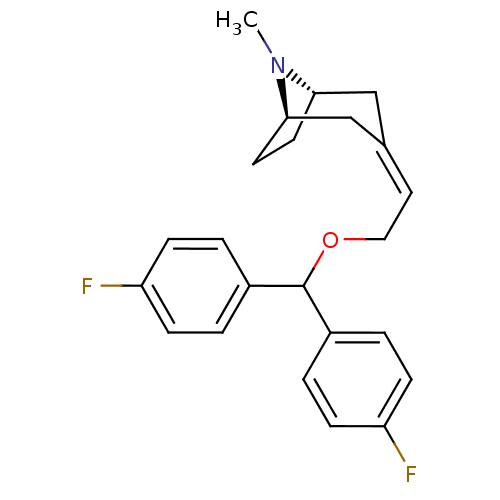
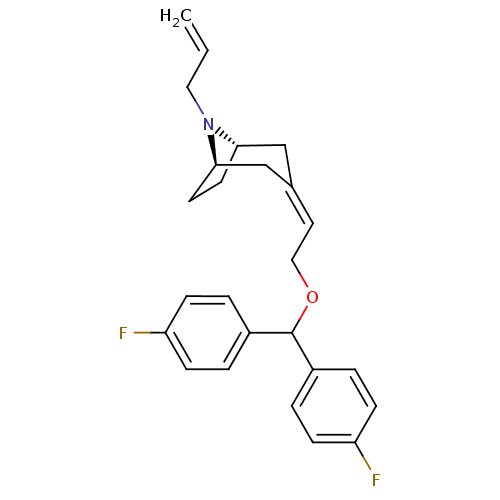
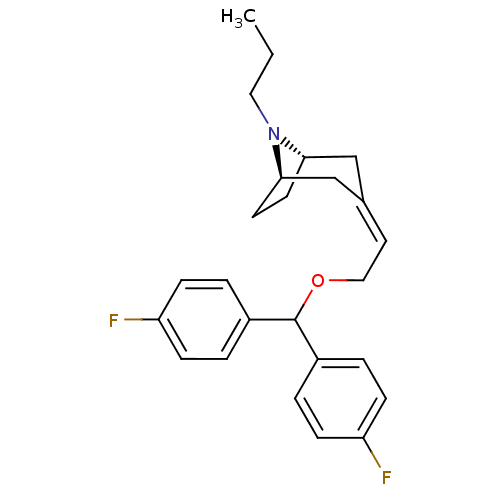
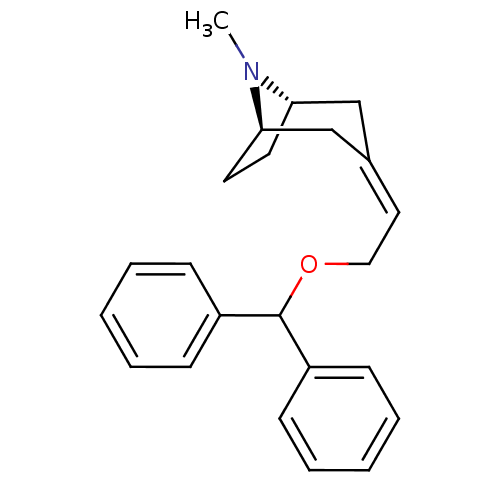
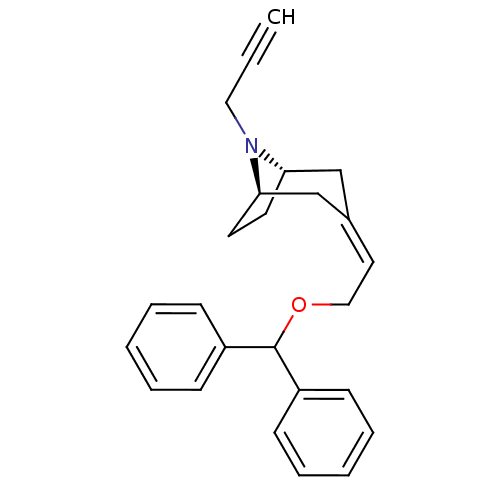
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 50034254
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 10.6nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 132nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 143nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 169nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 245nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 286nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 405nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 496nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 536nMAssay Description:Displacement of [3H]WIN35428 from DAT in Sprague-Dawley rat brain homogenates after 1 hr by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 544nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 656nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 746nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.15E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.39E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 1.77E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 2.22E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 2.71E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 2.75E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 2.85E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 3.12E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 3.15E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 3.34E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 3.46E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.58E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 4.09E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.24E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.30E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in Sprague-Dawley rat brain cortex membranes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 5.42E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.46E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.96E+3nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain homogenates after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
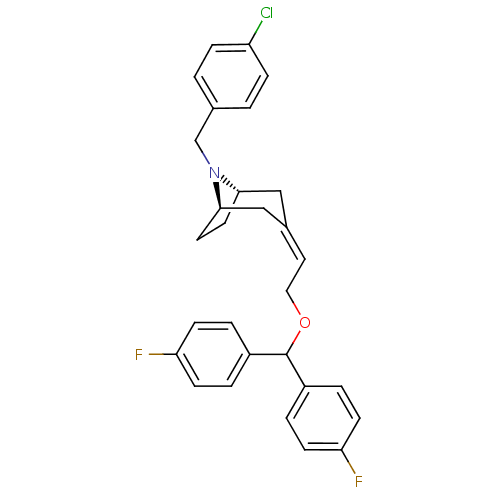
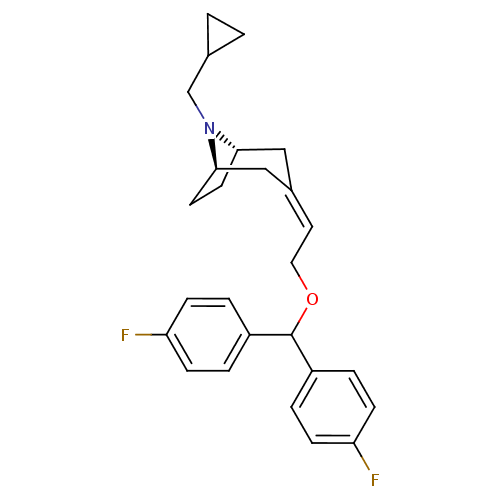

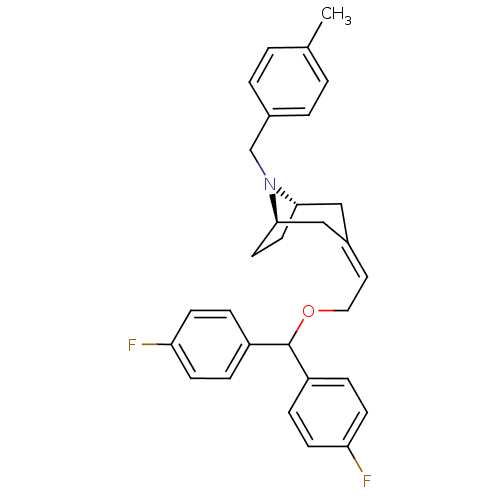


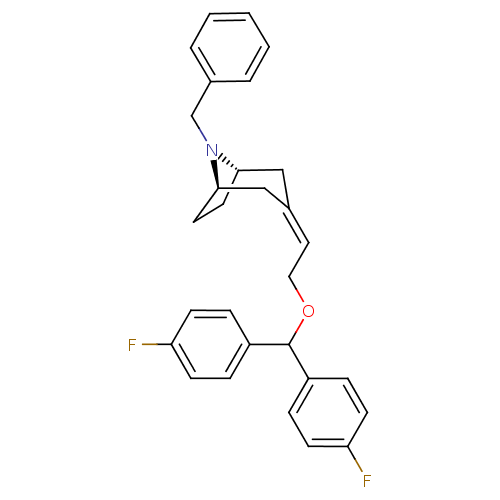
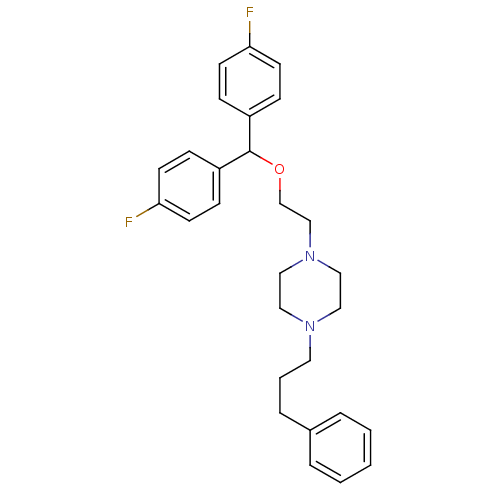
 3D Structure (crystal)
3D Structure (crystal)