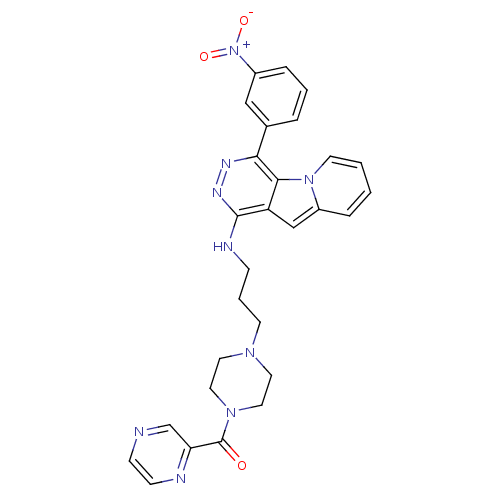
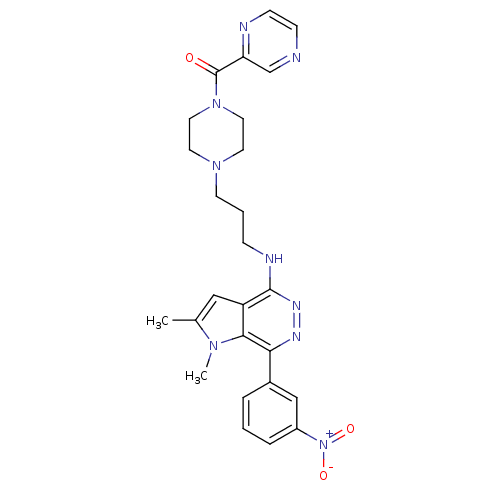
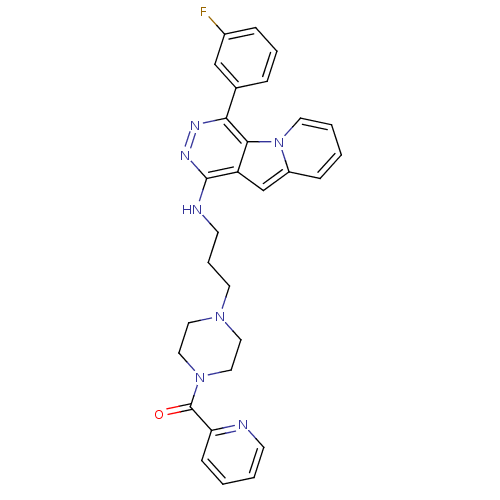
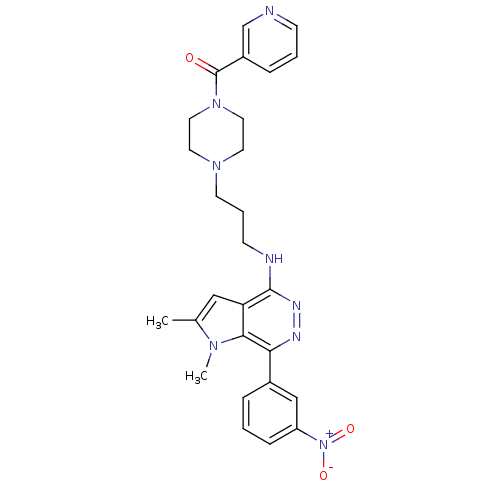
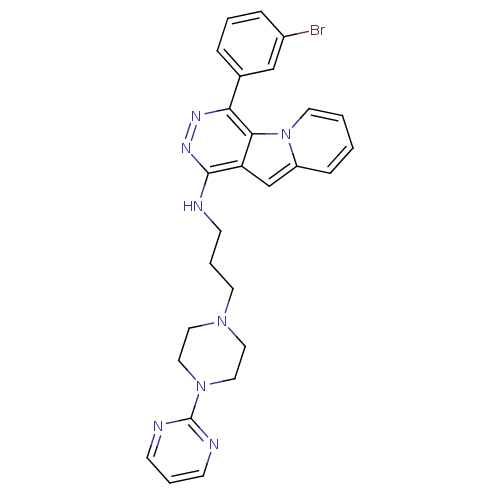
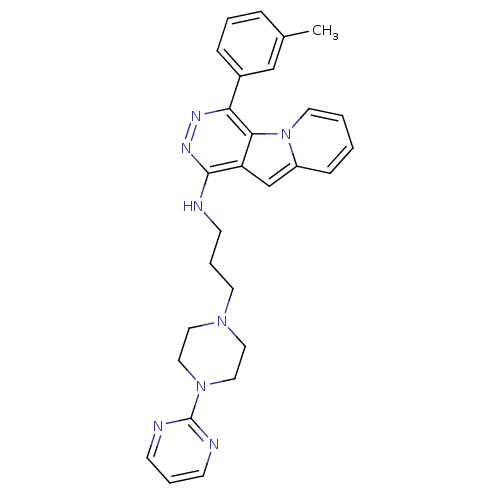
Report error Found 62 Enz. Inhib. hit(s) with all data for entry = 50031512
Affinity DataKi: 5.60nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 147nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 210nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 222nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 238nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 241nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 389nMAssay Description:Inhibition of PDE4B3 expressed in CHO cells assessed as isoproterenol-induced [125I]cAMP accumulation after 15 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 413nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 432nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 977nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.03E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.44E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.01E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.35E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.40E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.44E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.44E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.90E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of PDE4D4 expressed in CHO cells assessed as isoproterenol-induced [125I]cAMP accumulation after 15 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.08E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.20E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.50E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+3nMAssay Description:Inhibition of PDE4B3 expressed in CHO cells assessed as isoproterenol-induced [125I]cAMP accumulation after 15 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.37E+3nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.15E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.27E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMAssay Description:Inhibition of PDE4B3 expressed in CHO cells assessed as isoproterenol-induced [125I]cAMP accumulation after 15 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.42E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.53E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.75E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.90E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.22E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.62E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.86E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.12E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.22E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4B3 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.44E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.90E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 6.10E+4nMAssay Description:Displacement of [methyl-3H]rolipram from PDE4D4 expressed in CHO cells after 1 hr by scintillation countingMore data for this Ligand-Target Pair