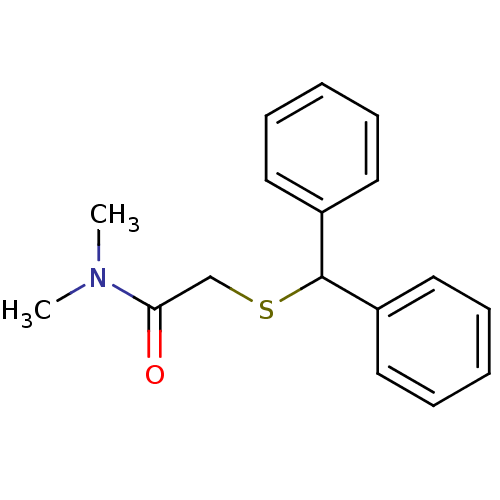
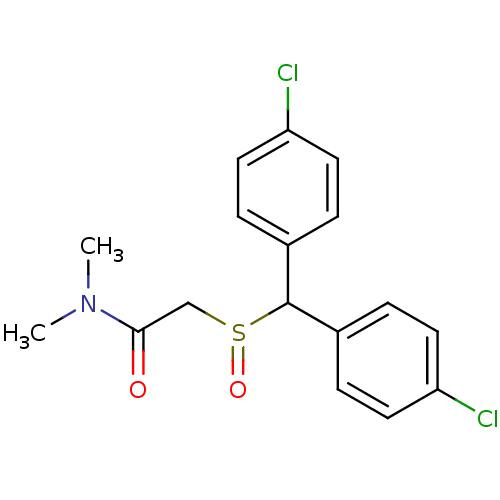
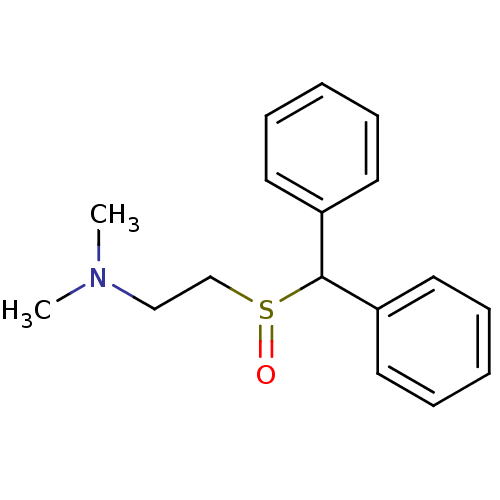
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50039387
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 72nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 100nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 194nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 286nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 406nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 600nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 919nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.57E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.65E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.93E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.19E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.20E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.23E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.35E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.44E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.45E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.52E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.66E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.89E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 3.21E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 3.26E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3.30E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 4.51E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 5.98E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 7.64E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 9.51E+3nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.06E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.27E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.42E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.62E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 1.65E+4nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.92E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.13E+4nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.23E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 2.59E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 3.32E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 3.46E+4nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3.62E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 3.90E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 4.25E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rat)
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
National Institute On Drug Abuse-Intramural Research Program
Curated by ChEMBL
Affinity DataKi: 4.58E+4nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5.21E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.77E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair