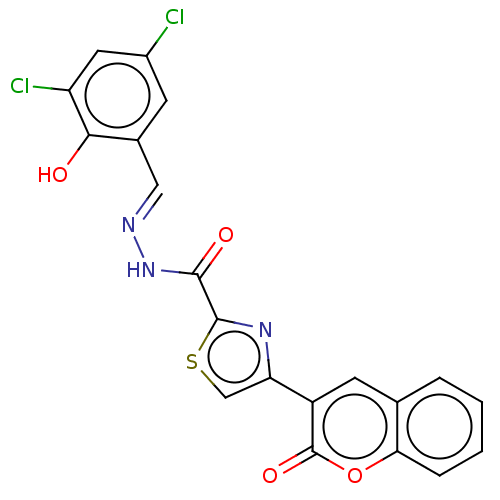
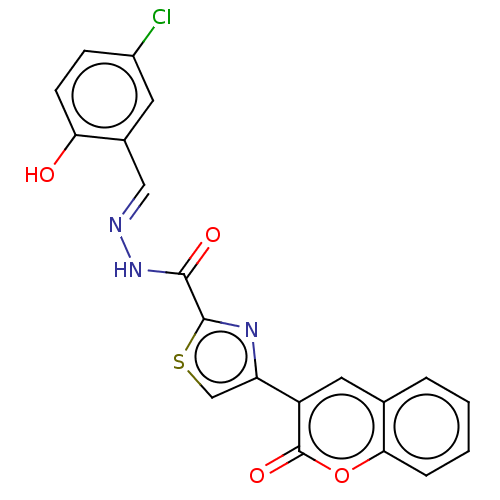
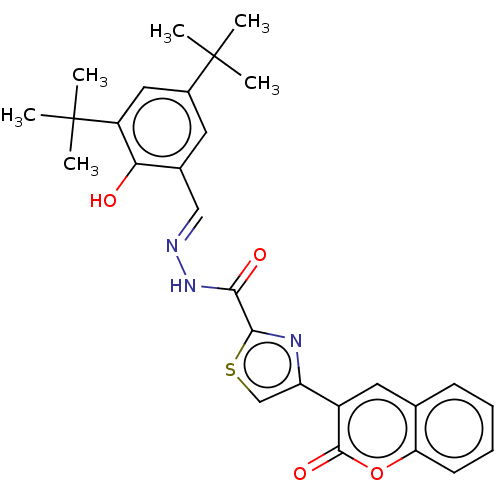
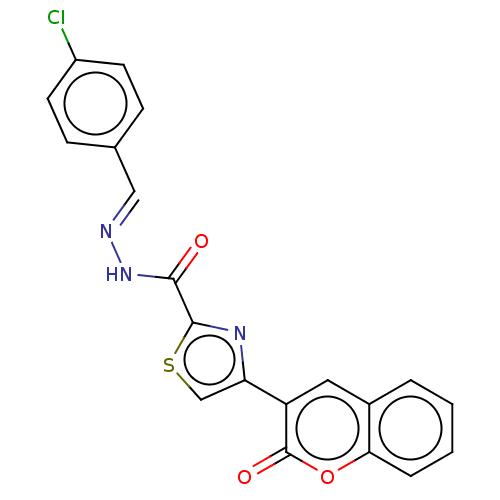
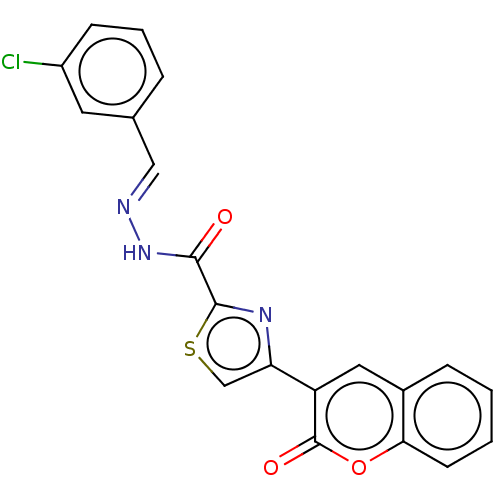
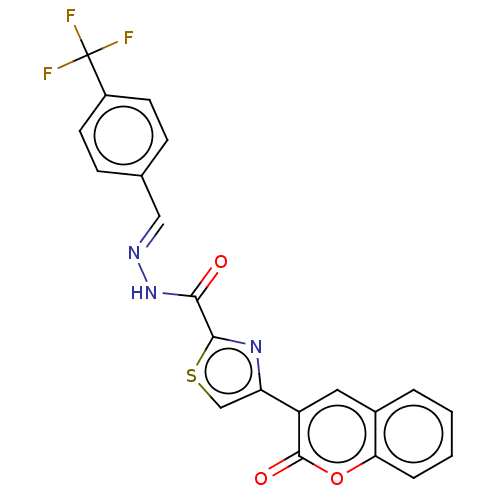
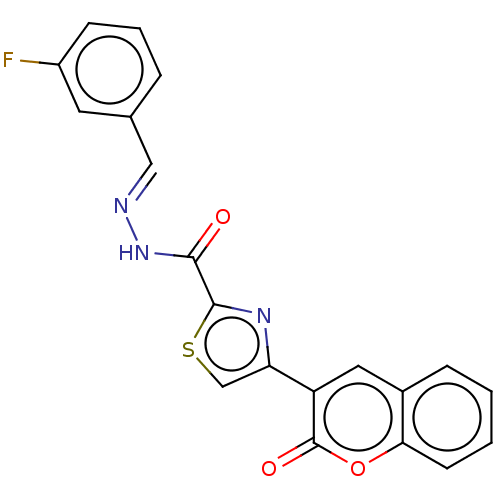
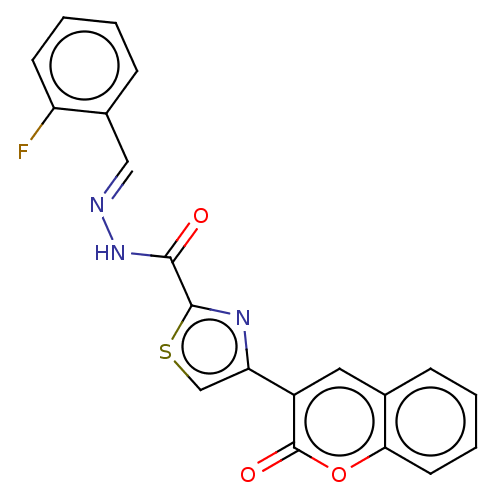
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 7211
Affinity DataIC50: 6.24E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 8.23E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.83E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 2.26E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 2.36E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 2.52E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 2.99E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 3.86E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 4.33E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 6.43E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair
Affinity DataIC50: 8.17E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed by u...More data for this Ligand-Target Pair