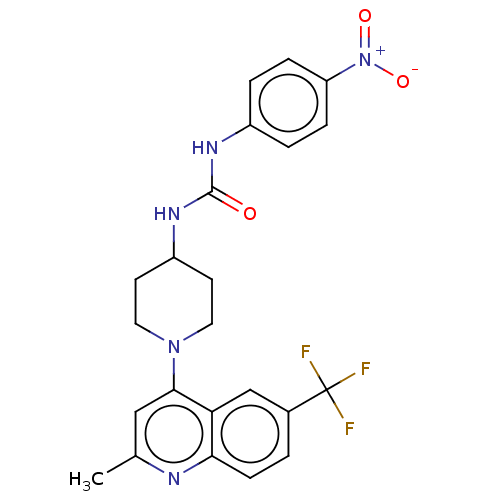
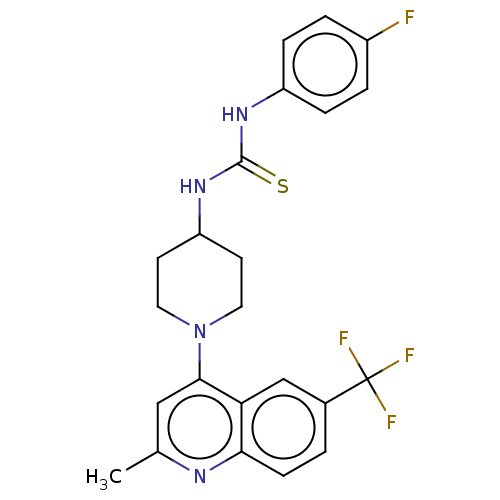
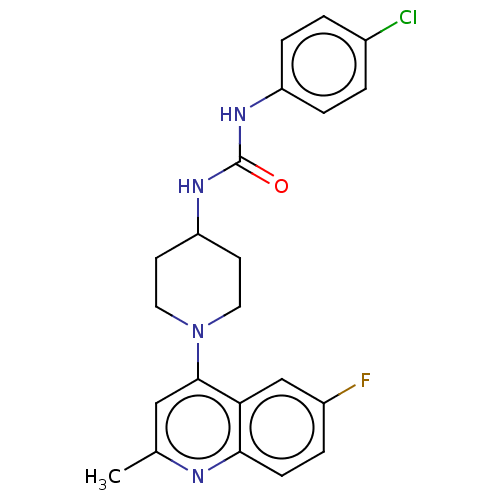
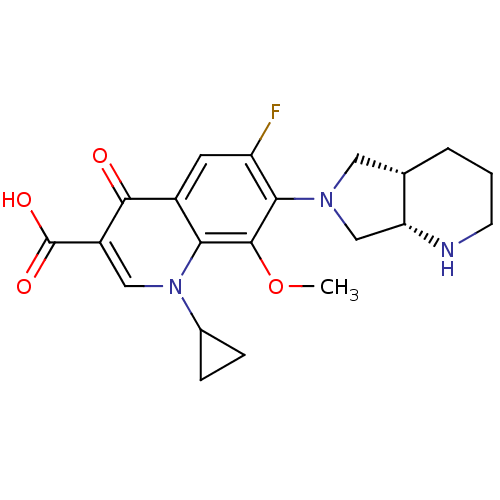
Report error Found 50 Enz. Inhib. hit(s) with all data for entry = 50045536
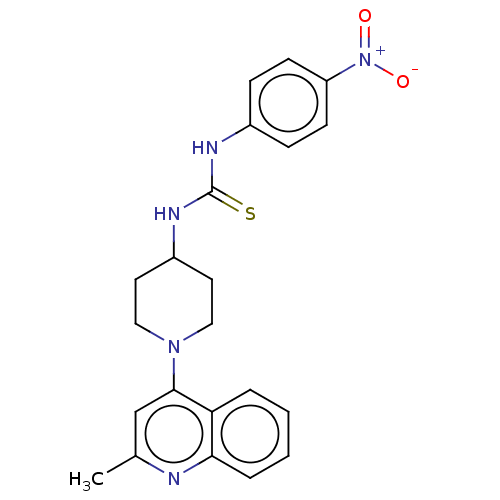
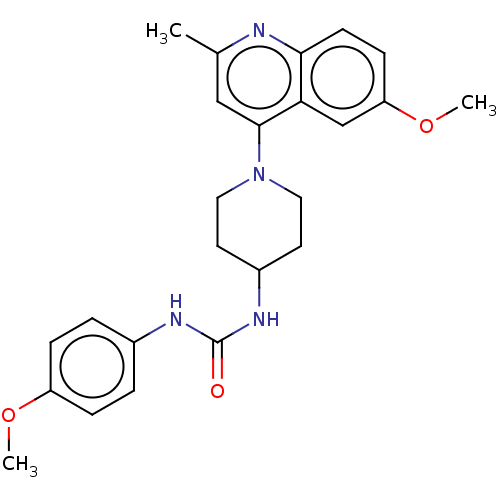
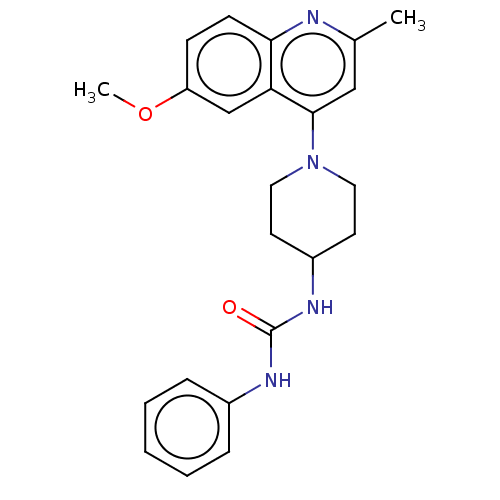
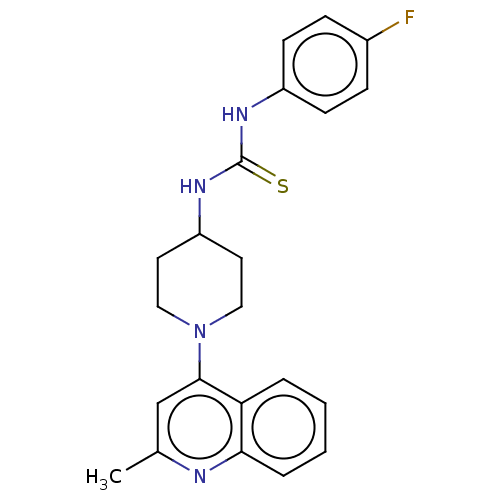
Affinity DataIC50: 180nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
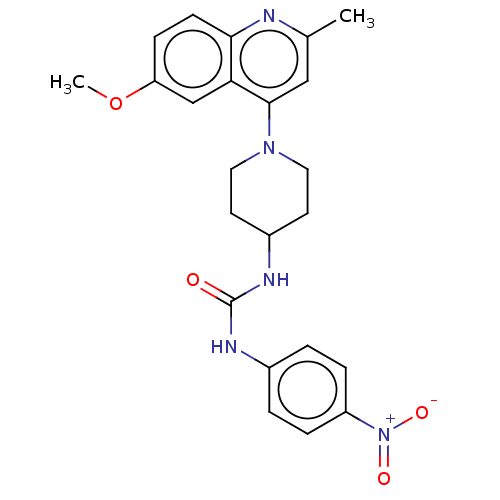
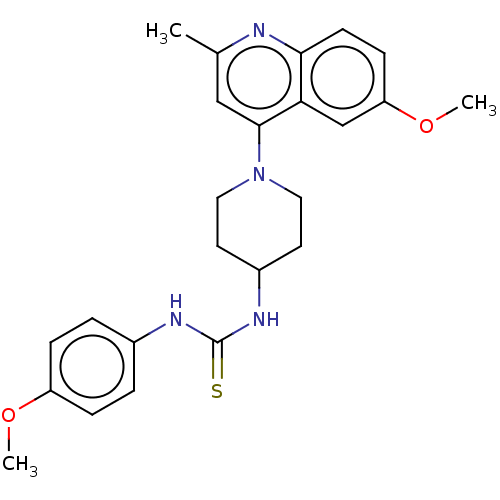
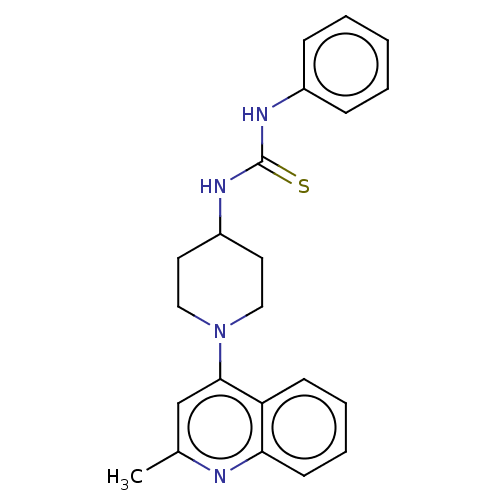
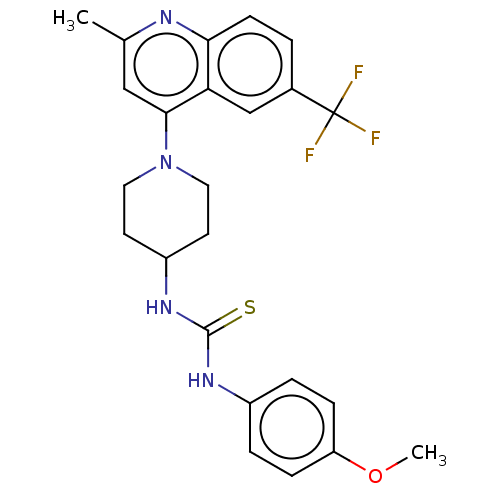
Affinity DataIC50: 950nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
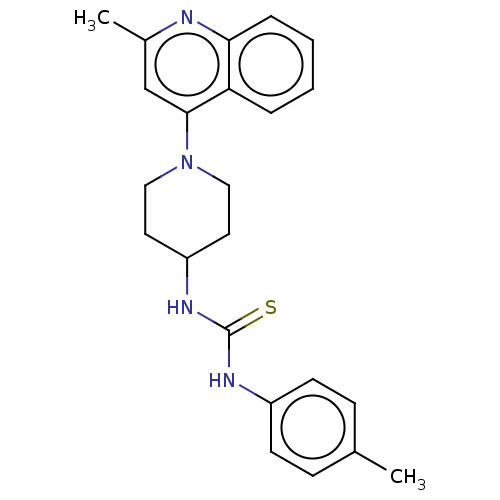
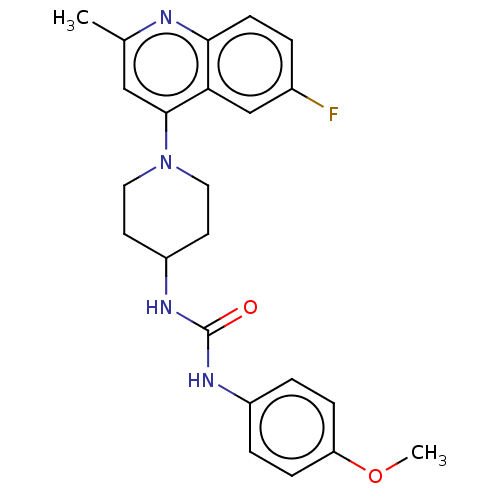
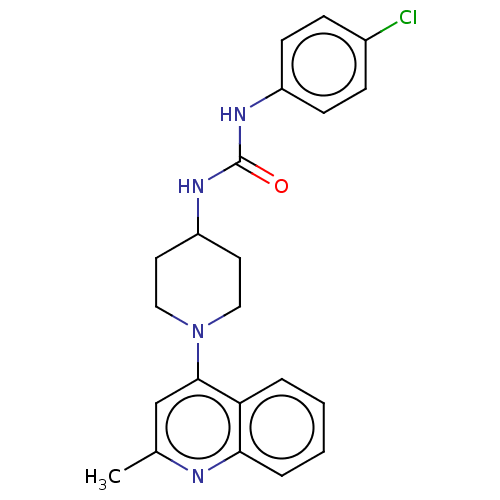
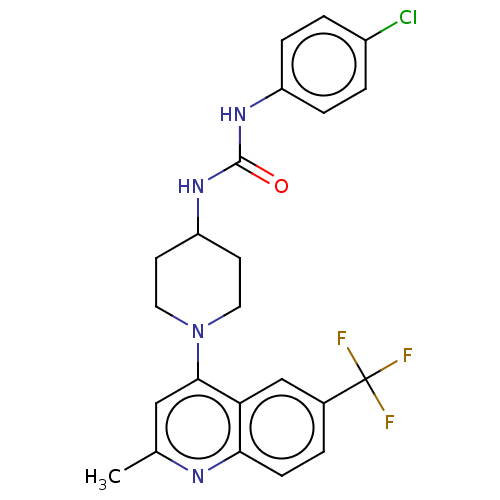
Affinity DataIC50: 1.78E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
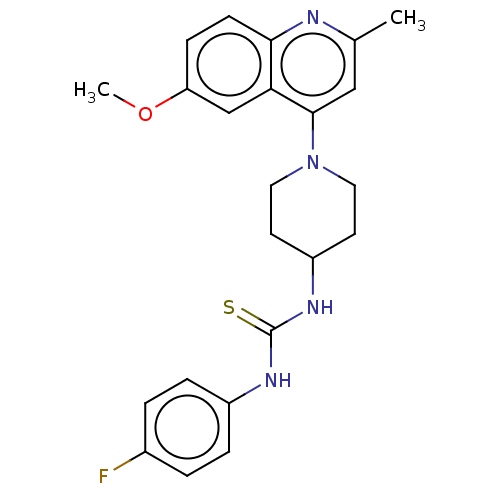
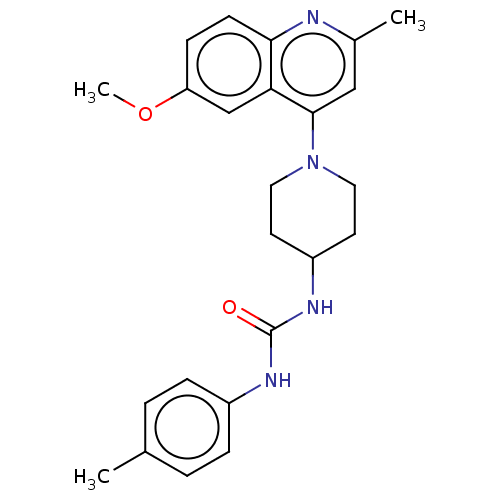
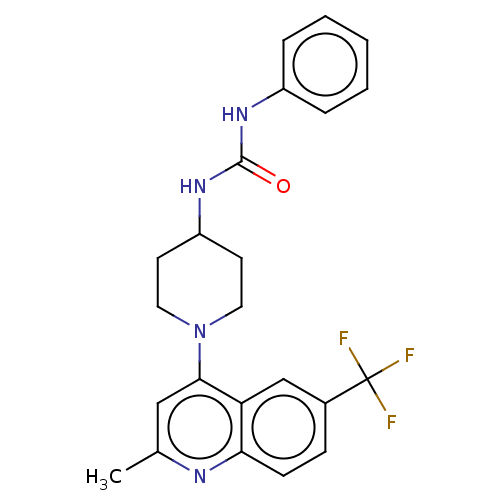
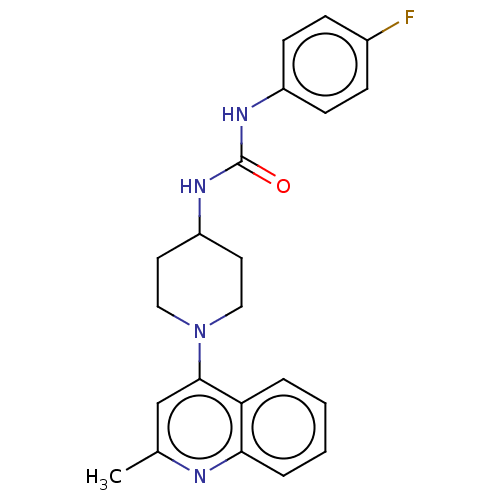
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.77E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 6.83E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 6.99E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 7.26E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 9.25E+3nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.27E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.29E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.49E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.65E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.94E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.98E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.47E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 2.98E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.49E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 3.95E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.11E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.12E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.18E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.59E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.76E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.85E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 5.62E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 5.93E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 6.82E+4nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair