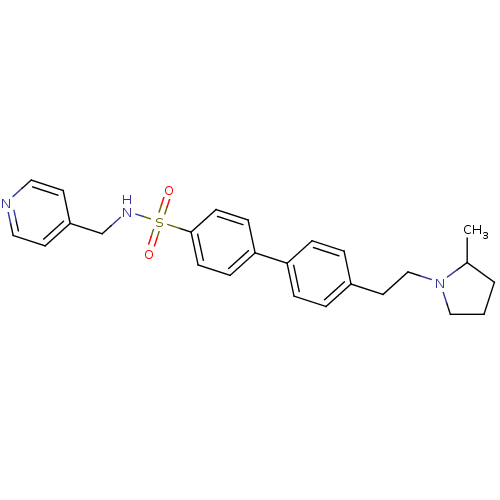
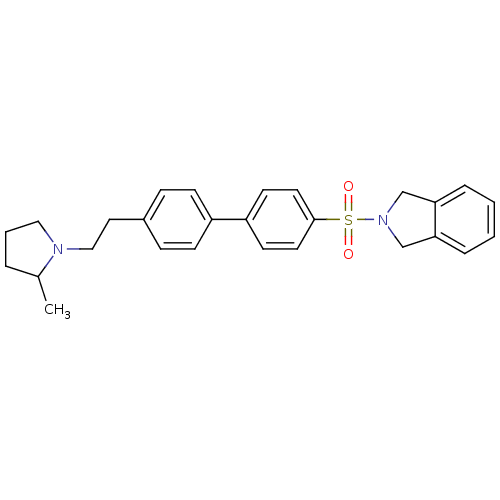
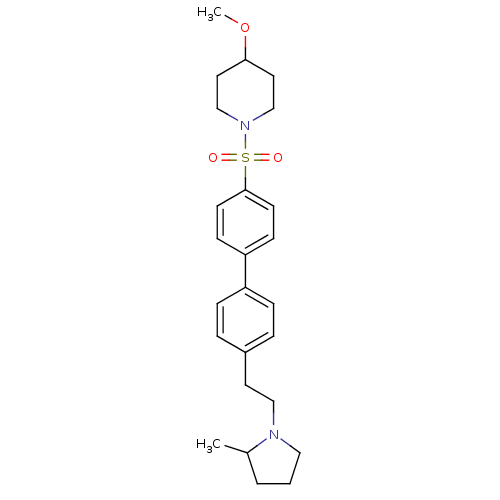
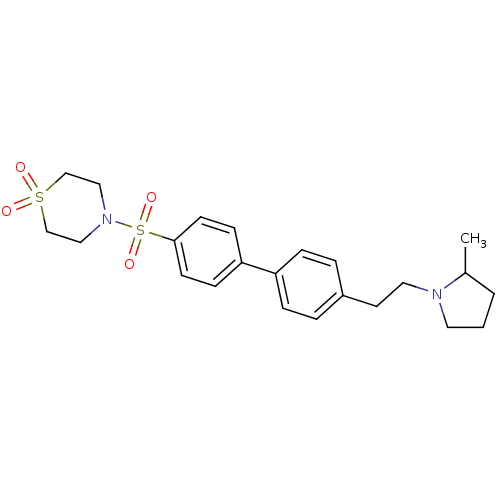
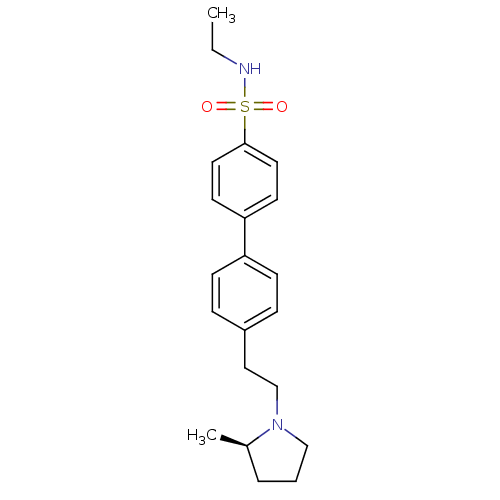
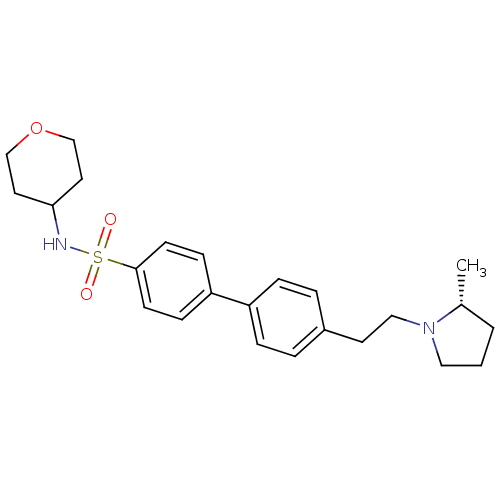
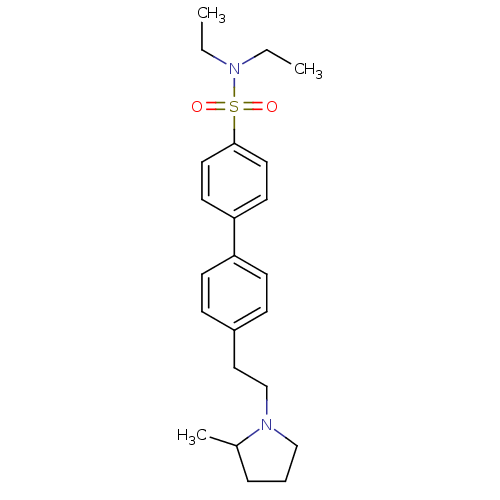
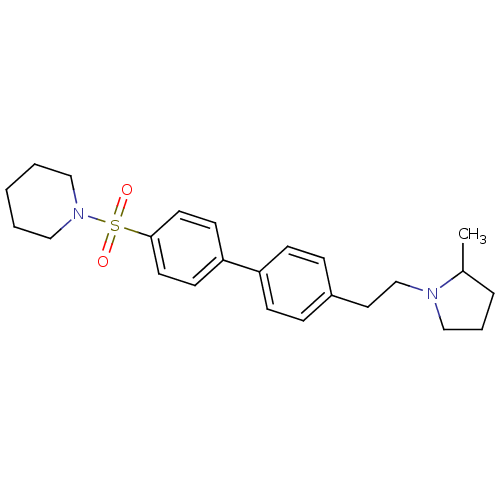
Report error Found 30 Enz. Inhib. hit(s) with all data for entry = 50030514
Affinity DataKi: 0.182nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.295nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.501nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.646nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.933nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inverse agonist activity against human histamine H3 receptor by [35S]GTPgamma binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inverse agonist activity against human histamine H3 receptor by [35S]GTPgamma binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inverse agonist activity against human histamine H3 receptor by [35S]GTPgamma binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.41nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.58nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.62nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.78nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.86nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 2.29nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 2.45nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 2.51nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.27nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.57nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.57nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 5.13nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.92nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.08nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 7.94nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 11.0nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 12.6nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 16.2nMAssay Description:Displacement of N-[3H]methylhistamine from histamine H3 receptor in rat cortex membraneMore data for this Ligand-Target Pair