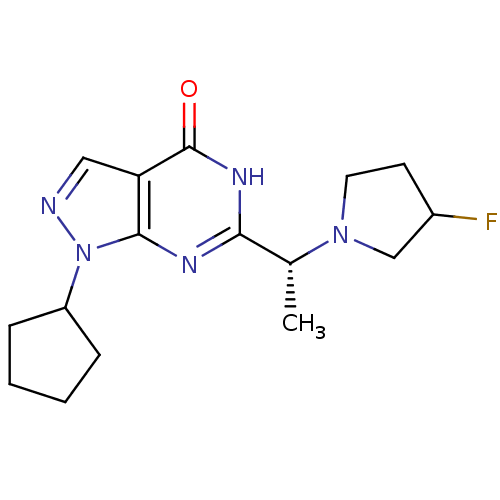
Report error Found 32 Enz. Inhib. hit(s) with all data for entry = 50040715
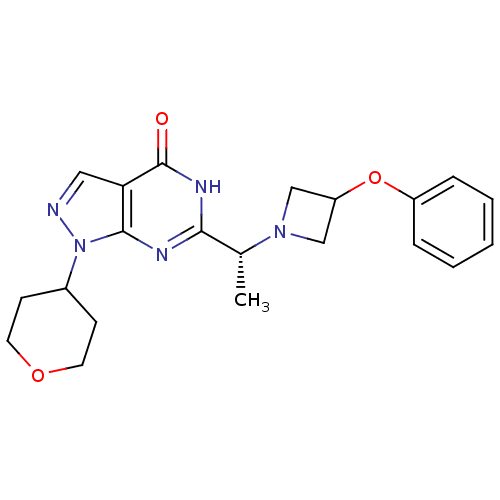
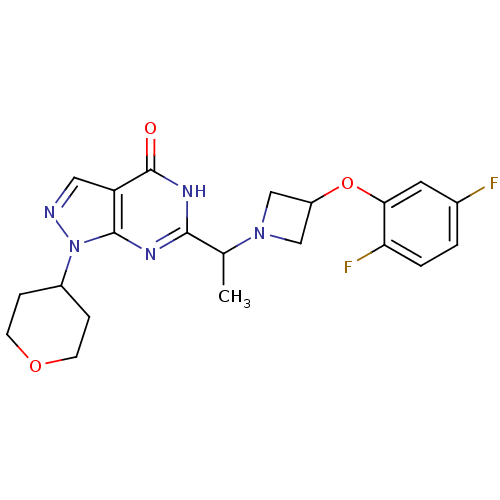
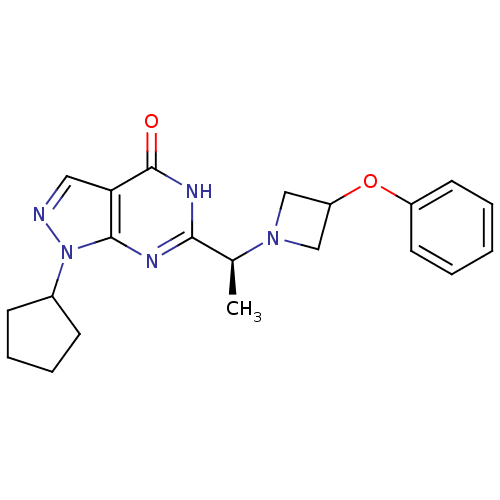
Affinity DataIC50: 7nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
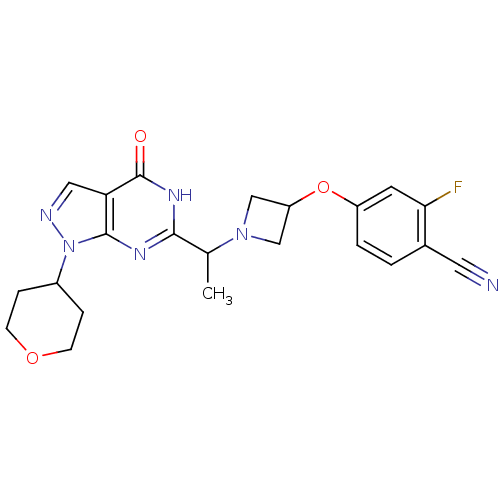
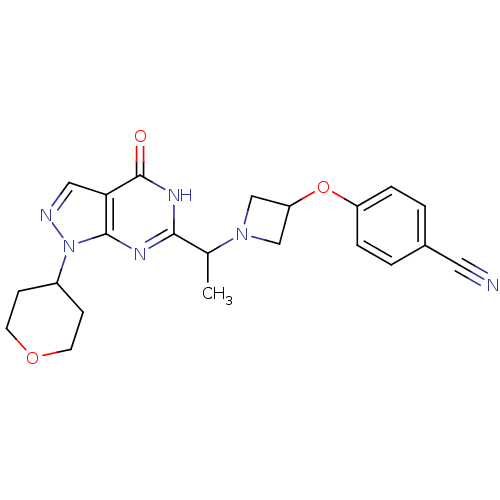
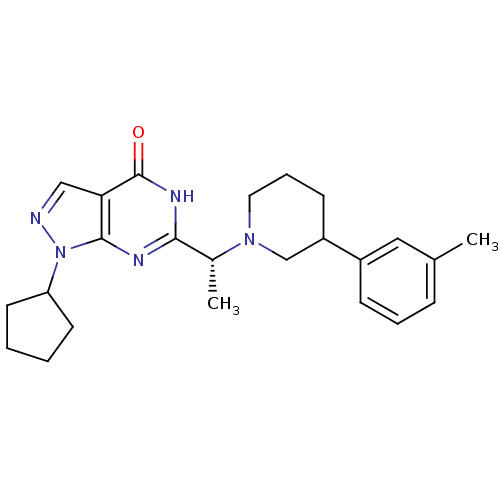
Affinity DataIC50: 11nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
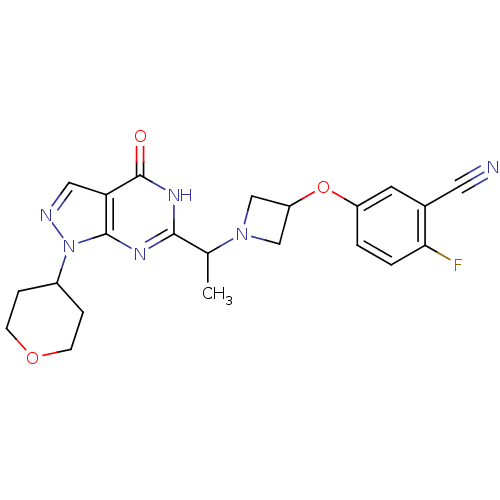
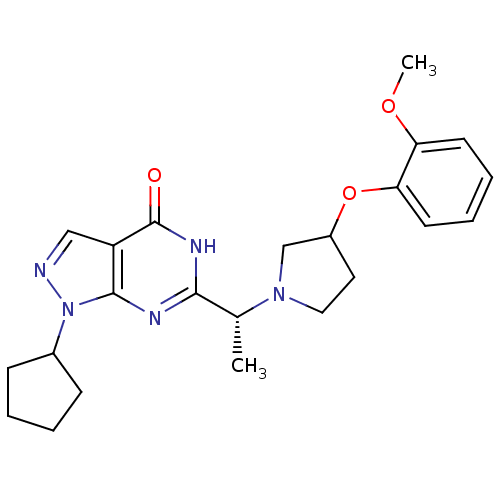
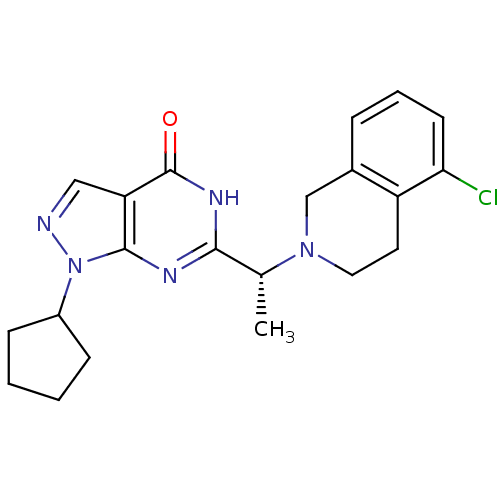
Affinity DataIC50: 19nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
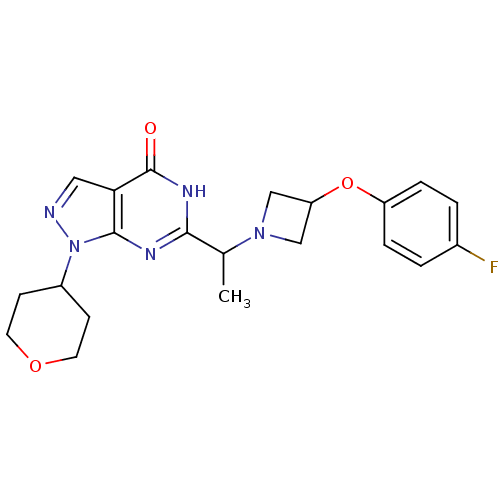
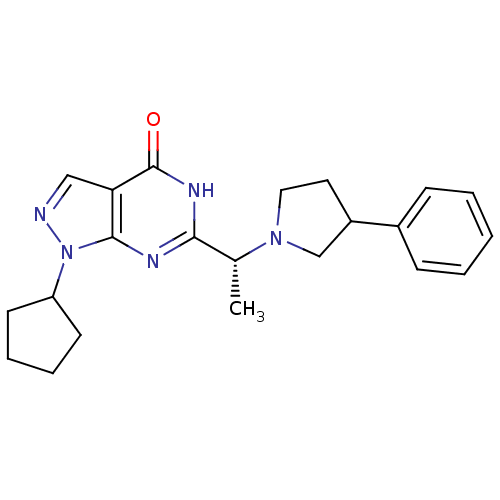
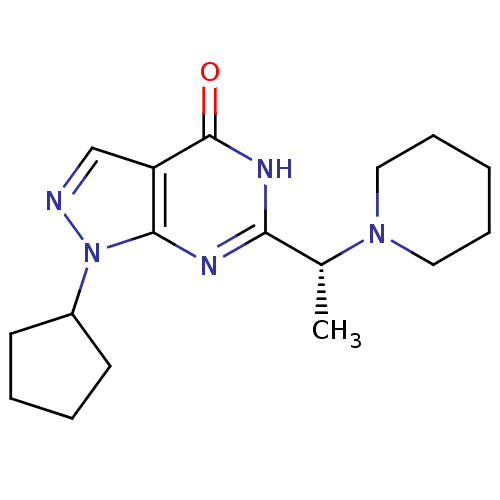
Affinity DataIC50: 22nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataIC50: 148nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 293nMAssay Description:Inhibition of human DATMore data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataIC50: 593nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataIC50: 986nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE1CMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE9A using [3H]cGMP as substrate after 30 mins by scintillation proximity assayMore data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C(Human)
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE1CMore data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of human DATMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Inhibition of human DATMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Inhibition of human DATMore data for this Ligand-Target Pair
Affinity DataIC50: 5.35E+4nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
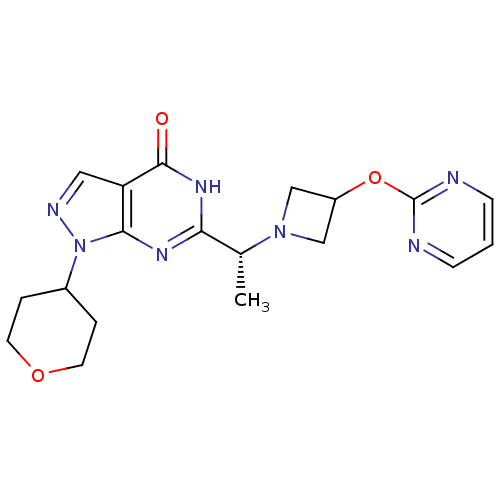
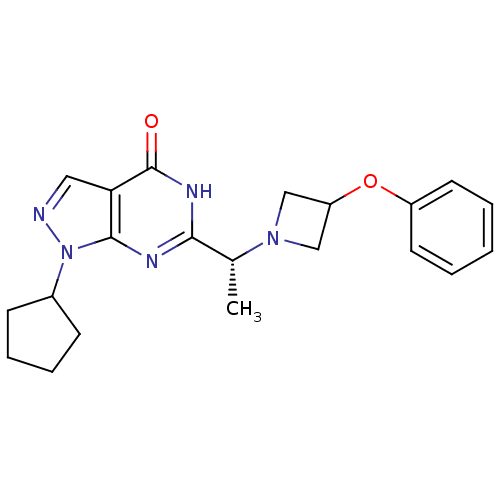
 3D Structure (crystal)
3D Structure (crystal)