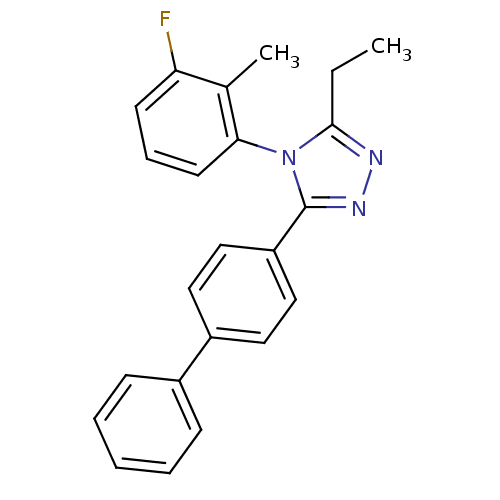
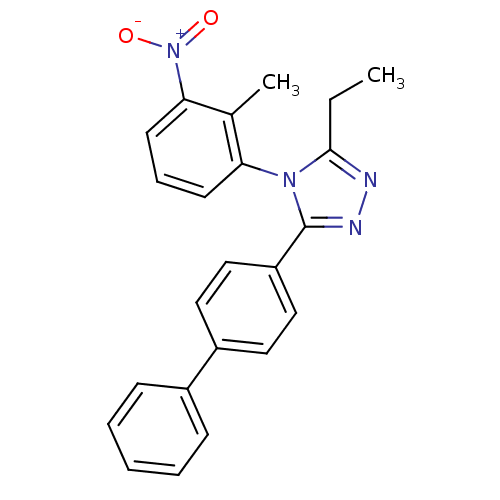
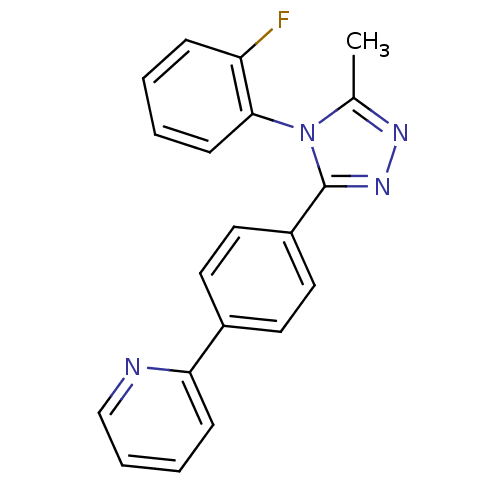
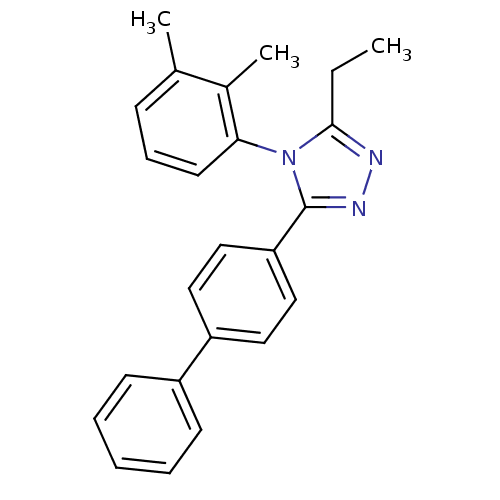
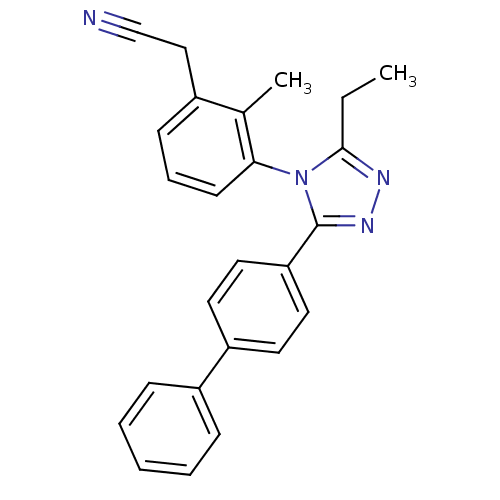
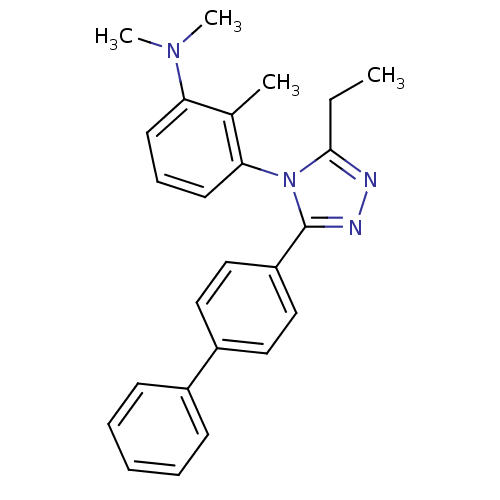
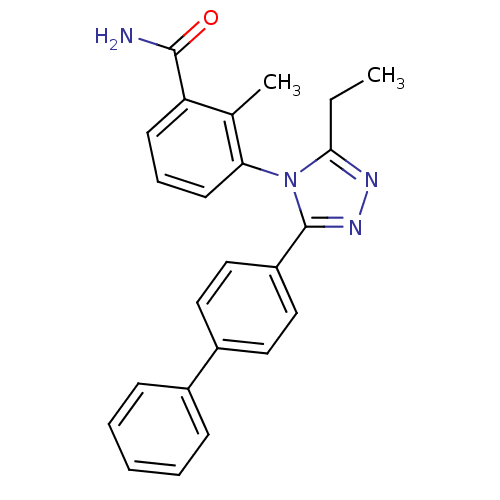
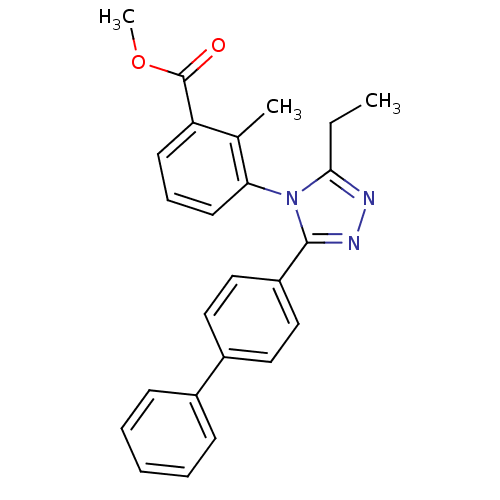
Report error Found 50 Enz. Inhib. hit(s) with all data for entry = 50043356
Affinity DataIC50: 8.90nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 590nMAssay Description:Inhibition of GlyT1 in human SK-N-MC cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 860nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 890nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 940nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair