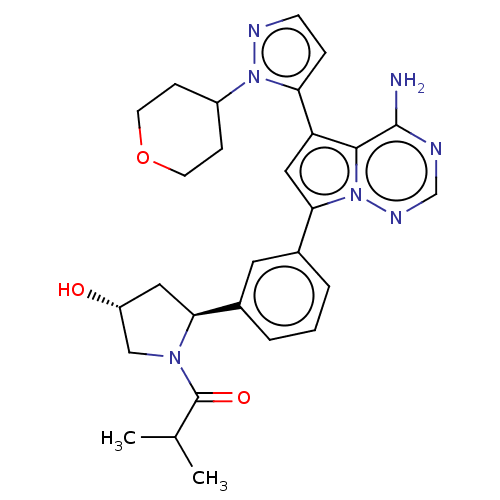
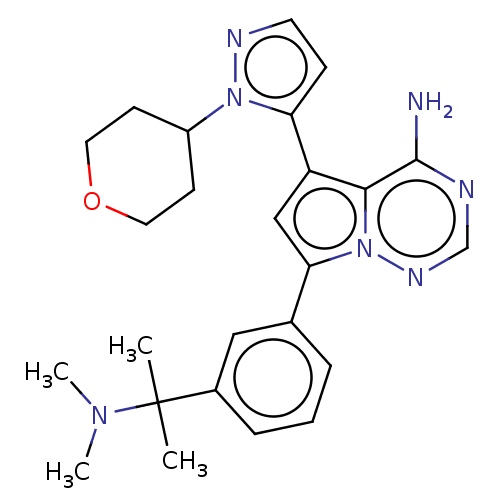
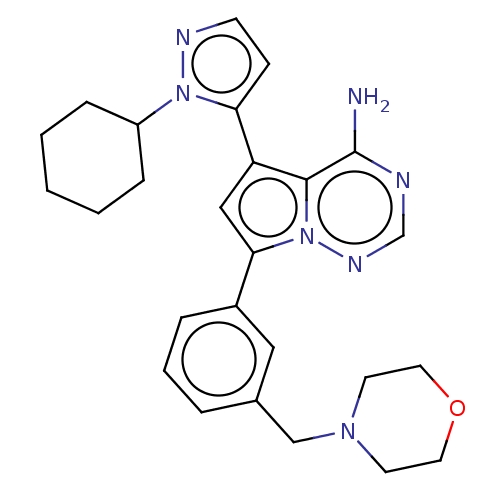
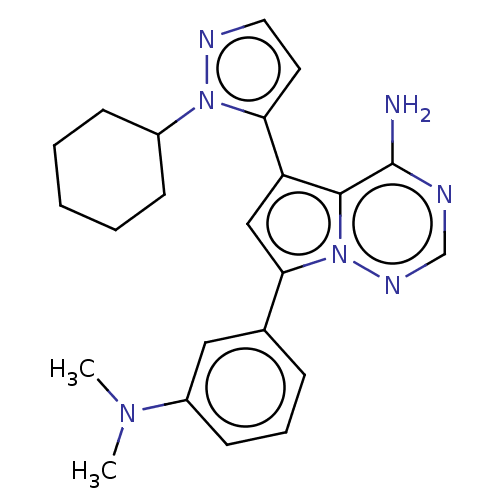
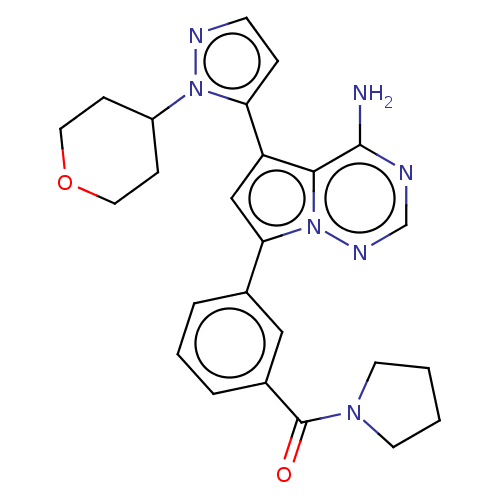
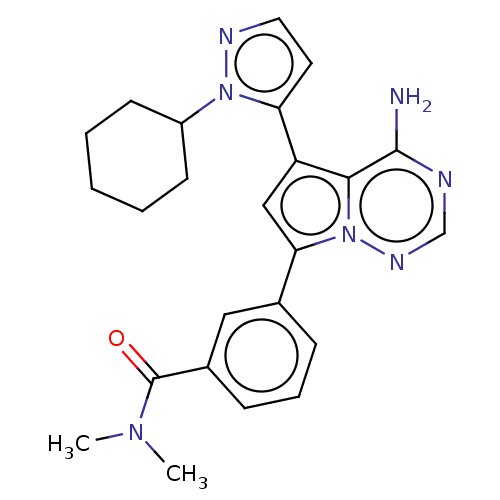
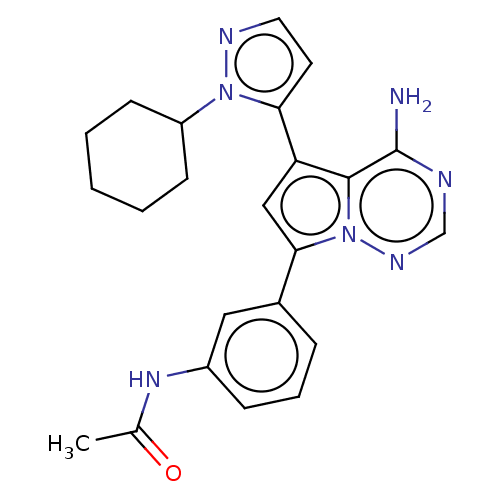
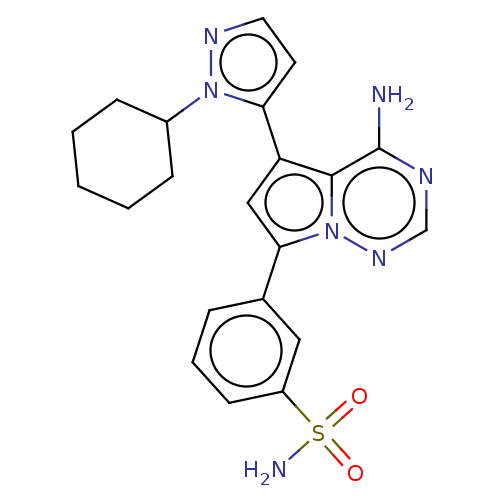
Report error Found 102 Enz. Inhib. hit(s) with all data for entry = 50009890
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of PI3Kdelta in BCR-stimulated human B cells assessed as suppression of CD86 expression after 18 hrs by FACS analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:PS as substrate incubated for 3 hrs by ADP-Glo assayMore data for this Ligand-Target Pair