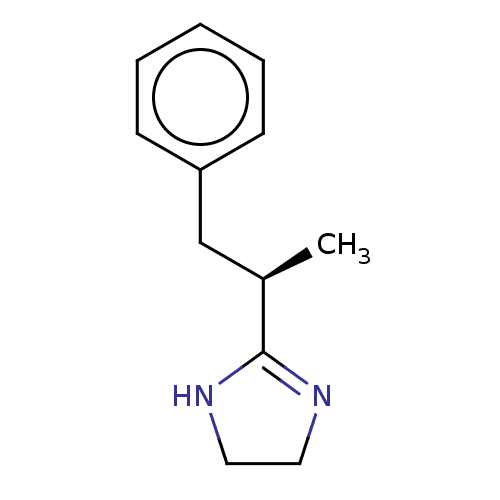
Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 50046020
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 98nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 126nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 155nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 195nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 309nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: <1.00E+3nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: <1.00E+3nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.05E+3nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.24E+3nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 6.31E+3nMAssay Description:Displacement of [125I]-p-iodoclonidin from I1-imidazoline receptor in rat PC12 cell membranesMore data for this Ligand-Target Pair