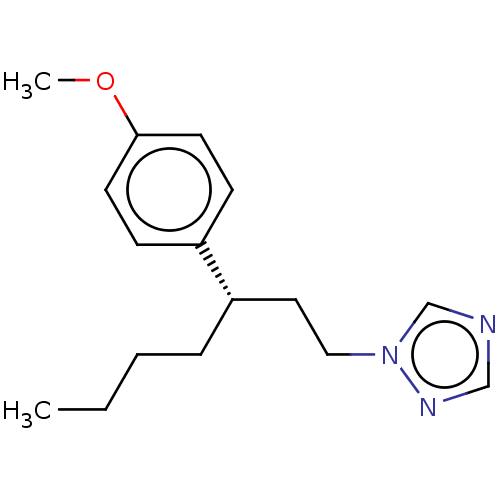
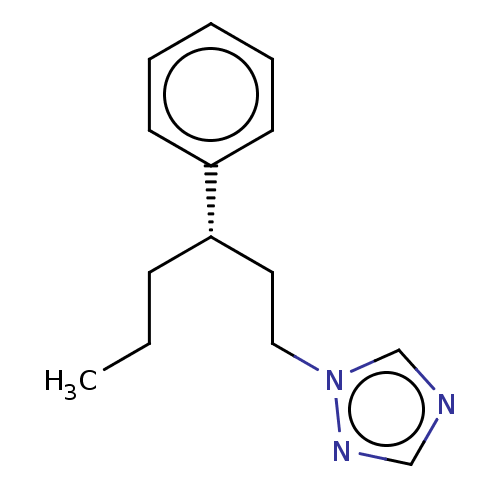
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 50004820
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 34nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 46nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 49nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 63nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 81nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 90nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 120nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 140nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 150nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 150nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 150nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 180nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 210nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 230nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 250nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 320nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 340nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 350nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 505nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 640nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 760nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 990nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 1.14E+3nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 1.22E+3nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair
TargetCytochrome P450 14alpha-demethylase(Green mold)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataKd: 1.66E+3nMAssay Description:Binding affinity to Penicillium digitatum CYP51 expressed in Escherichia coli BL21 (DE3) cells by binding spectrum methodMore data for this Ligand-Target Pair