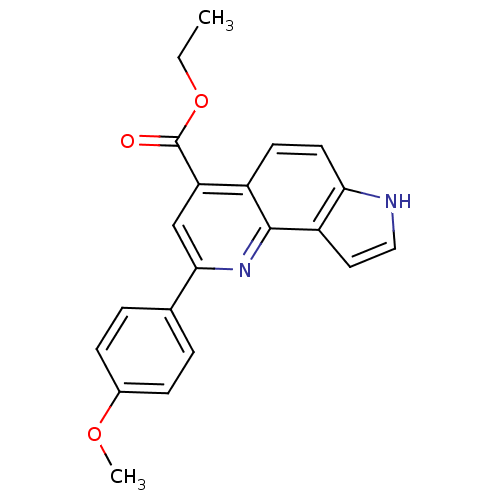
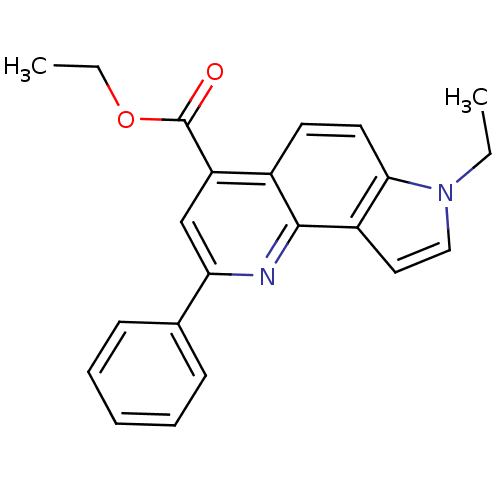
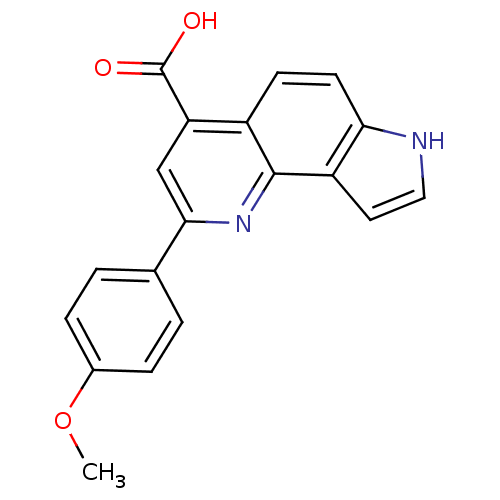
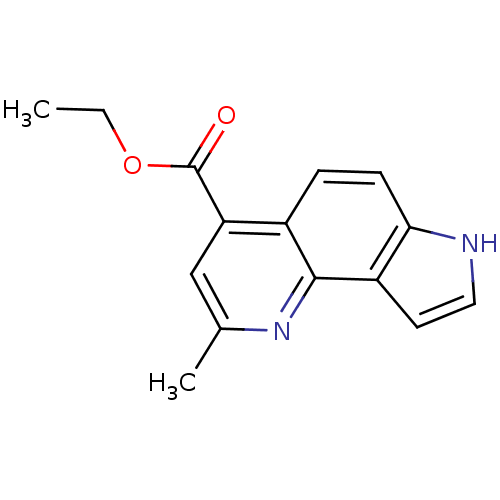
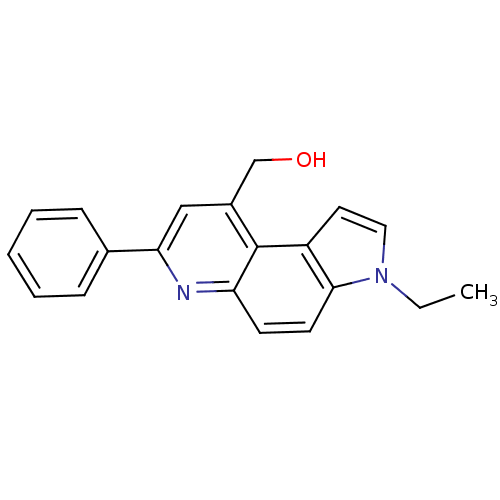
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 50043517
Affinity DataIC50: 3.10nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 75nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 454nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 550nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 590nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 922nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+3nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.47E+3nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+3nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.11E+3nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+3nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 4.15E+3nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human CYP11B1 using [1,2-3H]-11-deoxycorticosterone as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human CYP17 expressed in Escherichia coli using progesterone as substrate NADPH as cofactorMore data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of aromatase (unknown origin) using 7-methoxy-4-trifluoromethylcoumarin as substrate after 30 mins by fluorimetric analysisMore data for this Ligand-Target Pair