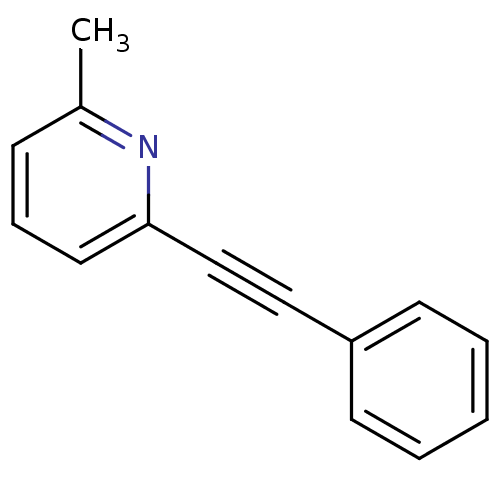
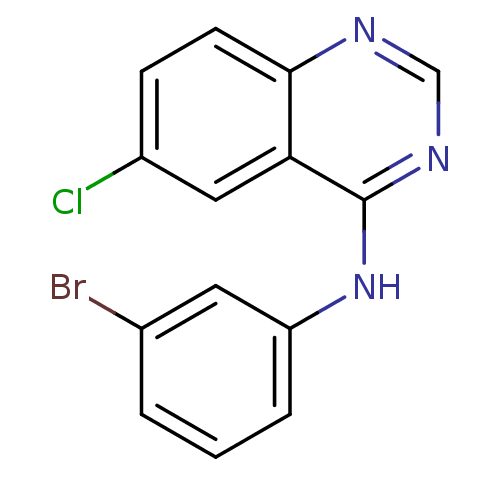
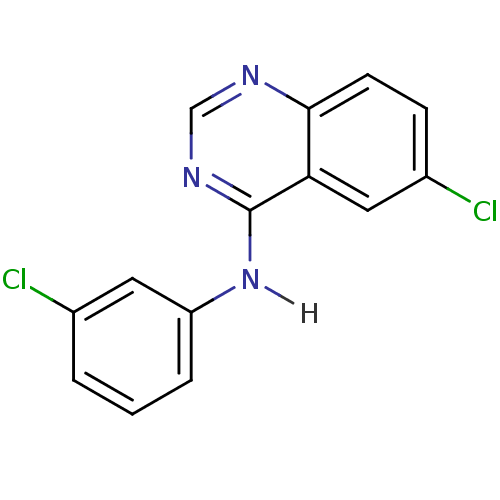
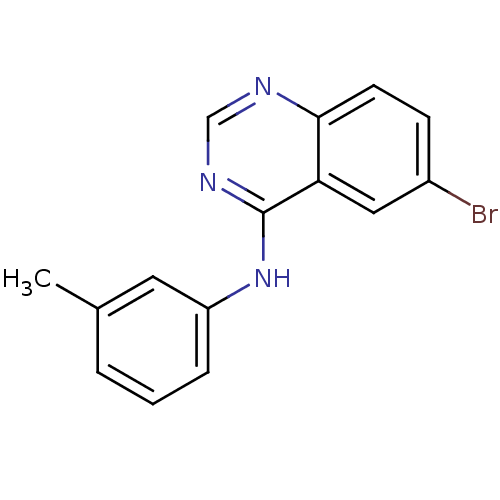
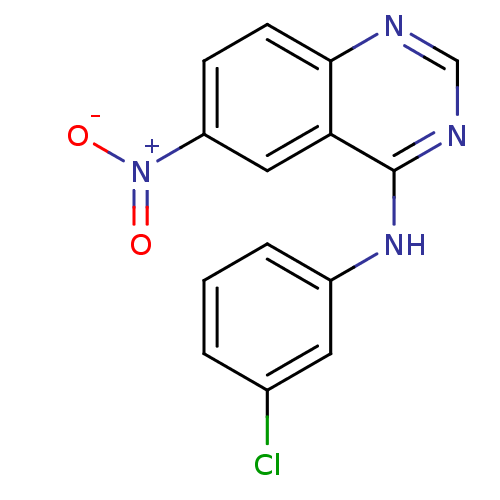
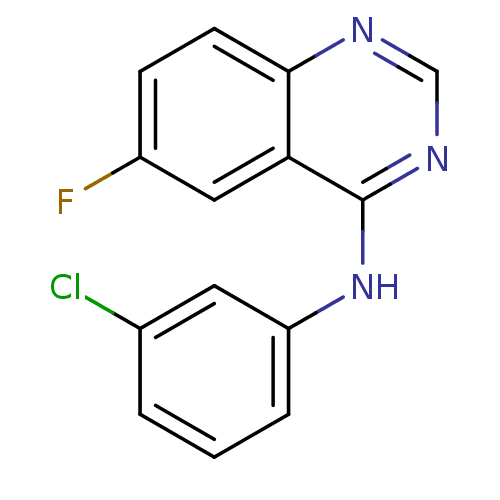
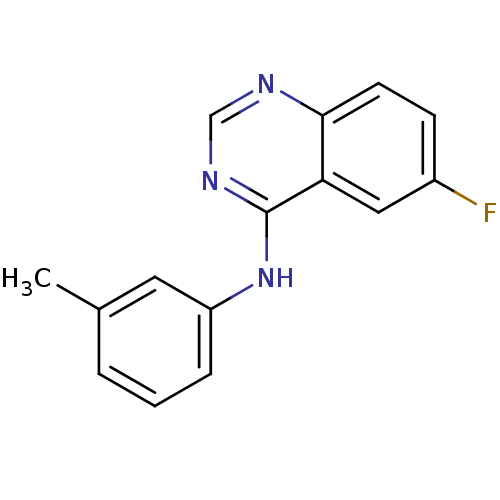
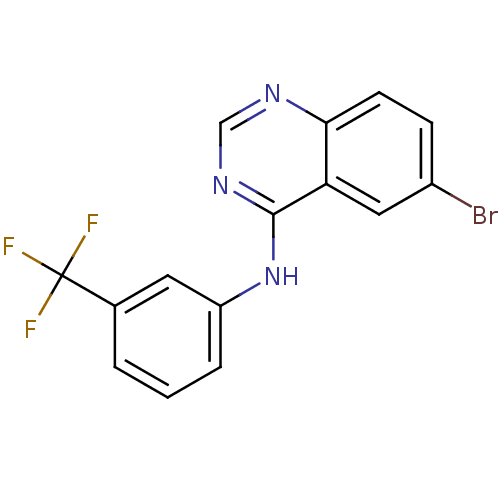
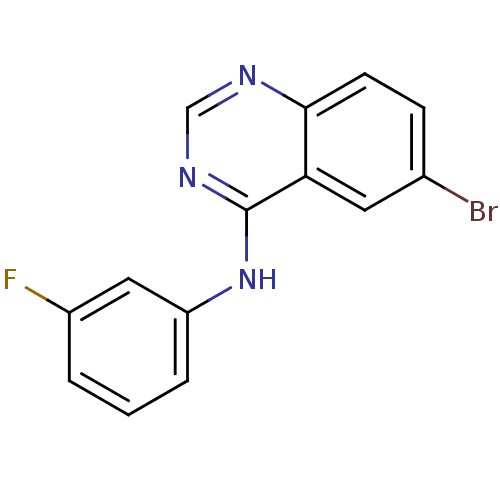
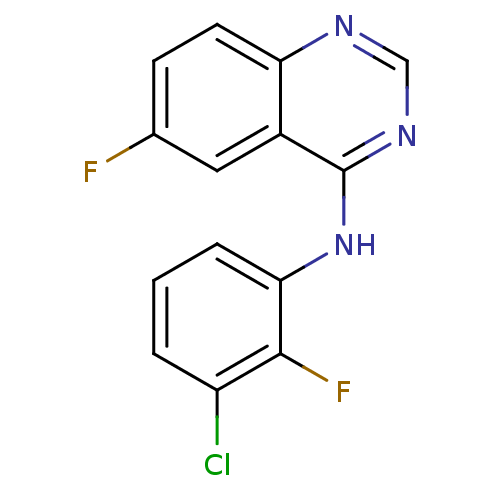
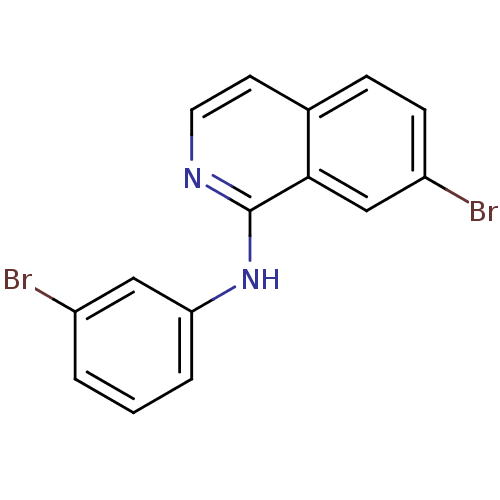
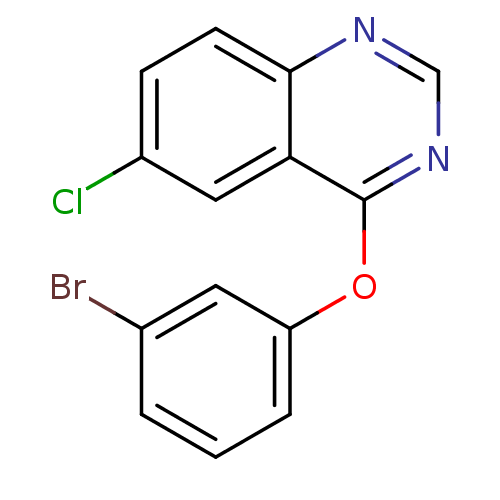
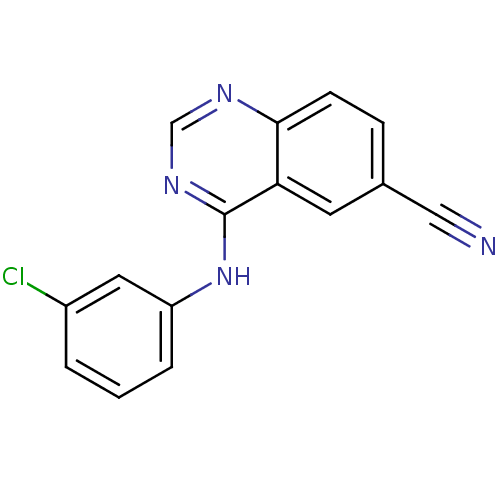
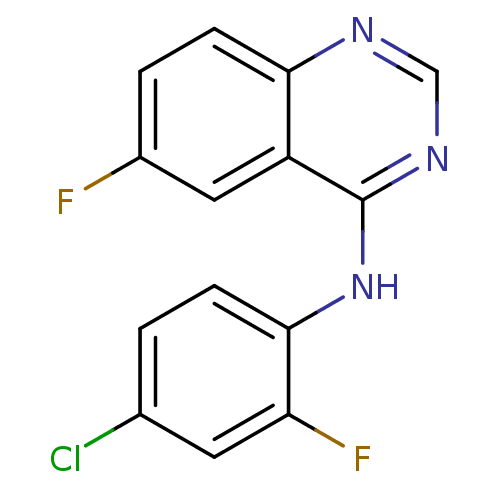
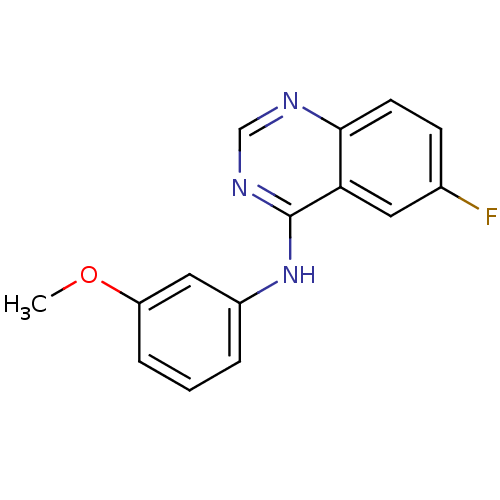
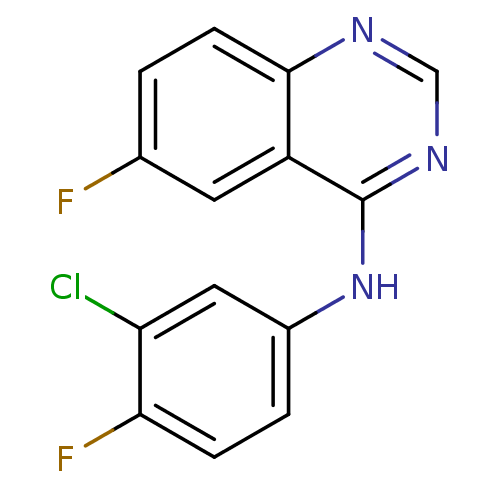
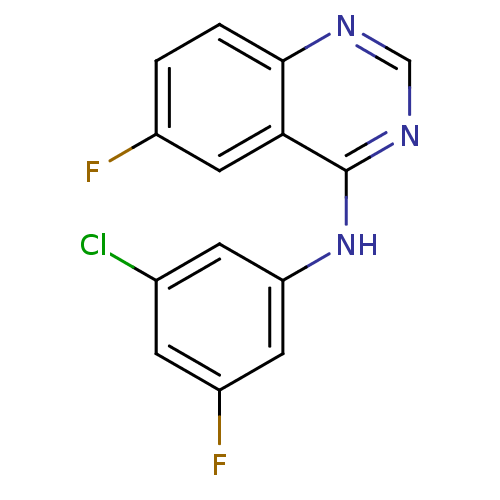
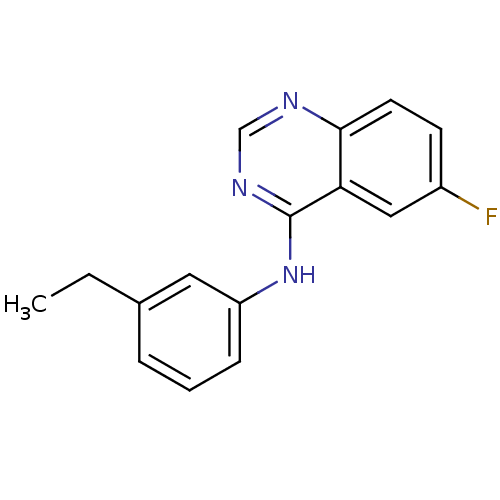
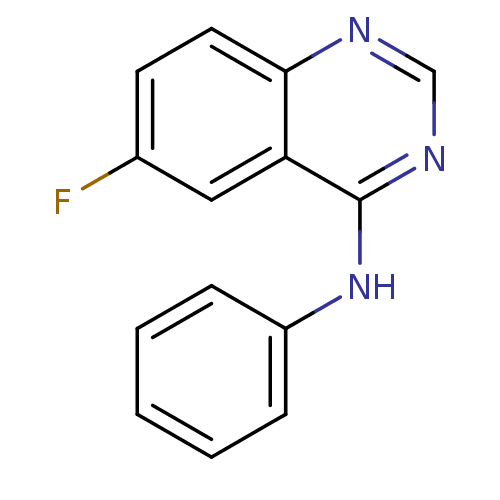
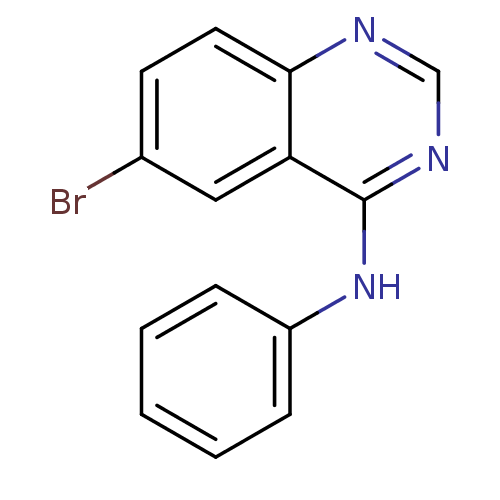
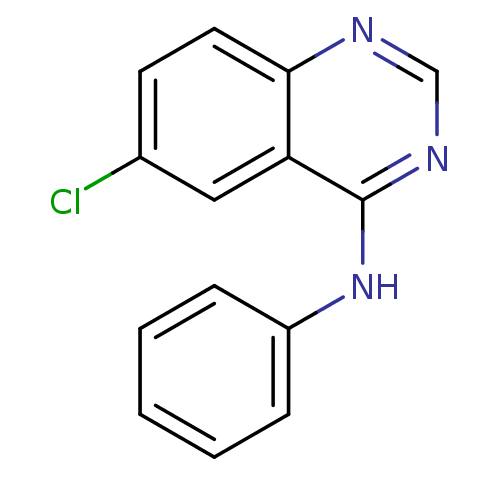
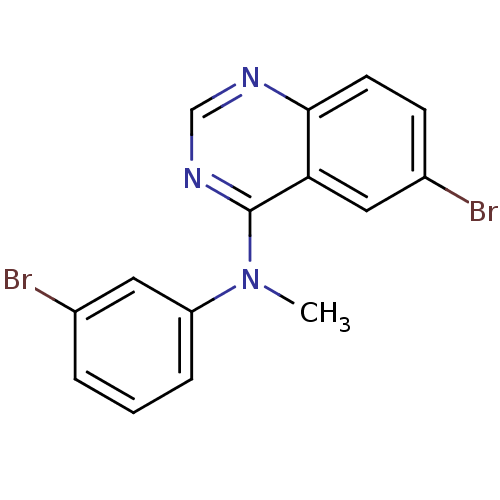
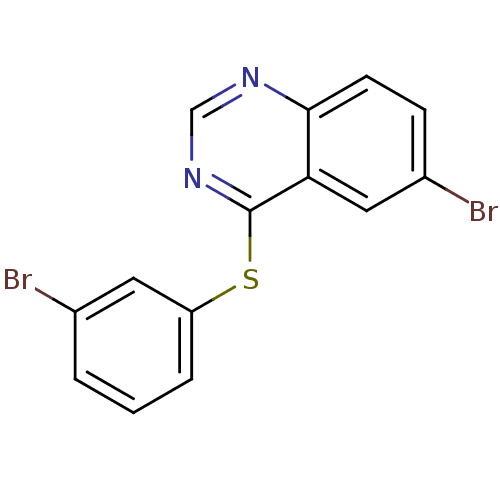
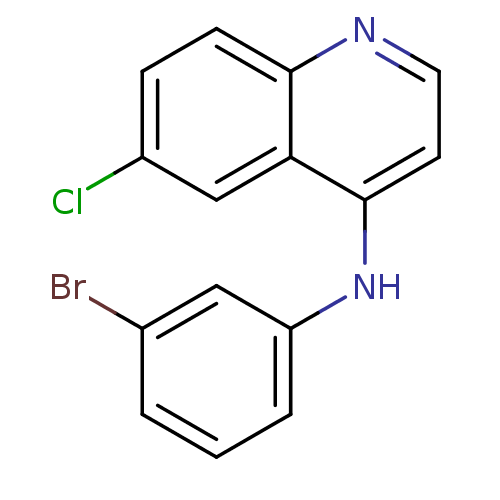
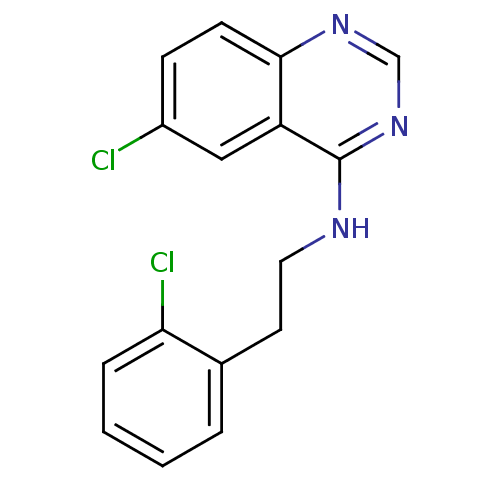
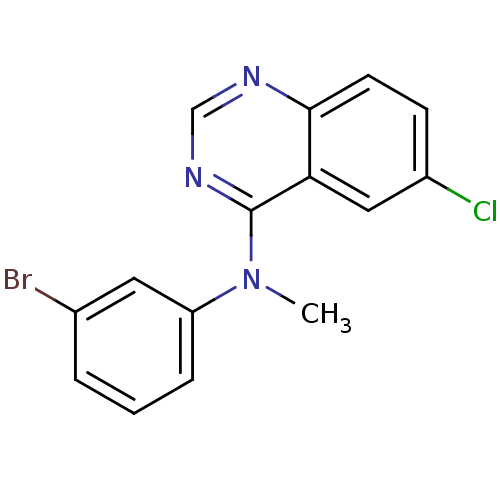
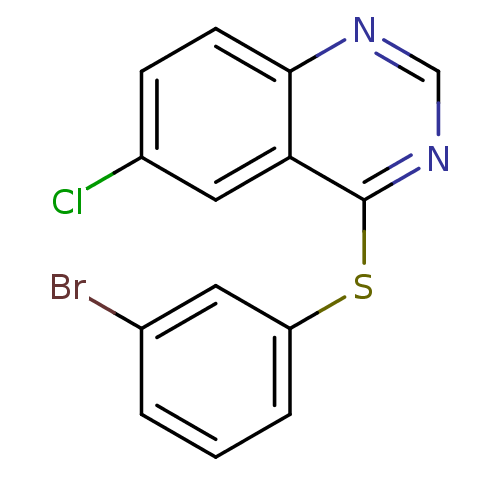
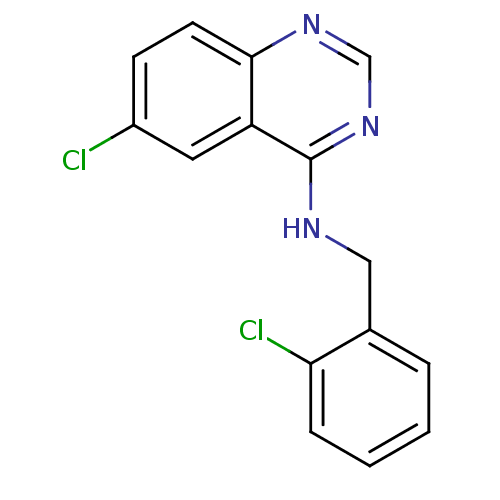
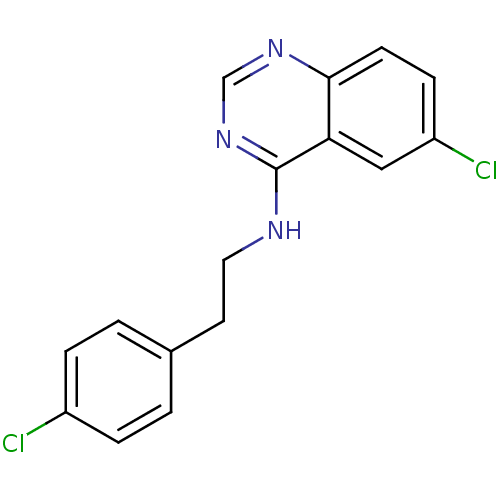
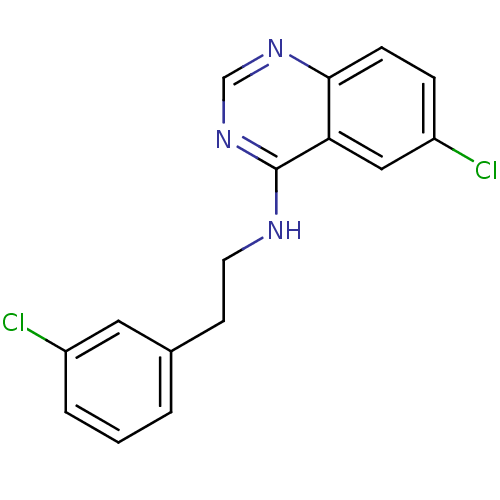
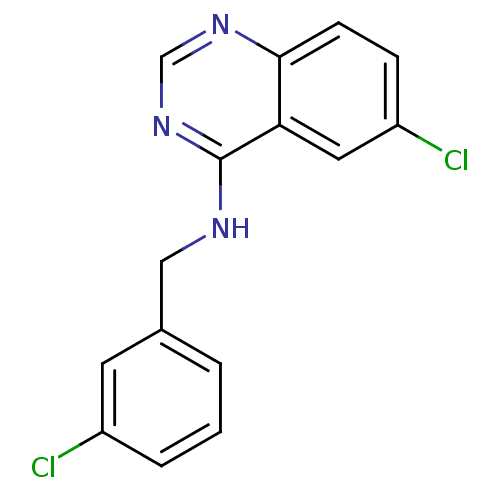
Report error Found 49 Enz. Inhib. hit(s) with all data for entry = 50031358
Affinity DataKi: 4.70nMAssay Description:Displacement of [3H]3methoxy-5-(pyridin-2-ylethynyl)pyridine from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 96nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataKi: 116nMAssay Description:Displacement of [3H]3methoxy-5-(pyridin-2-ylethynyl)pyridine from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 124nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 174nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 246nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataKi: 249nMAssay Description:Displacement of [3H]3methoxy-5-(pyridin-2-ylethynyl)pyridine from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 256nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 274nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 311nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 398nMAssay Description:Antagonist activity at rat mGluR1 expressed in BHK cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 713nMAssay Description:Antagonist activity at rat mGluR1 expressed in BHK cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataKi: 736nMAssay Description:Displacement of [3H]3methoxy-5-(pyridin-2-ylethynyl)pyridine from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 955nMAssay Description:Antagonist activity at rat mGluR1 expressed in BHK cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.35E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.37E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.72E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.97E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of EGFR autophosphorylation in human A431 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.33E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.57E+3nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulator activity at rat mGluR5 receptor expressed in HEK293 cells assessed as potentiation of glutamate-induced responseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Antagonist activity at rat mGluR5 expressed in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair