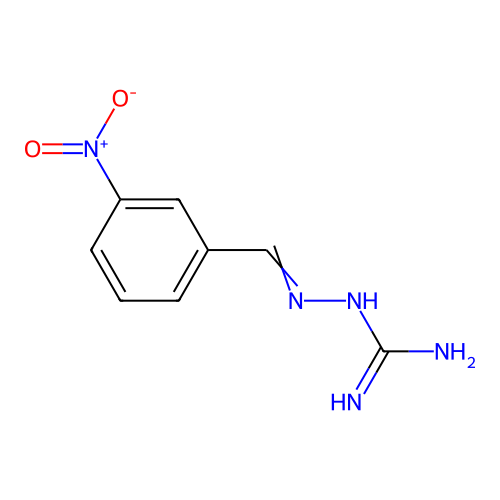
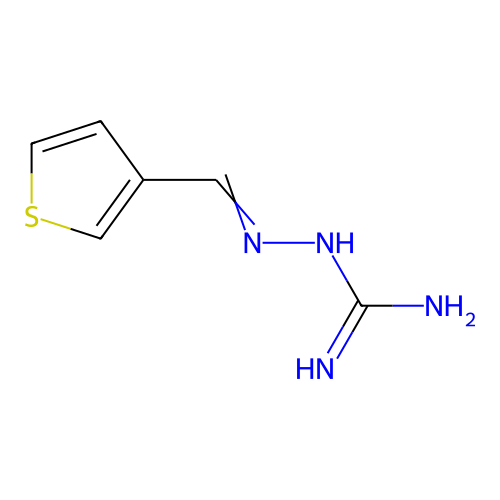
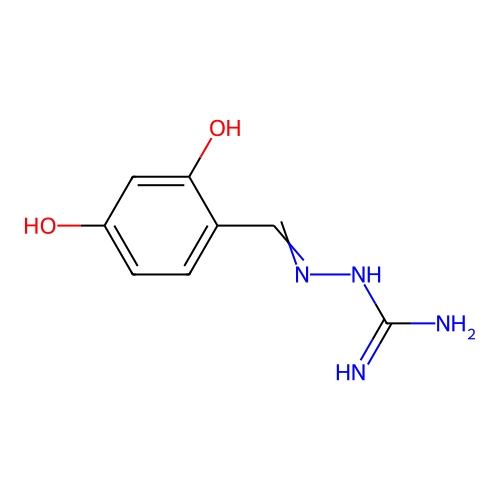
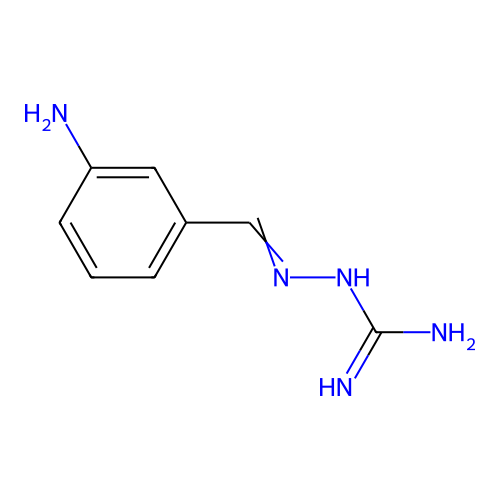
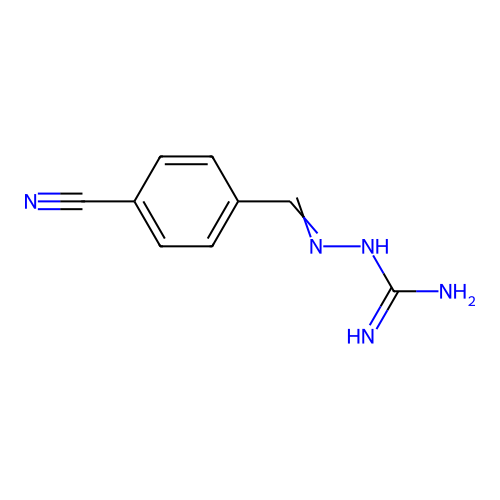
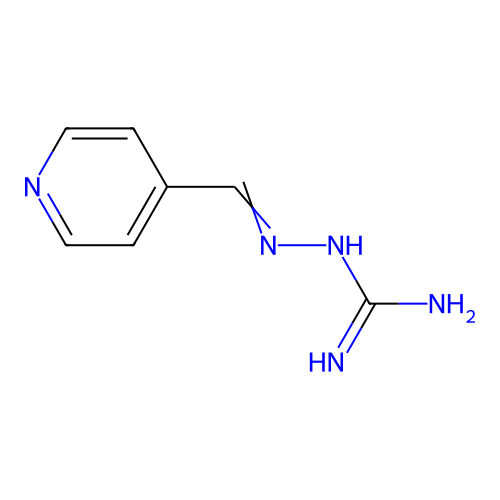
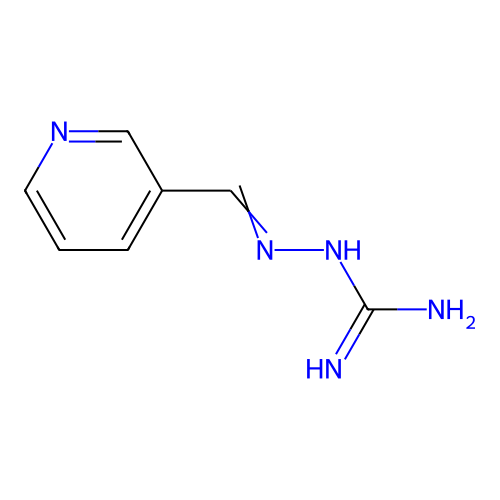
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 50005311
Affinity DataIC50: 3.63E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.03E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.08E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.76E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.94E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.51E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.71E+3nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.29E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.51E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.63E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.69E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.88E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.24E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.31E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.80E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.89E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.27E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.01E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.75E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.89E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.31E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.08E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.24E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.24E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.41E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.76E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.13E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 9.12E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 9.55E+4nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.24E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.29E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.37E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.76E+5nMAssay Description:Displacement of [3H]MK-801 from NMDA receptor complex (unknown origin) after 40 minsMore data for this Ligand-Target Pair