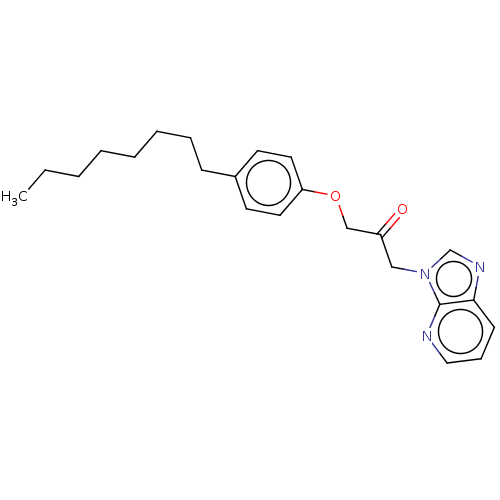
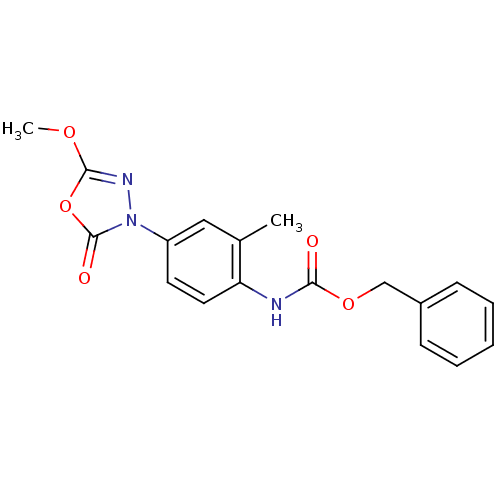
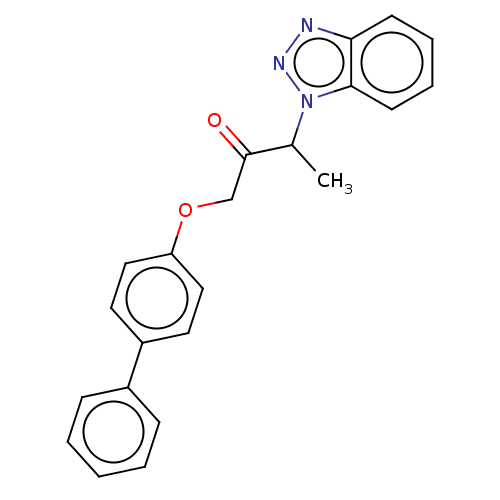
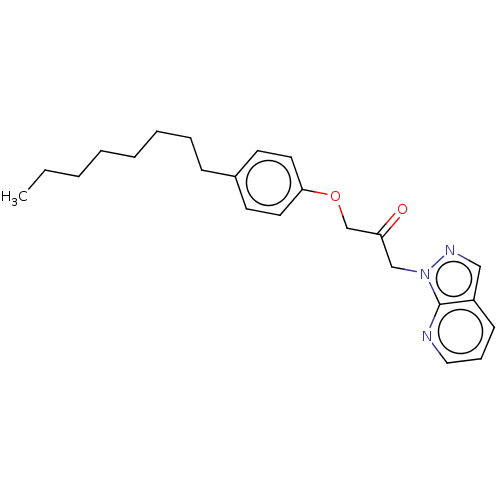
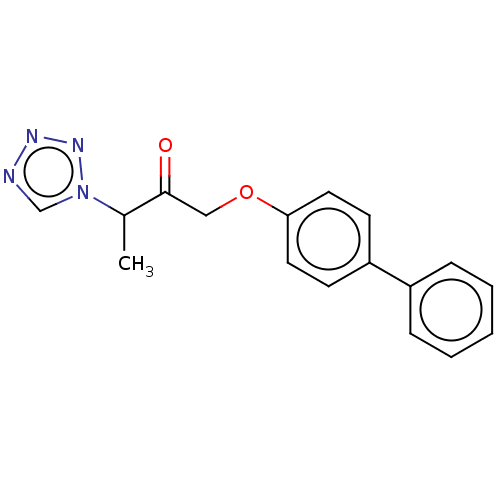
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 50048198
Affinity DataIC50: 2.90nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 93nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of recombinant human MAGL using 1,3-dihydroxypropan-2-yl 4-pyren-1-yl-butanoate as substrate after 45 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 620nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 840nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of FAAH in rat brain microsomes using N-(2-hydroxyethyl)-4-pyren-1-ylbutanamide as substrate after 60 mins by fluorescence-based reversed ...More data for this Ligand-Target Pair