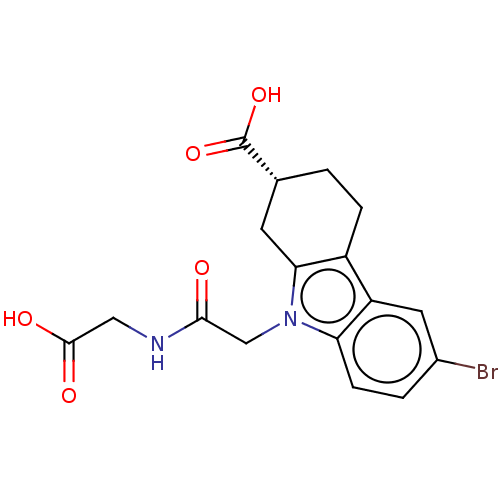
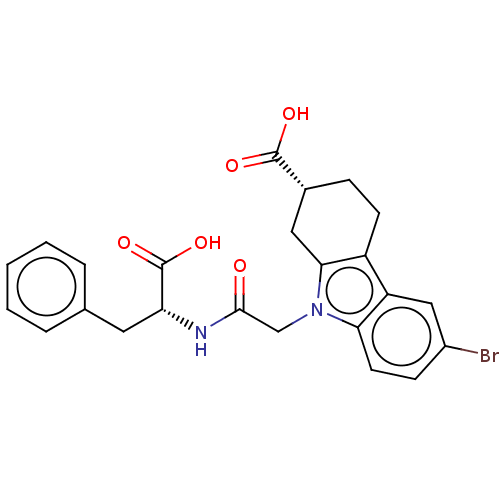
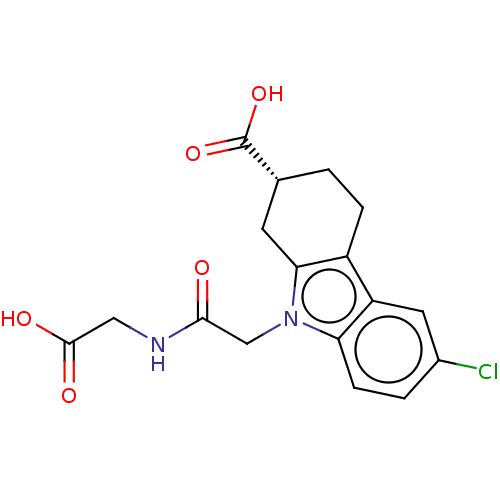
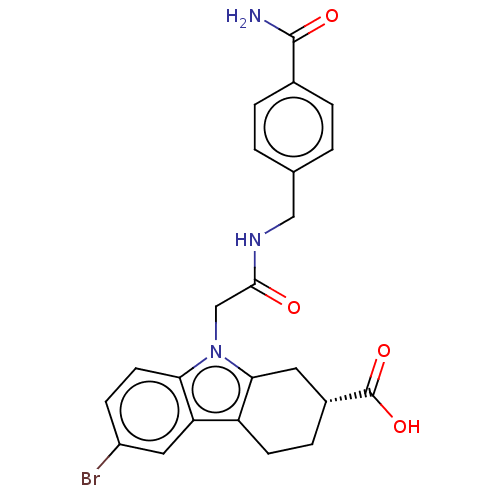
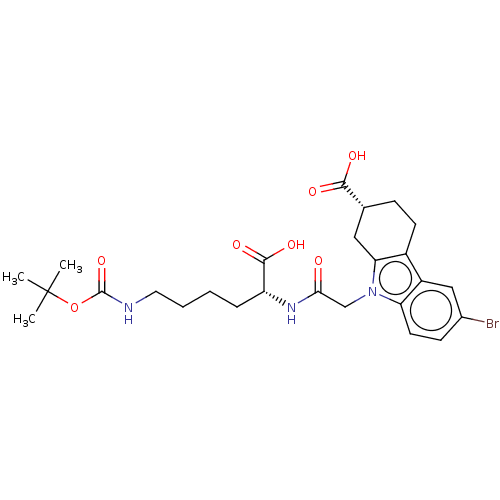
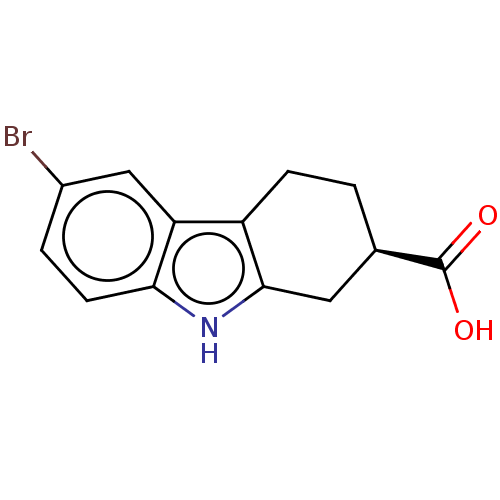
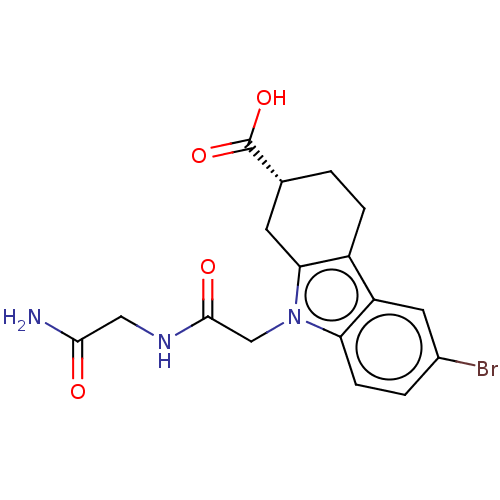
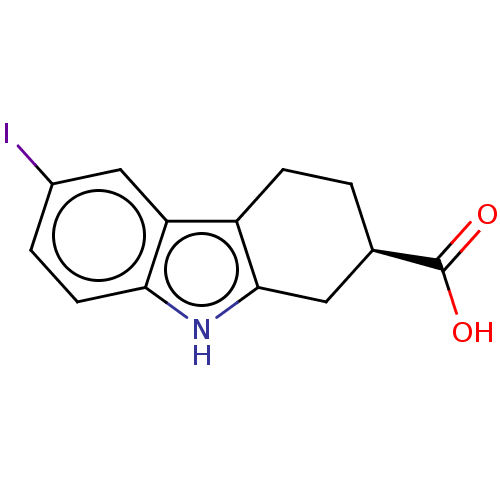
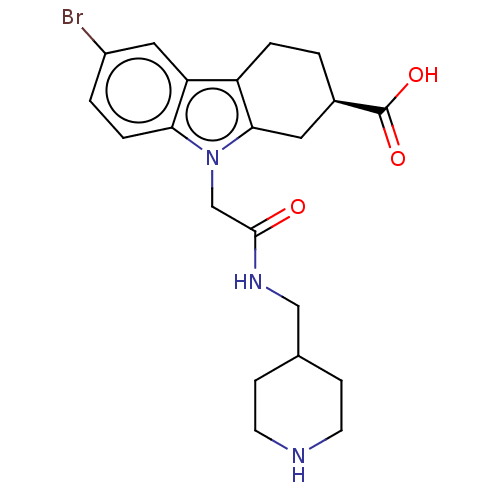
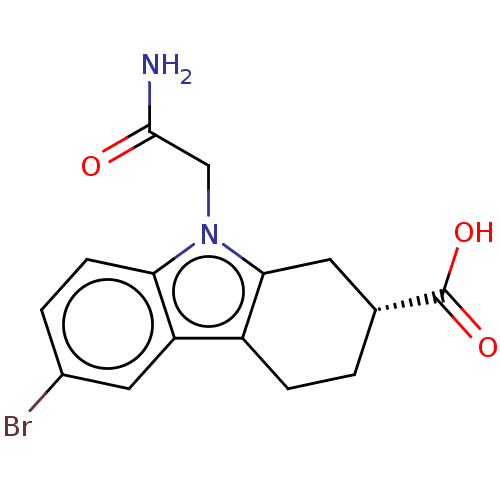
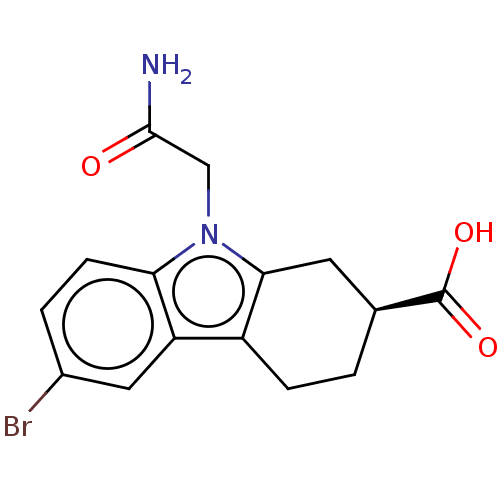
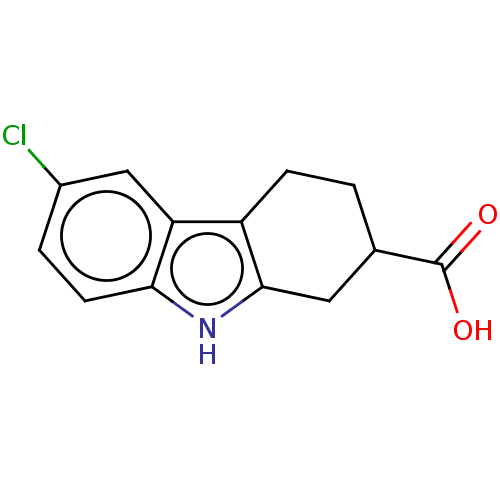
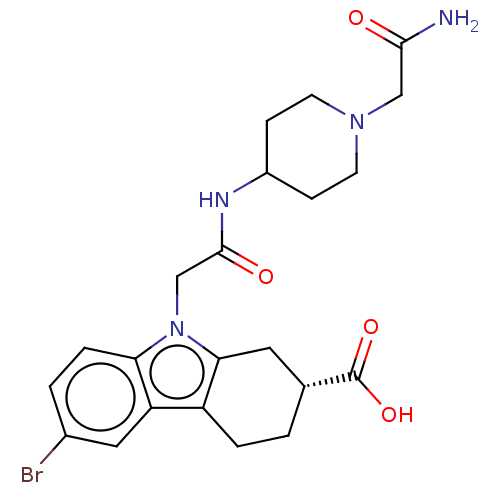
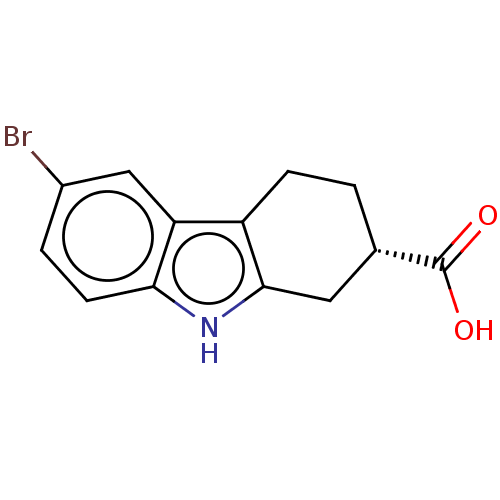
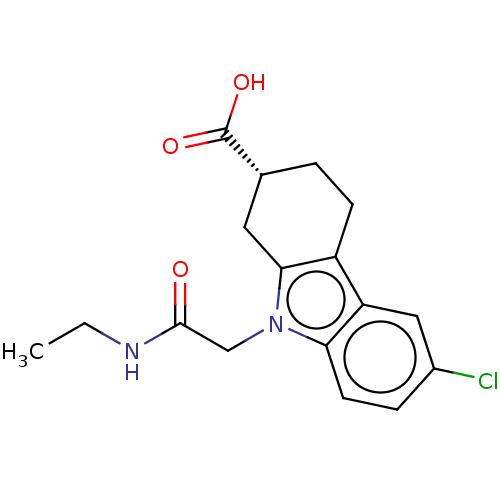
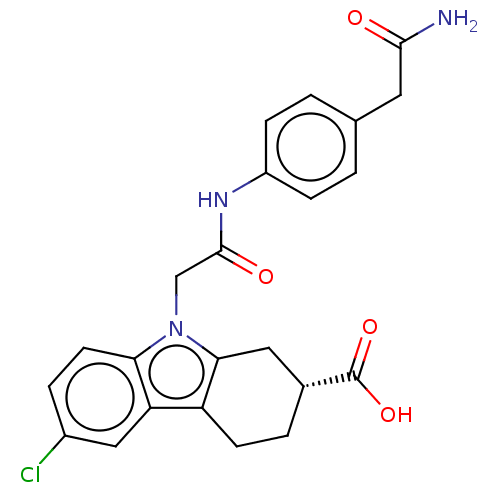
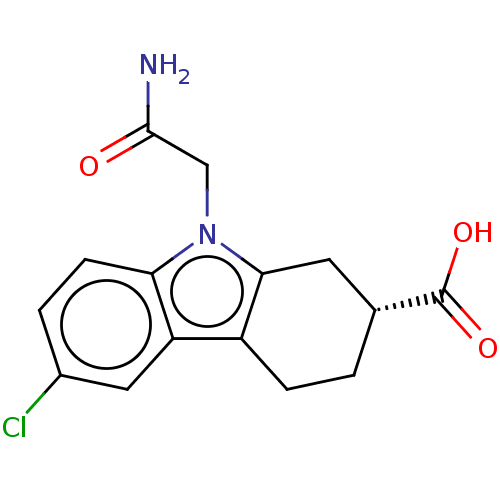
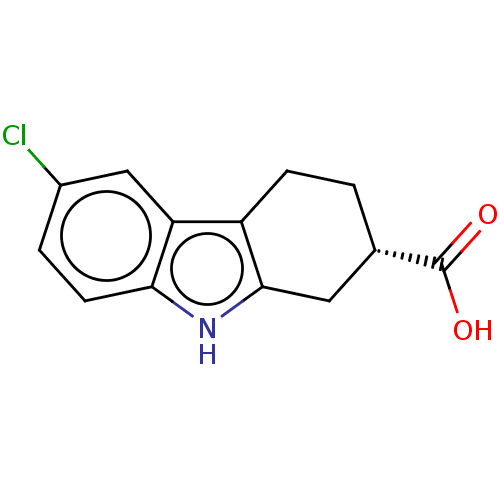
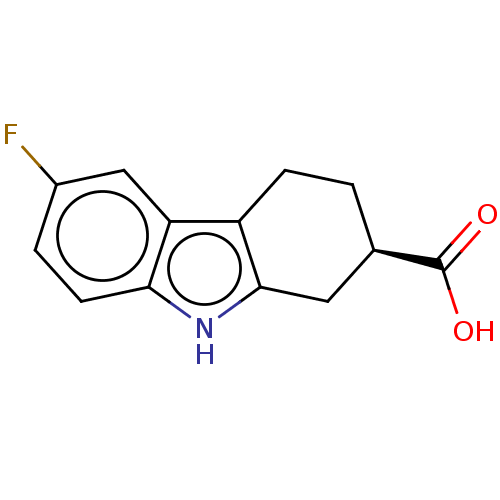
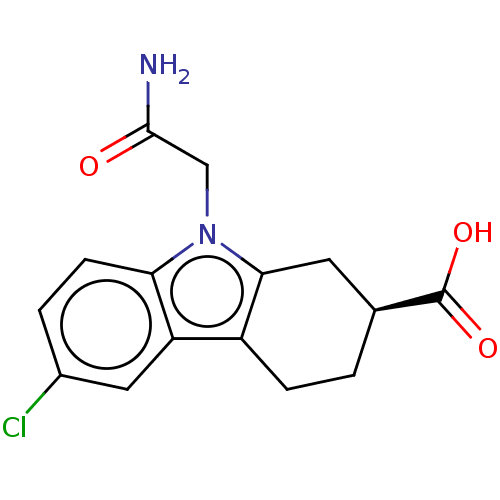
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 50046152
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKd: 9.00E+3nMAssay Description:Binding affinity to Escherichia coli DNA sliding clamp protein at 298K by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKd: 1.30E+4nMAssay Description:Binding affinity to Escherichia coli DNA sliding clamp protein at 298K by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKd: 1.40E+4nMAssay Description:Binding affinity to Escherichia coli DNA sliding clamp protein at 298K by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.50E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKd: 2.40E+4nMAssay Description:Binding affinity to Escherichia coli DNA sliding clamp protein at 298K by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 2.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 3.60E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 4.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 6.40E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.40E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 7.40E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 8.30E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 8.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 9.50E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 9.70E+4nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.15E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.16E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.34E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.37E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.50E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.57E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.58E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.66E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.71E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.71E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.75E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.82E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.93E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 2.01E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 2.04E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.08E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 2.46E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.46E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.85E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.99E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.07E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.28E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.47E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.62E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.68E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.42E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 4.54E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBeta sliding clamp(Escherichia coli (strain K12))
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
The University of Wollongong and The Illawarra Health and Medical Research Institute
Curated by ChEMBL
Affinity DataKi: 4.93E+5nMAssay Description:Inhibition of Escherichia coli DNA sliding clamp protein using N-fluorescein (FAM)-QLDLF-OH tracer by fluorescence polarization assayMore data for this Ligand-Target Pair
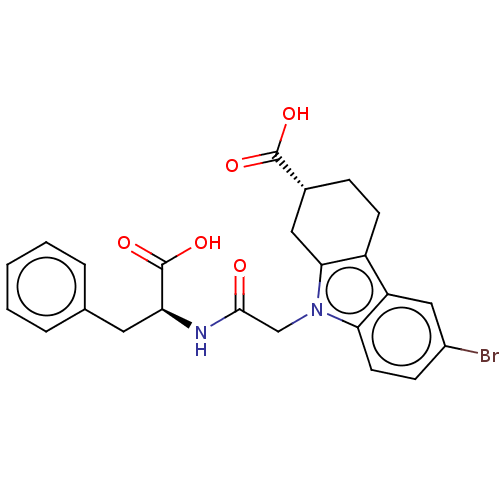
 3D Structure (crystal)
3D Structure (crystal)