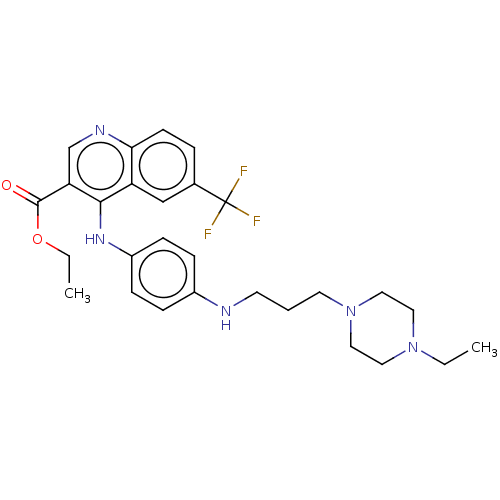
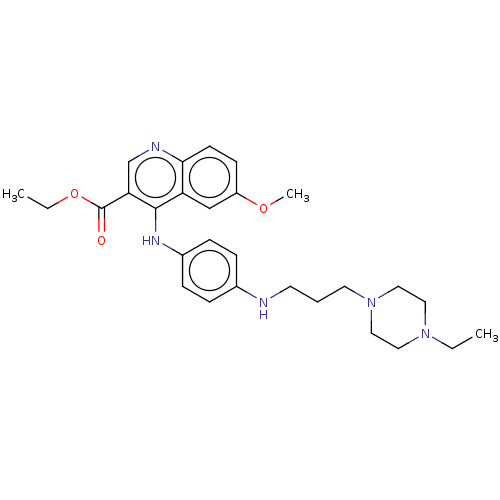
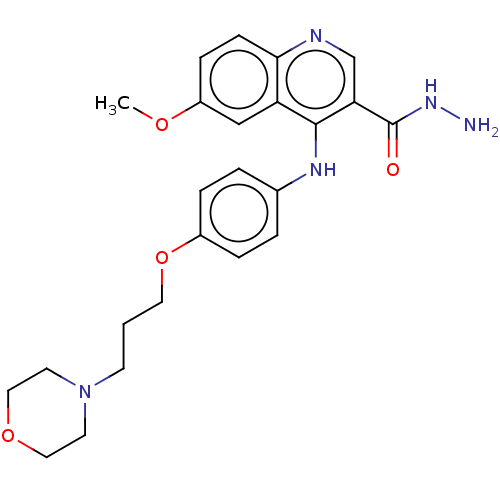
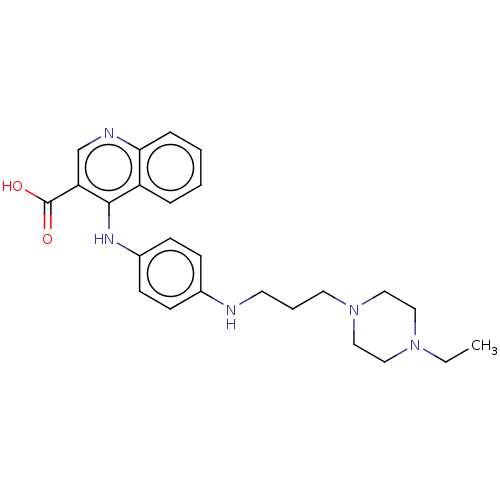
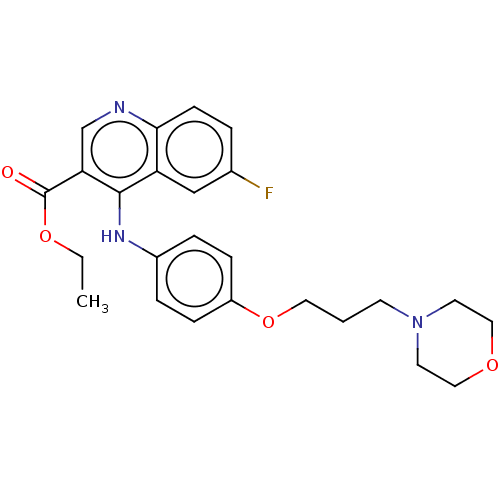
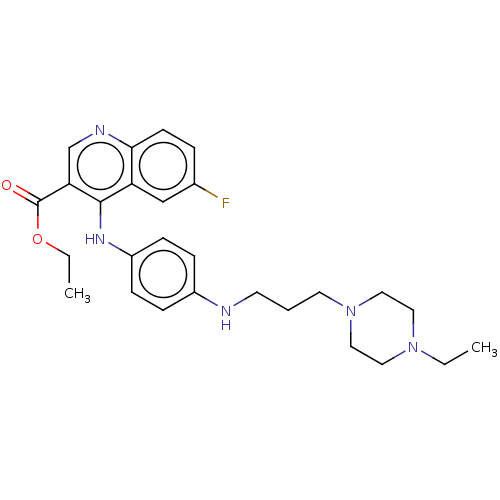
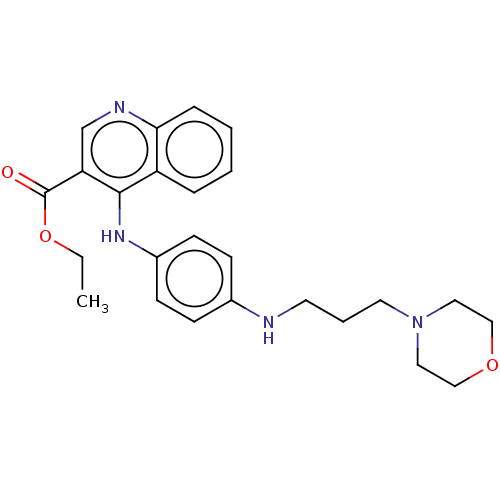
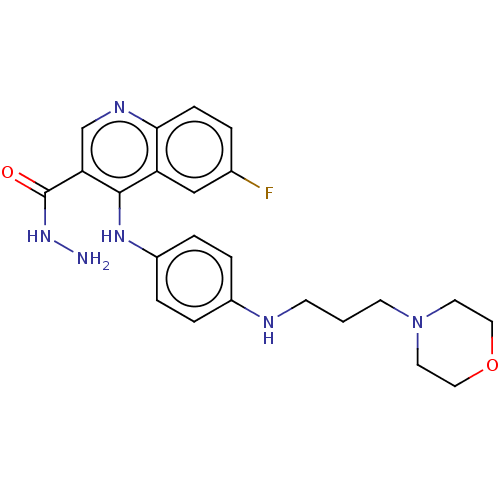
Report error Found 48 Enz. Inhib. hit(s) with all data for entry = 50046700
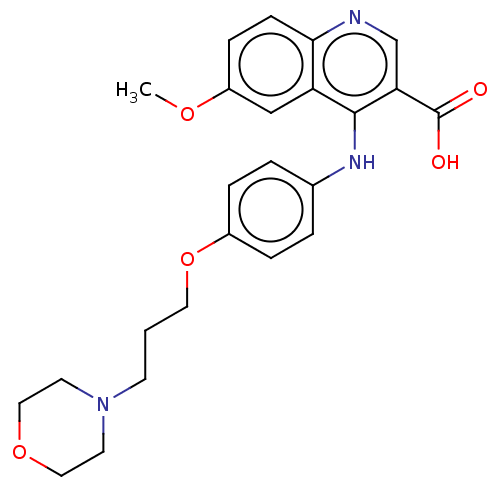
Affinity DataIC50: 180nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
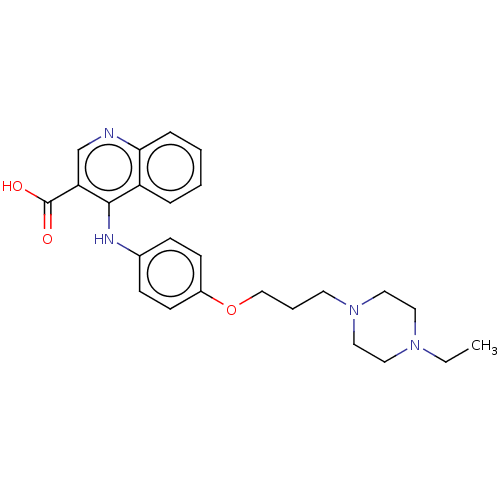
Affinity DataIC50: 860nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
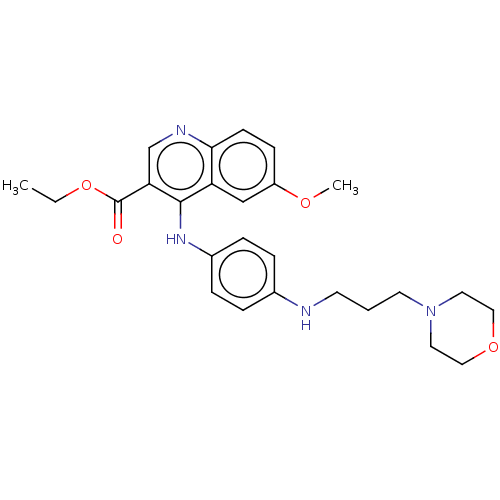
Affinity DataIC50: 970nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
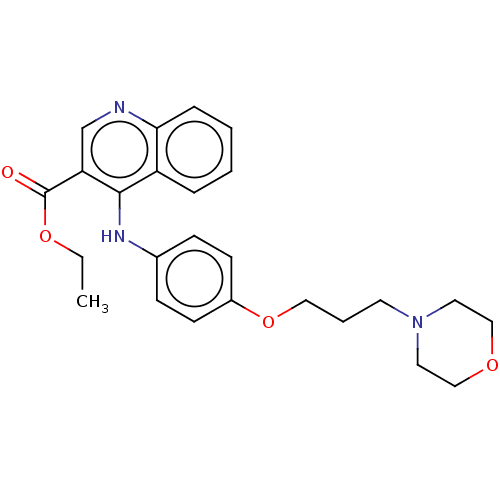
Affinity DataIC50: 970nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.92E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 6.62E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 6.82E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 7.89E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 7.91E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 8.82E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 9.55E+3nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.49E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.69E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.69E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.79E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.87E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.09E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.28E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.36E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.63E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.78E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.84E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.13E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.87E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.88E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.89E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 4.49E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 4.73E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in Escherichia coli BL21 (DE3) pLysS cells assessed as reduction in ATPase activity incu...More data for this Ligand-Target Pair