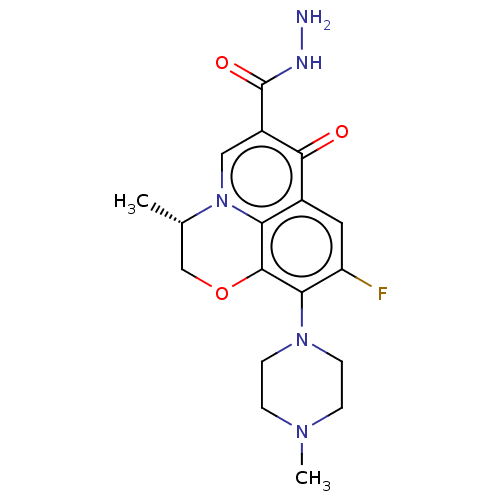
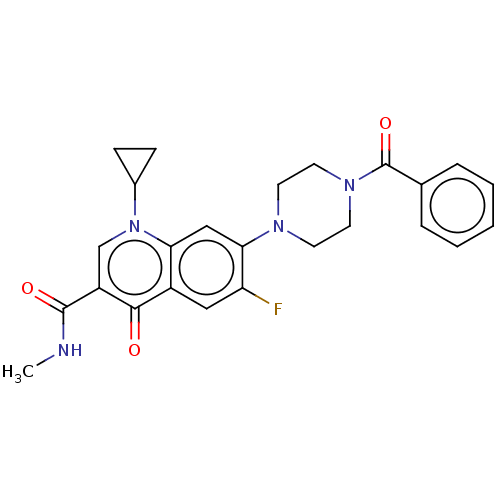
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 7822
Affinity DataIC50: 1.22E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.29E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.08E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.26E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.89E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.92E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.17E+3nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.89E+4nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.93E+4nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.72E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.12E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.12E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.49E+5nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.41E+6nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+6nMAssay Description:The assay mixture, containing 50 μl (2 mg/ml) of enzyme and 100 μl of different concentration of the tested agents, was added to 850 μ...More data for this Ligand-Target Pair

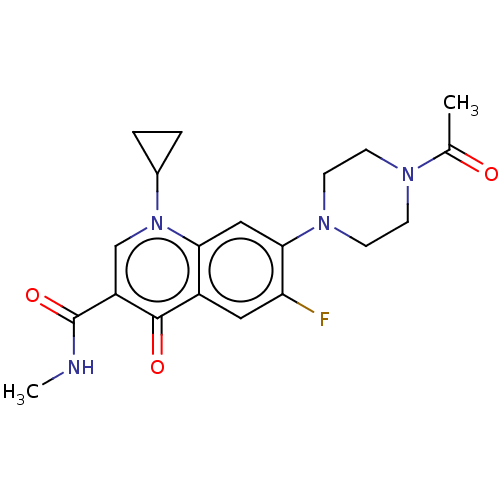









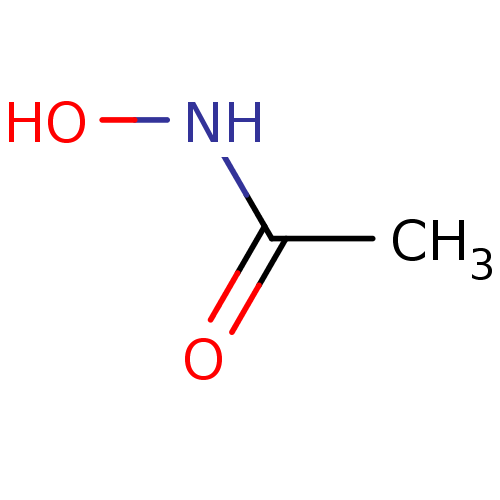
 3D Structure (crystal)
3D Structure (crystal)