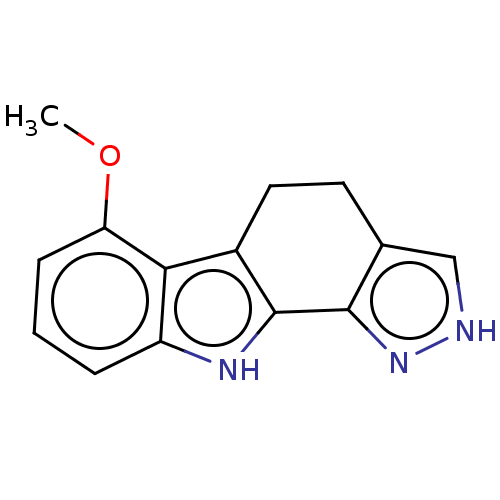
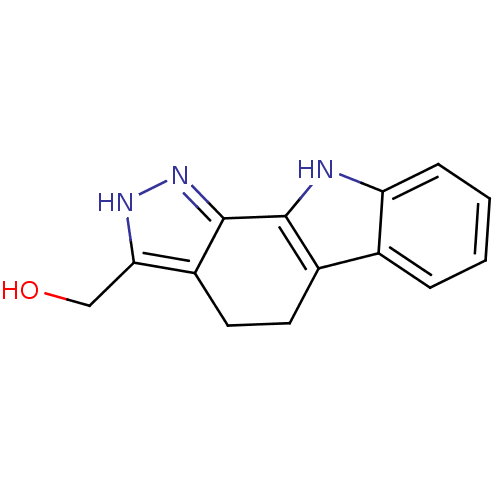
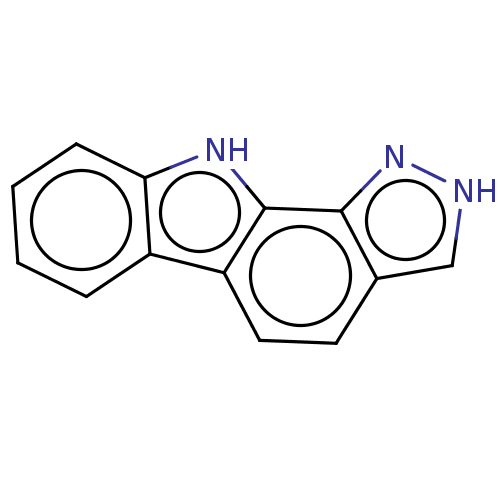
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 50005574
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
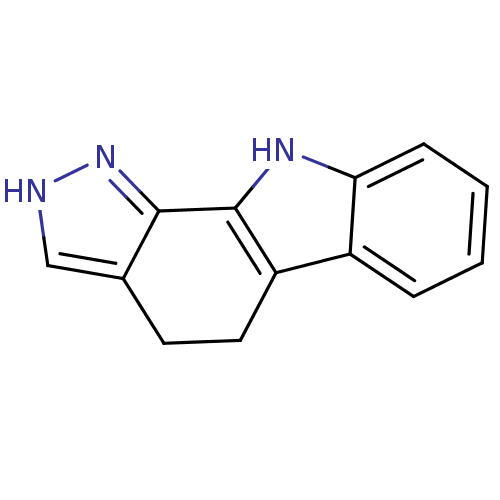
Affinity DataEC50: 14nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
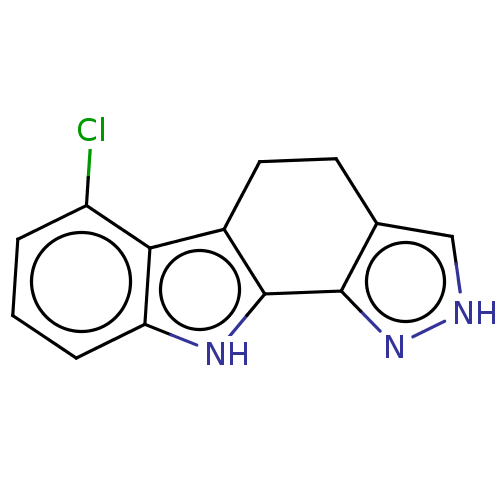
Affinity DataEC50: 47nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
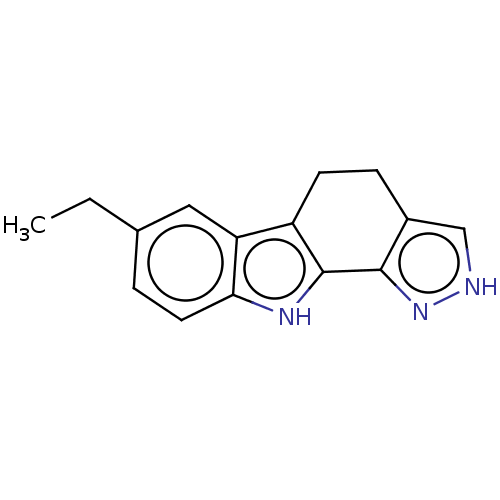
Affinity DataEC50: 80nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
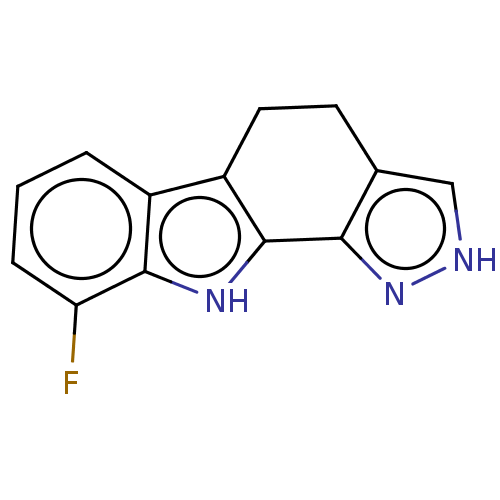
Affinity DataEC50: 100nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 110nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 110nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 140nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 150nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 170nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 250nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 250nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 500nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 640nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 670nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 760nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 780nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 810nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 860nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 1.45E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 2.11E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 2.64E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 3.62E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 4.86E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 5.79E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair
TargetMitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1(Human)
Chinese Academy of Sciences
Curated by ChEMBL
Chinese Academy of Sciences
Curated by ChEMBL
Affinity DataEC50: 6.32E+3nMAssay Description:Inhibition of RIP1/JNK2 (unknown origin)-mediated necrosisMore data for this Ligand-Target Pair