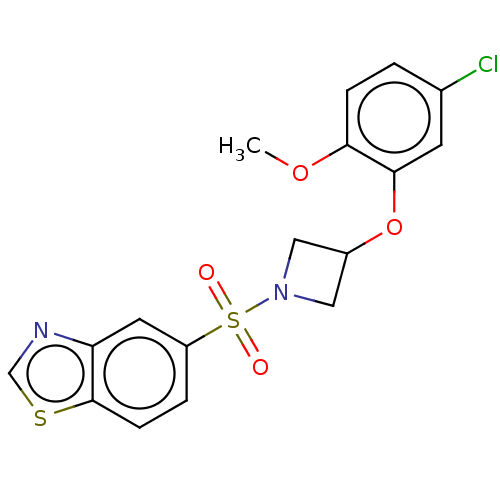
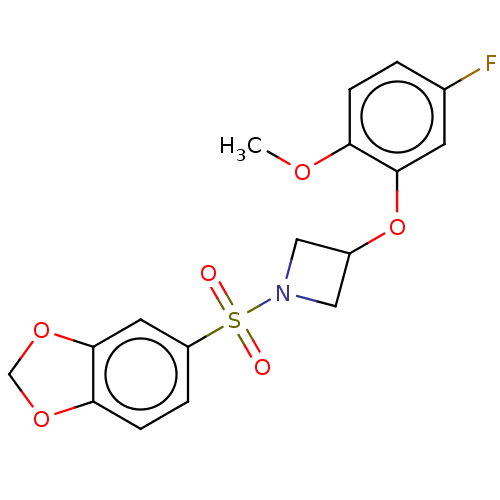
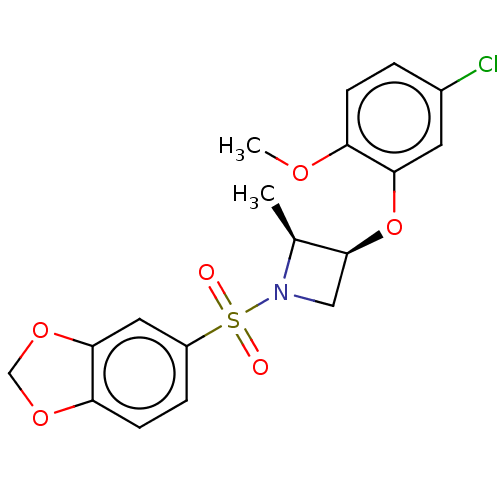
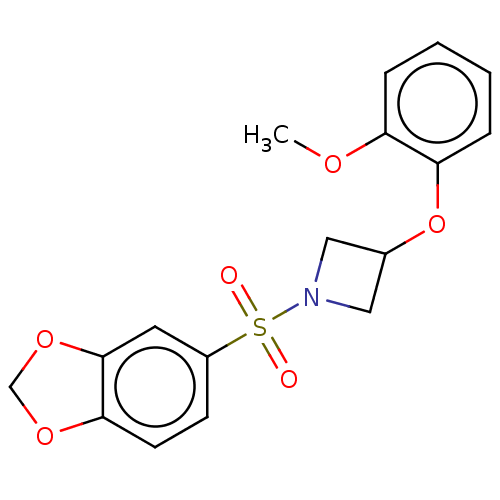
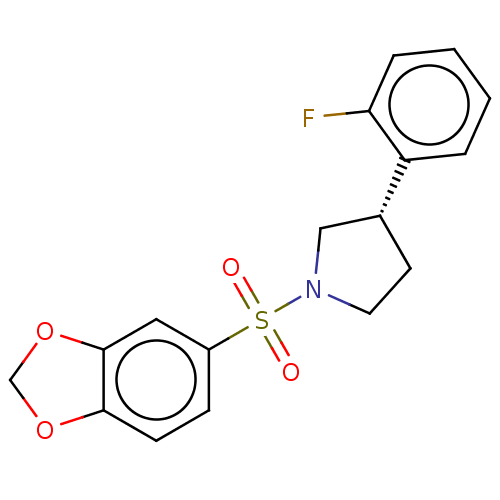
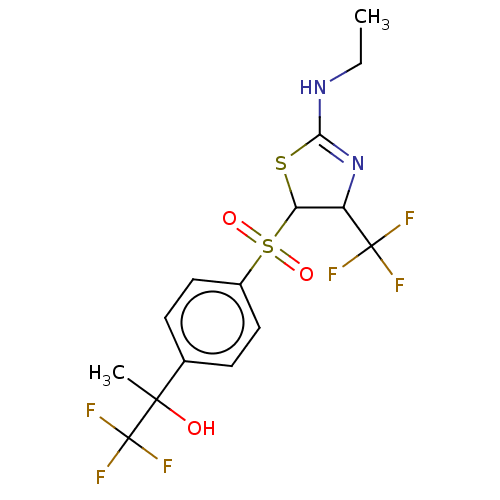
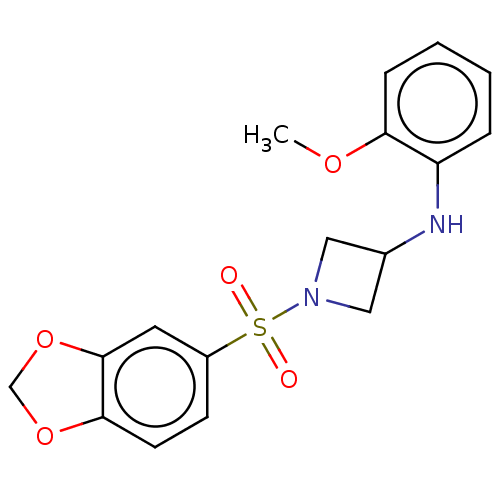
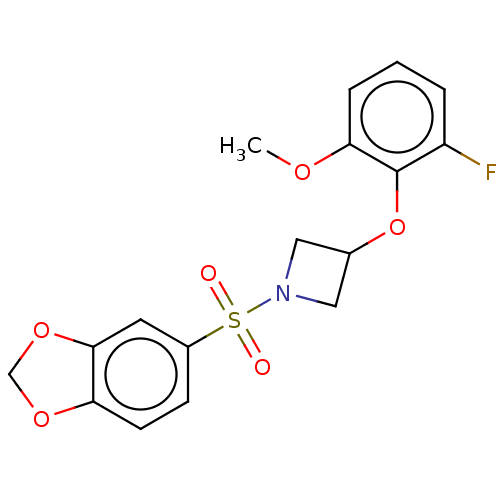
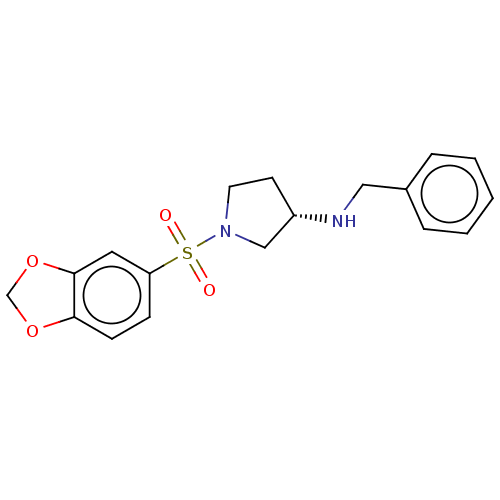
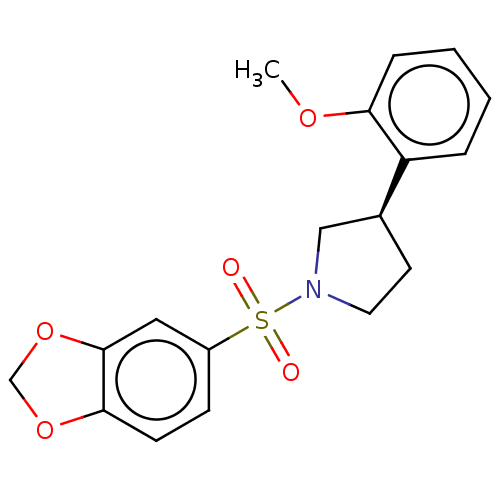
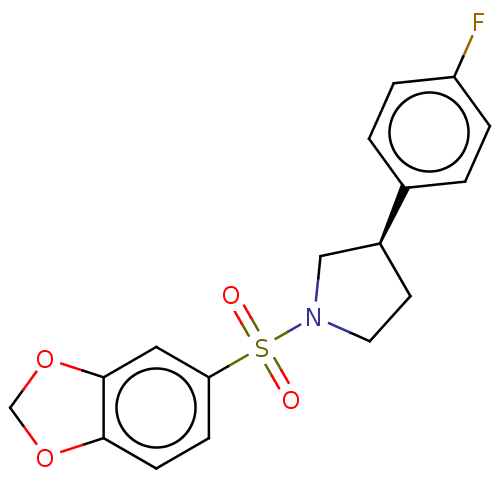
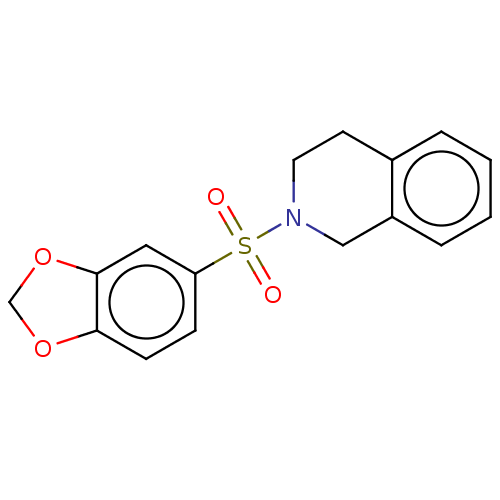
Report error Found 186 Enz. Inhib. hit(s) with all data for entry = 50002884
Affinity DataKd: 70nMAssay Description:Binding affinity to human GlyRalpha3cryst by SPR methodMore data for this Ligand-Target Pair
Affinity DataEC50: 300nMAssay Description:Positive allosteric modulation of human GlyRalpha1beta assessed as increase in glycine-induced chloride ion flux by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 300nMAssay Description:Binding affinity to human GlyRalpha3cryst by SPR methodMore data for this Ligand-Target Pair
Affinity DataEC50: 370nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 500nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 600nMAssay Description:Positive allosteric modulation of human GlyRalpha1beta assessed as increase in glycine-induced chloride ion flux by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 700nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 900nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 900nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.21E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.10E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.30E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.30E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.50E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.70E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.10E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.30E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.70E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.70E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.20E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.20E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.20E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.40E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 6.10E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 7.12E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 8.70E+3nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.24E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Positive allosteric modulation of human GlyRalpha3beta expressed in HEK293T cells assessed as increase in glycine-induced chloride ion flux measured ...More data for this Ligand-Target Pair