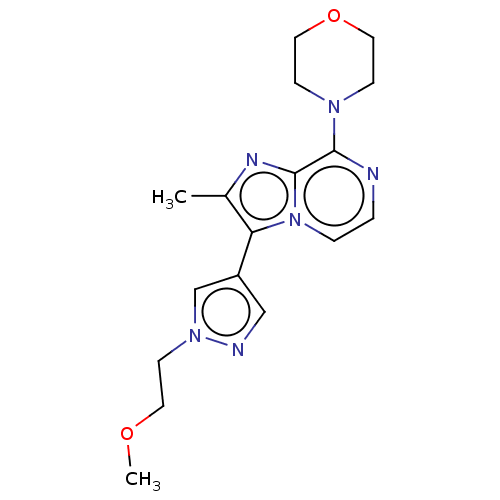
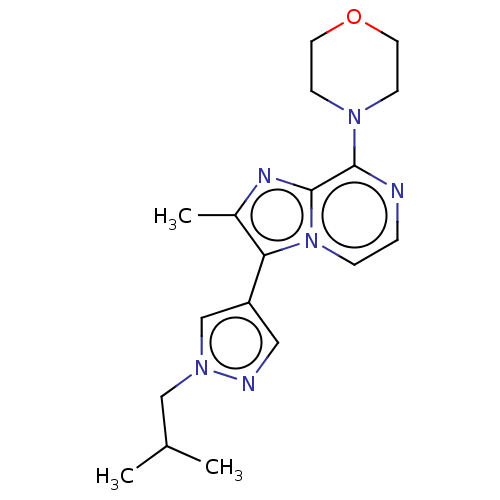
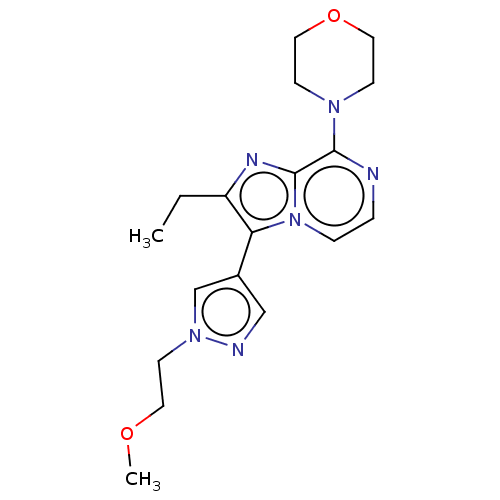
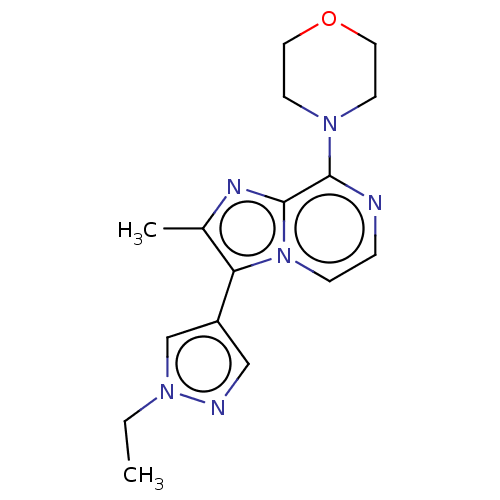
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 50044368
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
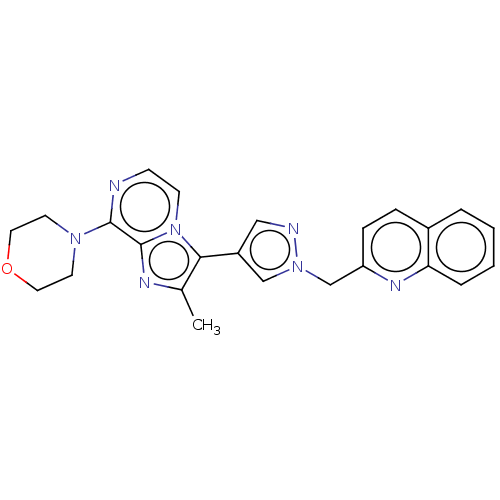
Affinity DataIC50: 0.646nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
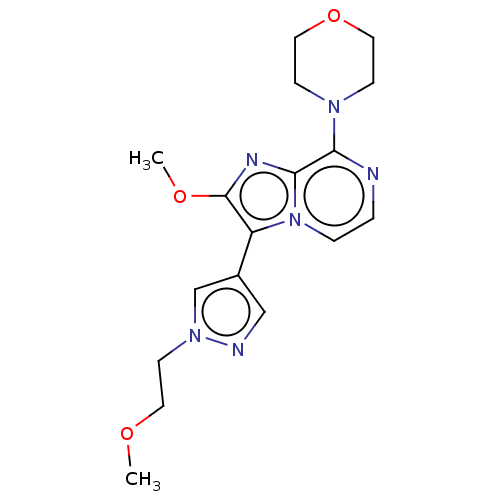
Affinity DataIC50: 47nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
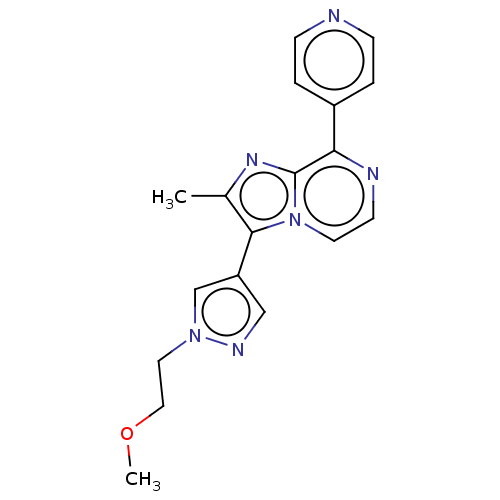
Affinity DataIC50: 54nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
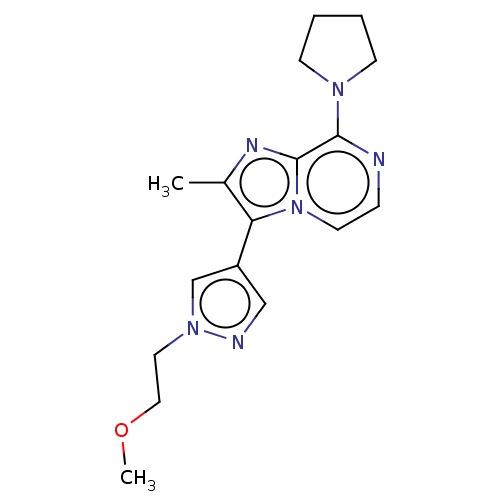
Affinity DataIC50: 60nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 65nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 66nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 85nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 102nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 112nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 112nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 138nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 141nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 145nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 155nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 158nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 234nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 234nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 257nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 309nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 347nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 501nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 562nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of rat recombinant PDE10A expressed in baculovirus infected Sf9 cells using [3H]cAMP as substrate after 60 mins by TopCount scintillation ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE5A (unknown origin)More data for this Ligand-Target Pair
TargetRod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha(Human)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of PDE6A (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Inhibition of PDE3A (unknown origin)More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 2.51E+3nMAssay Description:Inhibition of PDE1B (unknown origin)More data for this Ligand-Target Pair
TargetRod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha(Human)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PDE6A (unknown origin)More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PDE1B (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP2D6 (unknown origin)More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Janssen Pharmaceutica
Curated by ChEMBL
Janssen Pharmaceutica
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PDE5A (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP1A2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PDE3A (unknown origin)More data for this Ligand-Target Pair