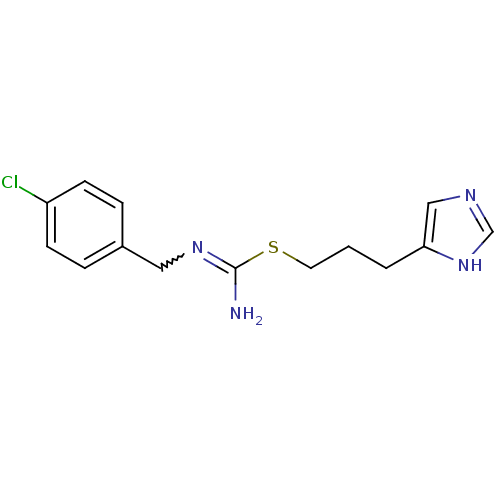
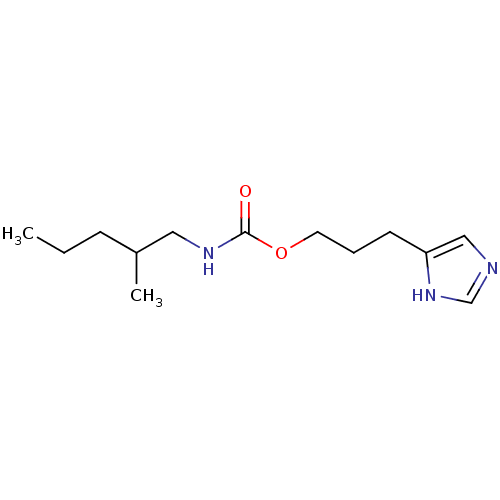
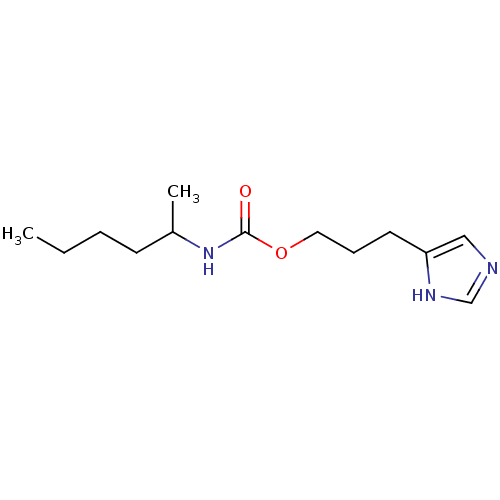
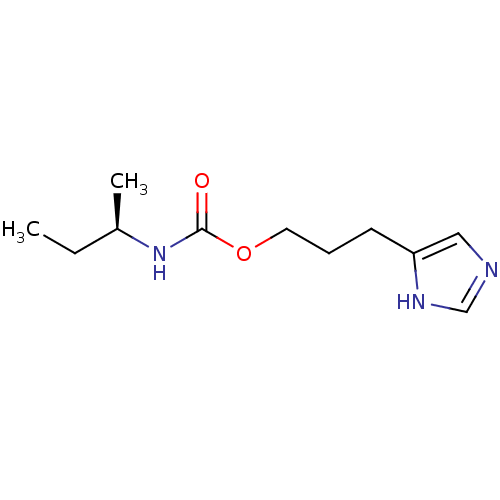
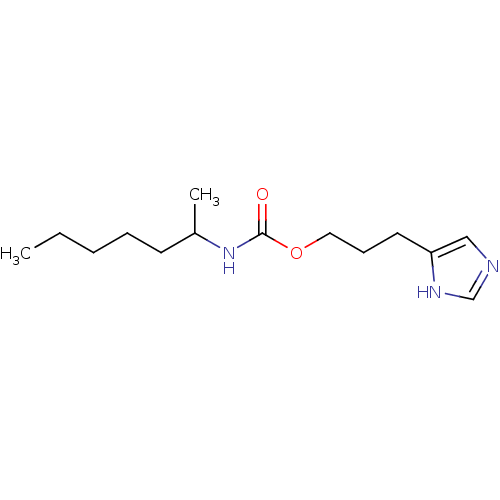
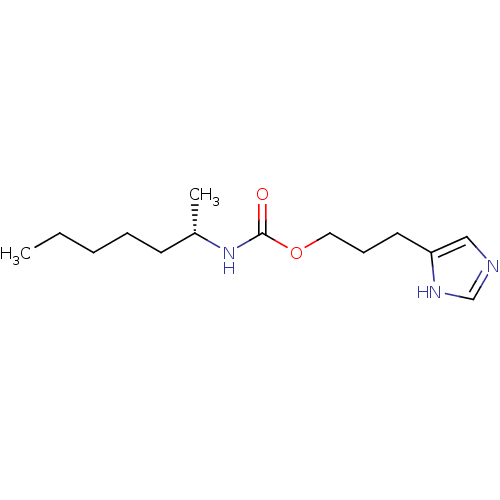
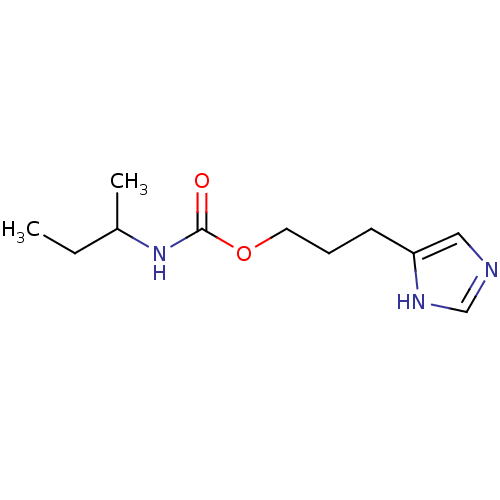
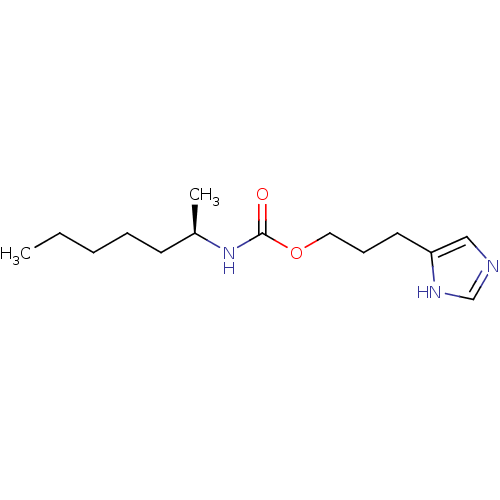
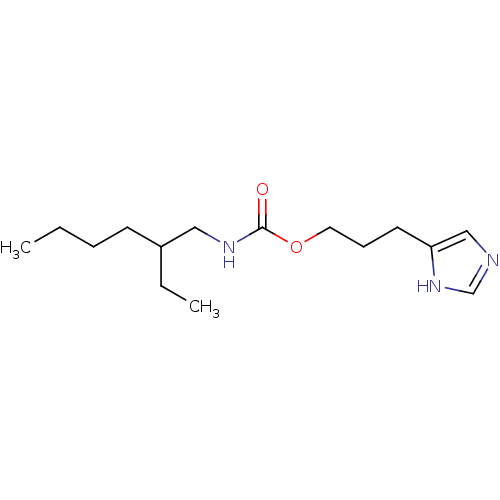
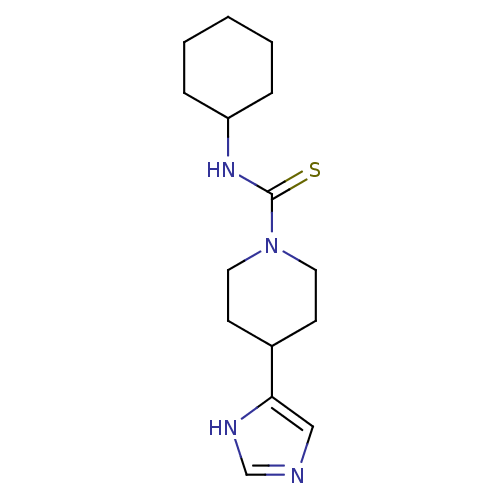
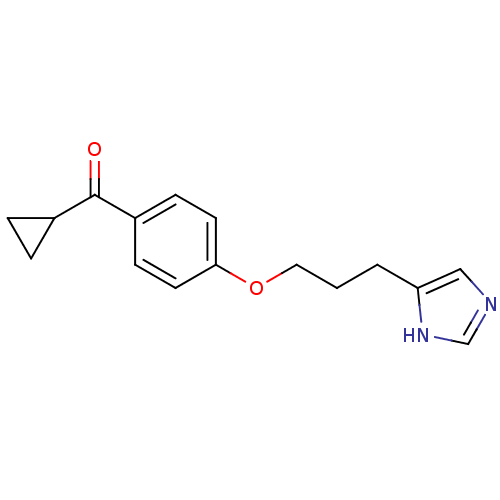
Report error Found 52 Enz. Inhib. hit(s) with all data for entry = 50031352
Affinity DataKi: 2.40nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.70nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.10nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [3H]histamine from histamine H3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 46nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 49nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 74nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 75nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 91nMAssay Description:Displacement of [3H]iodoproxyfan from human histamine H3 receptor expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 118nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 123nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 162nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 426nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 612nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 636nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair
Affinity DataKi: 695nMAssay Description:Displacement of [3H]histamine from human histamine H4 receptor expressed in SF9 cells coexpressing Galphai2, Gbeta1gamma2More data for this Ligand-Target Pair