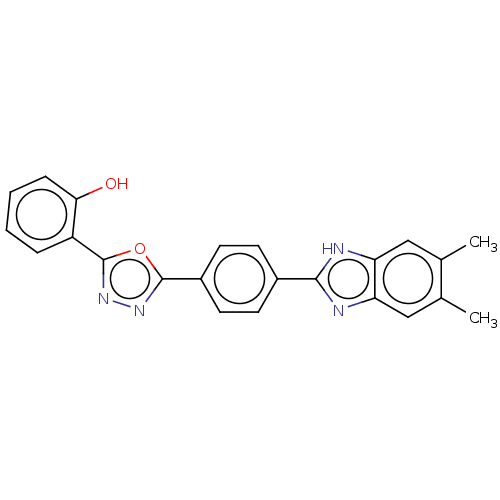
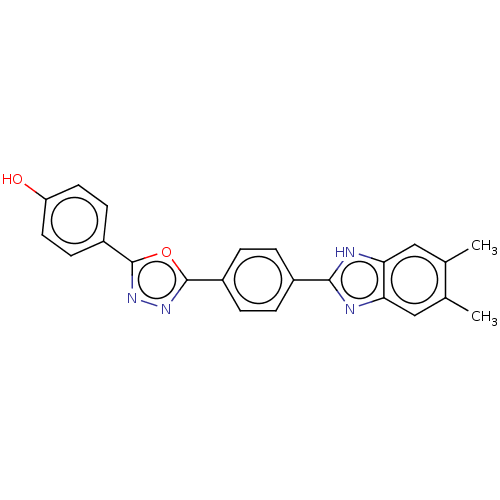
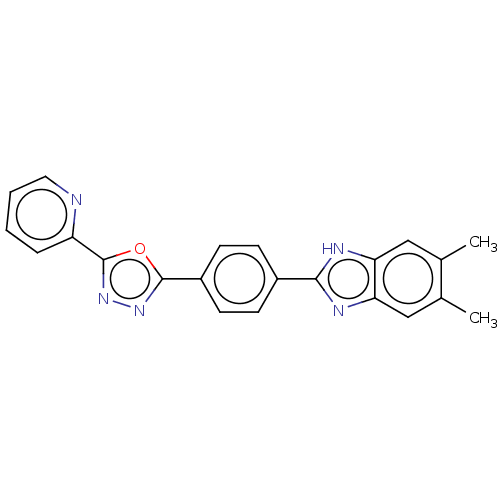
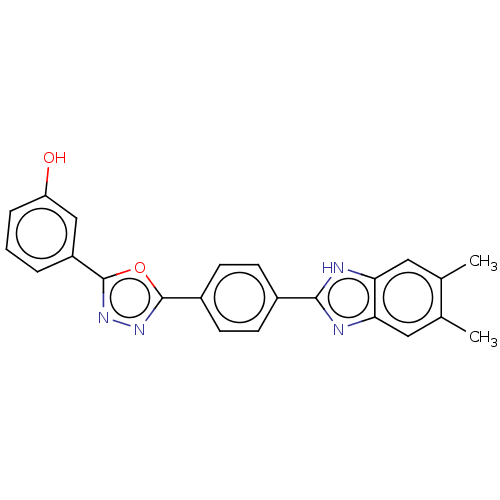
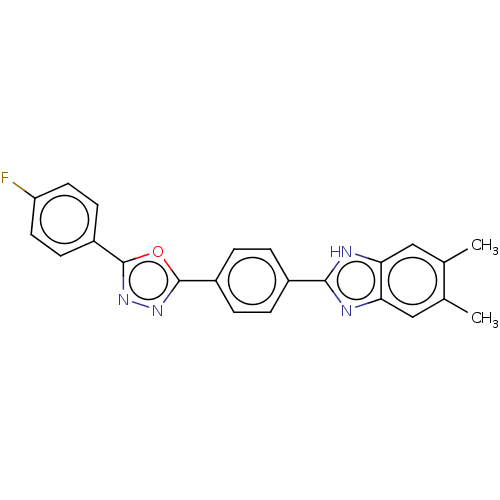
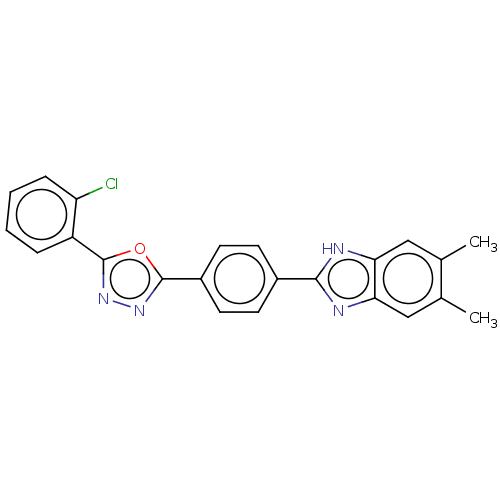
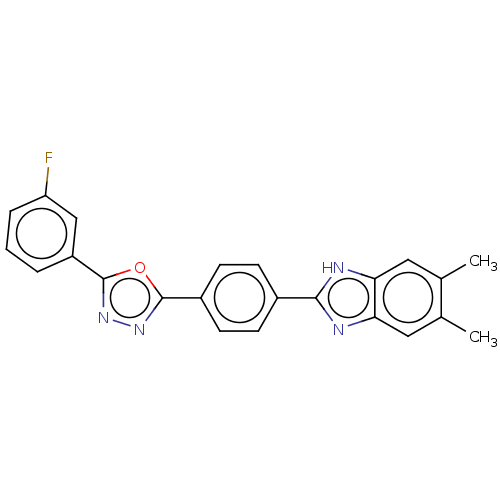
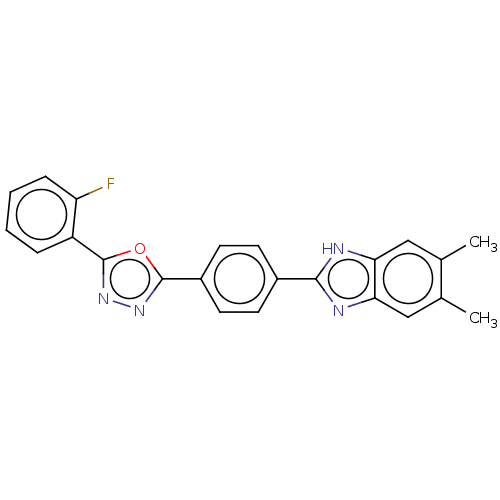
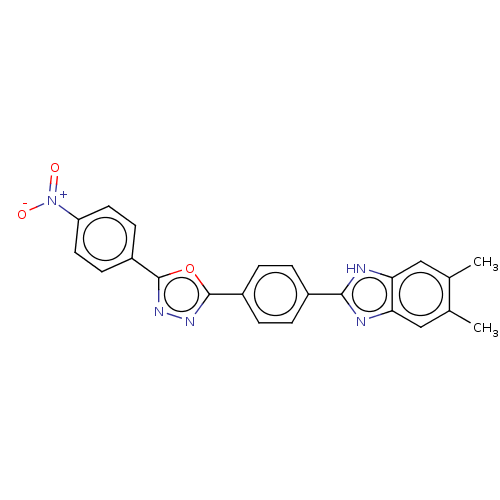
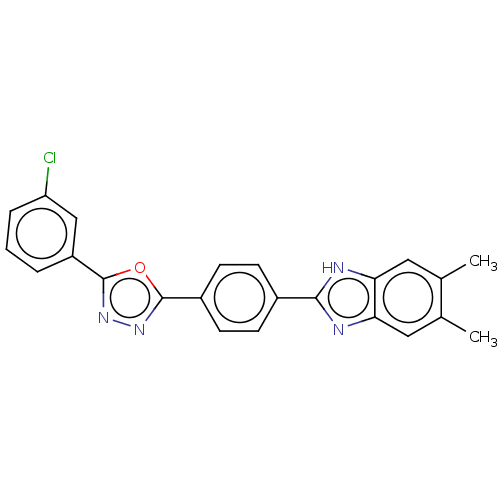
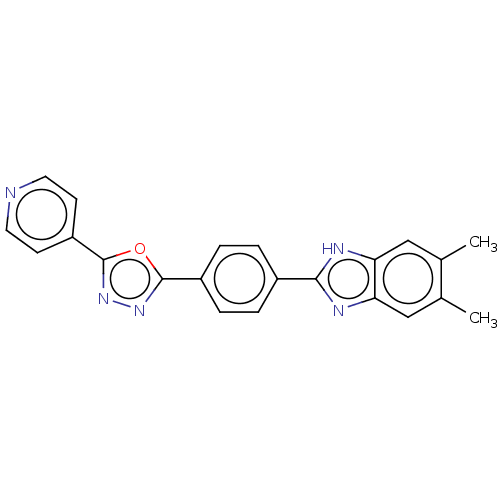
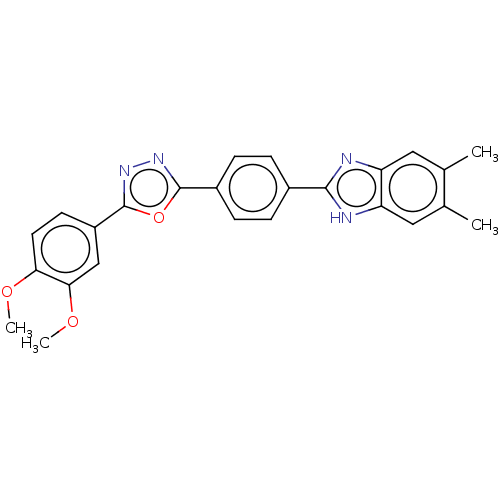
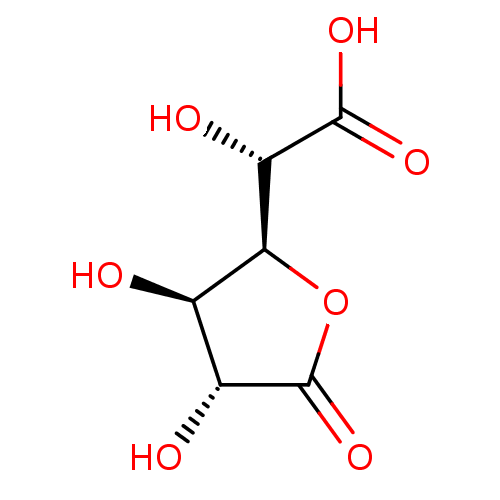
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50046038
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.12E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 8.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.31E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.92E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 2.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.11E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.21E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.61E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.84E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair