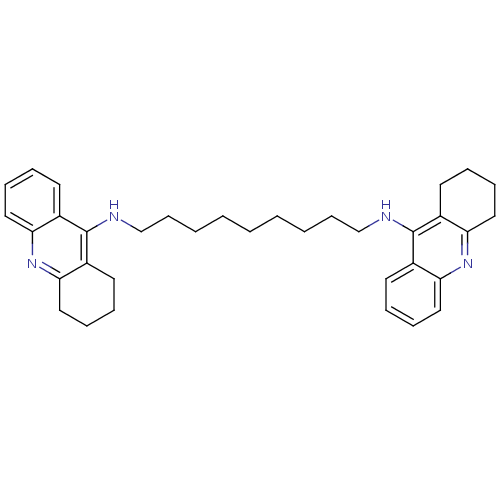
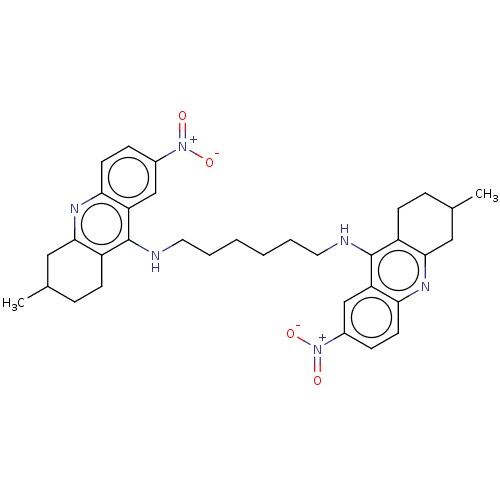
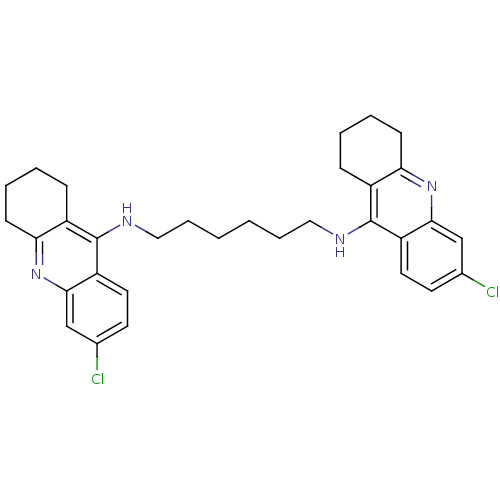
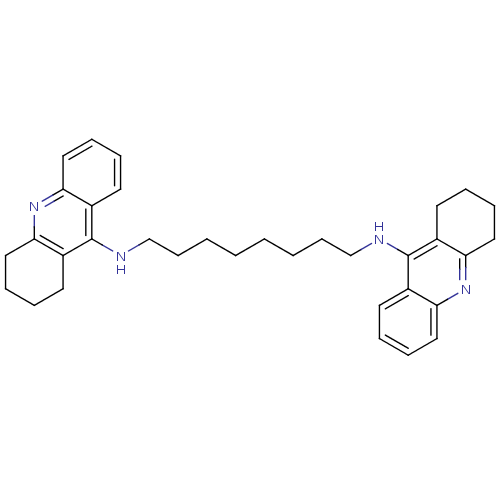
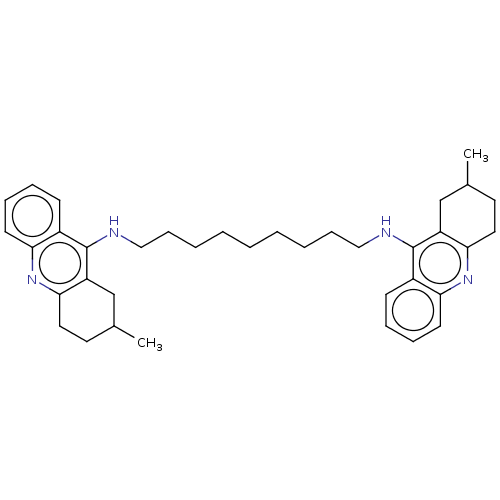
Report error Found 9 Enz. Inhib. hit(s) with all data for entry = 50001095
TargetTrypanothione reductase(Trypanosoma cruzi)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of Trypanosoma cruzi trypanothione reductase using varying levels of trypanothione disulfide as substrate by Lineweaver--burk ...More data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 7.40E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.33E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.69E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.81E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.94E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair
TargetCysteine protease(Trypanosoma brucei rhodesiense)
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Julius-Maximilians-University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate measured for 10 mins by spectrofluorometric analysisMore data for this Ligand-Target Pair