Report error Found 72 Enz. Inhib. hit(s) with all data for entry = 50012203
Affinity DataIC50: 13nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 58nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 75nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 76nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 88nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 101nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 102nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 103nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 113nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 113nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 132nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 164nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 167nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 172nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 191nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 195nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 196nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 198nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 201nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 216nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 217nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 222nMAssay Description:Displacement of FITC-geldanamycin from N-terminal HSP90alpha (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 232nMAssay Description:Displacement of FITC-geldanamycin from N-terminal GRP94 (unknown origin) after 30 mins by fluorescence polarization competitive binding assayMore data for this Ligand-Target Pair
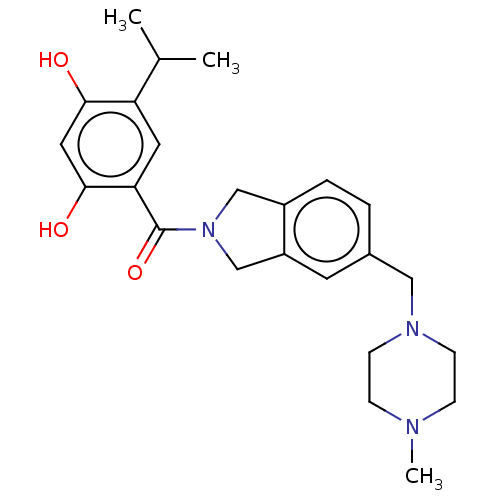
 3D Structure (crystal)
3D Structure (crystal)