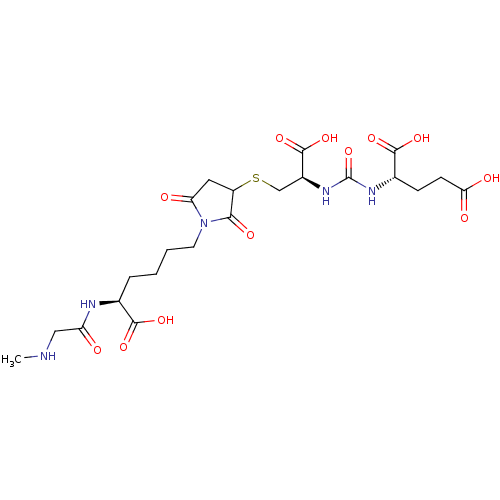
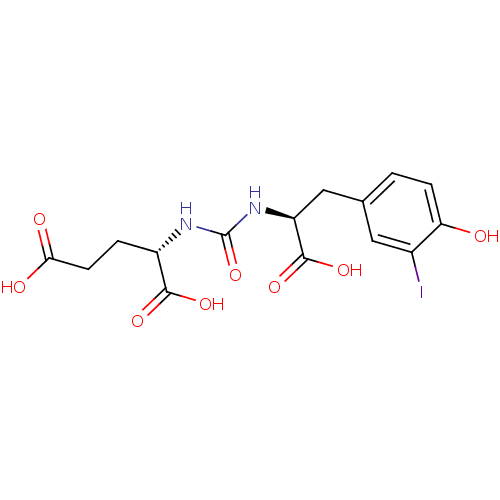
Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 50043444
Affinity DataKi: 2.10nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKd: 7.80nMAssay Description:Binding affinity to PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKd: 20nMAssay Description:Binding affinity to PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 95nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKd: 143nMAssay Description:Binding affinity to PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 376nMAssay Description:Displacement of [123I]-DCIT from PSMA in human LNCaP cells after 1 hr by gamma countingMore data for this Ligand-Target Pair
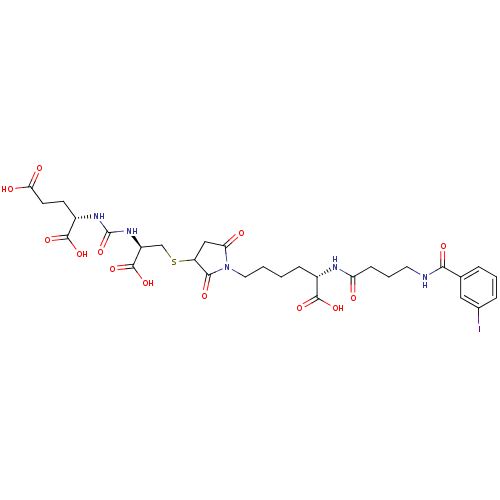

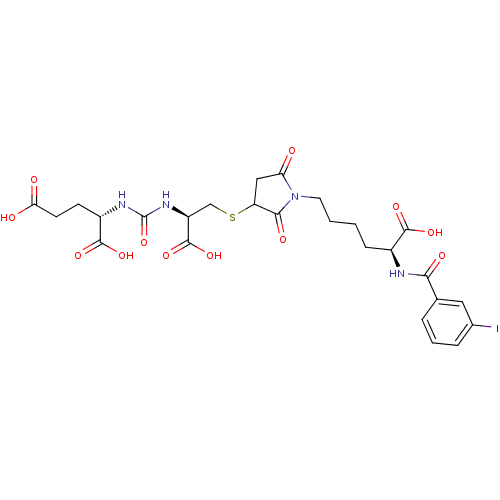



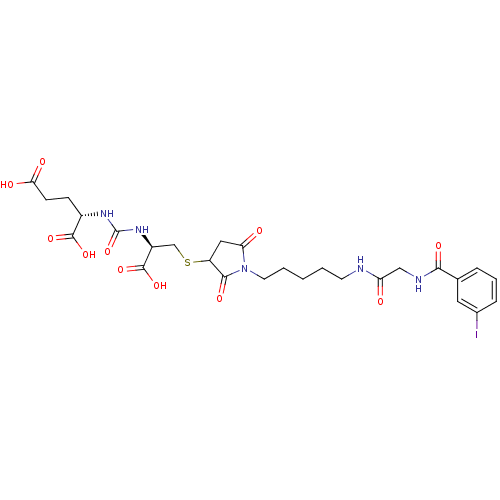
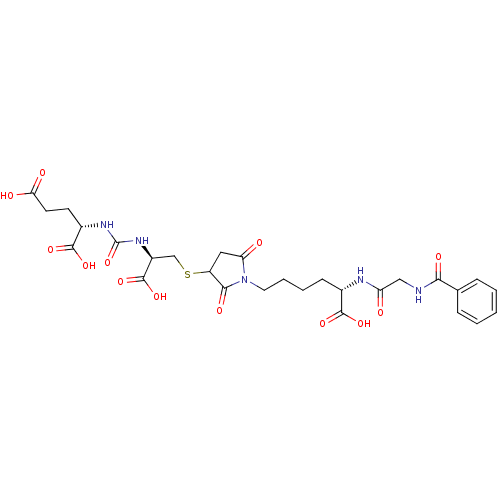

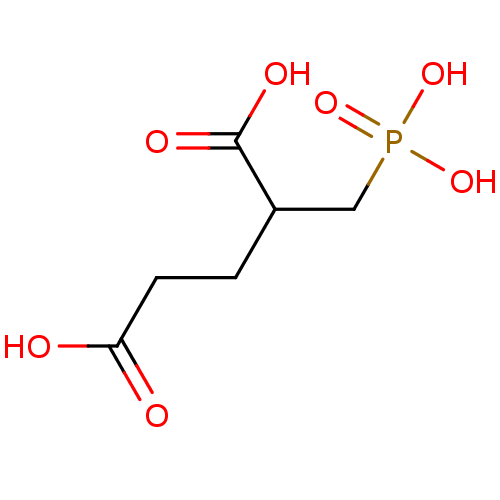
 3D Structure (crystal)
3D Structure (crystal)