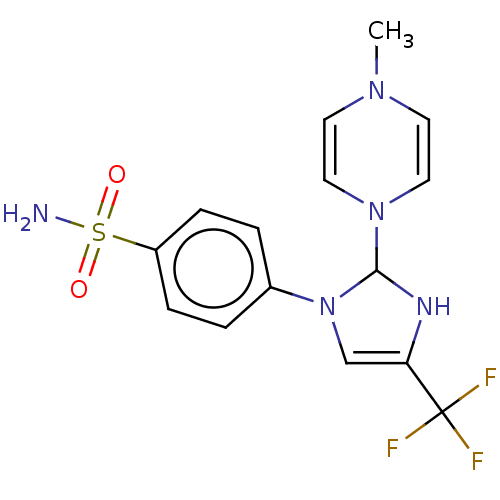
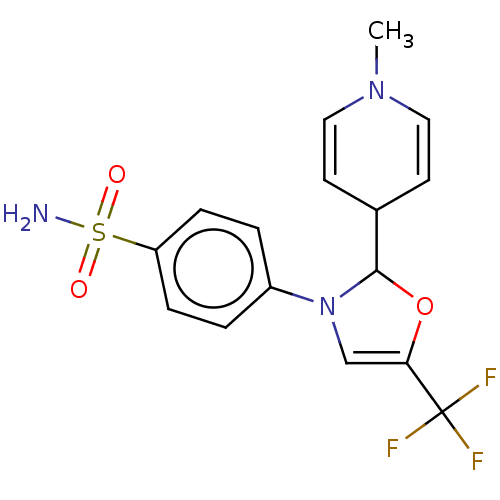
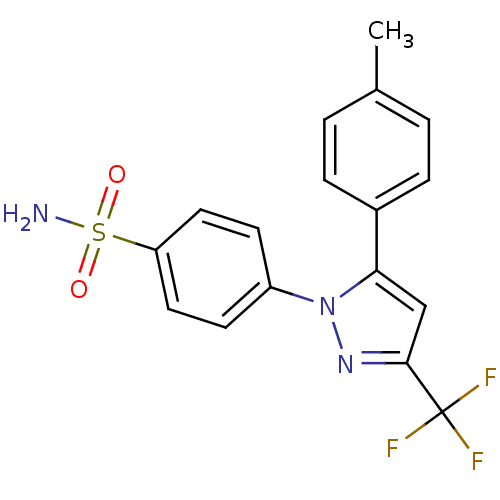
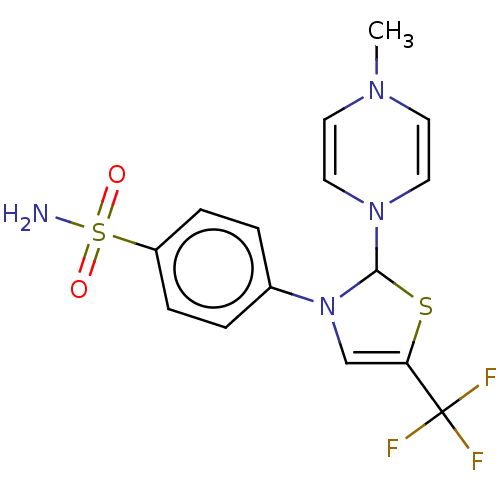
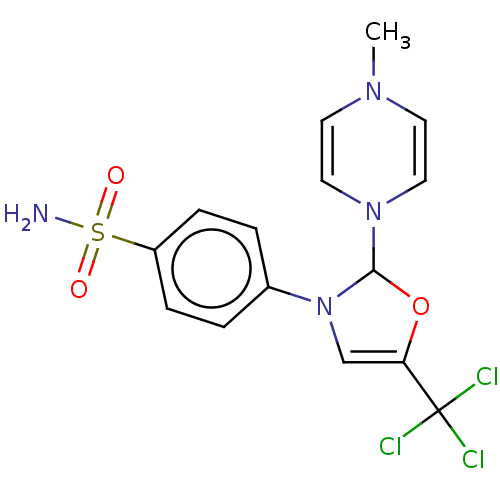
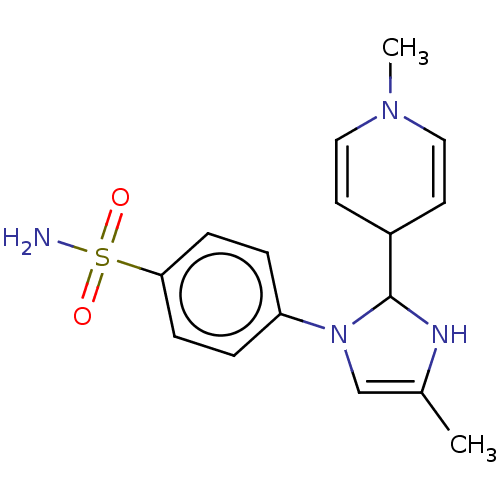
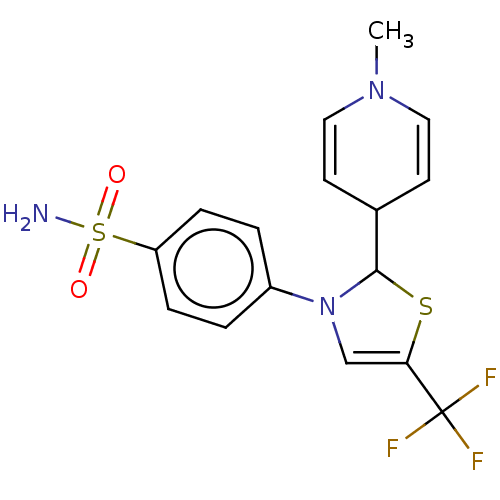
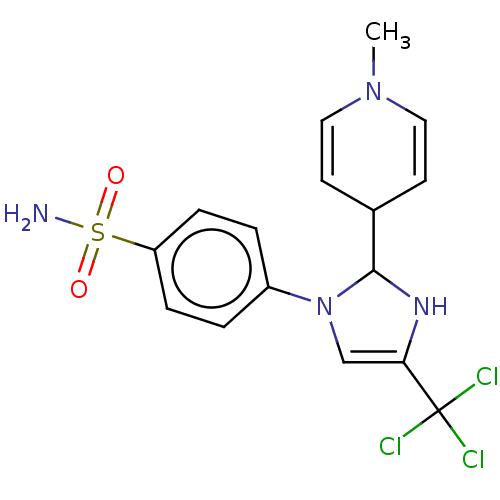
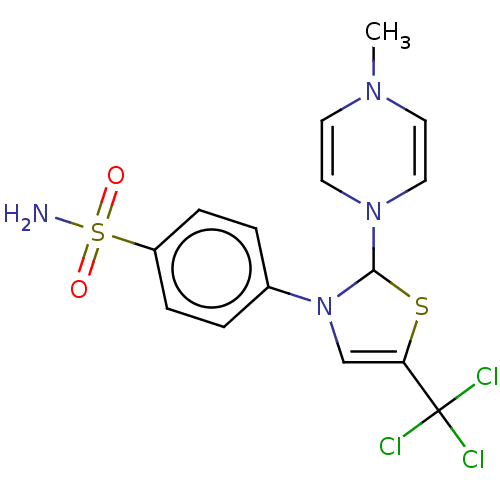
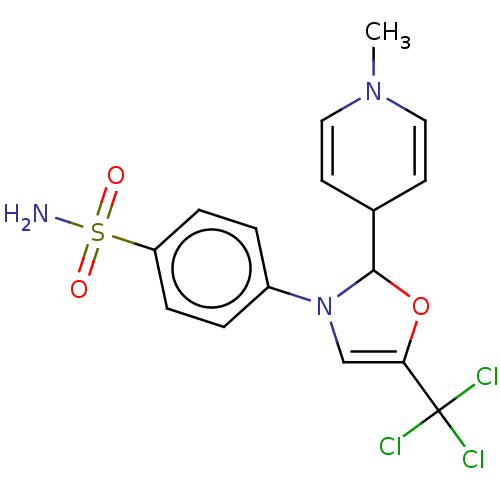
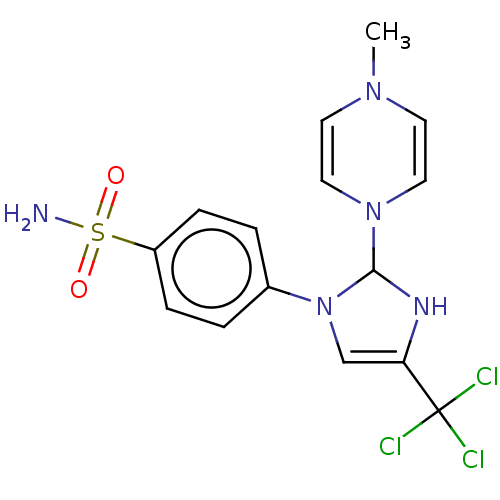
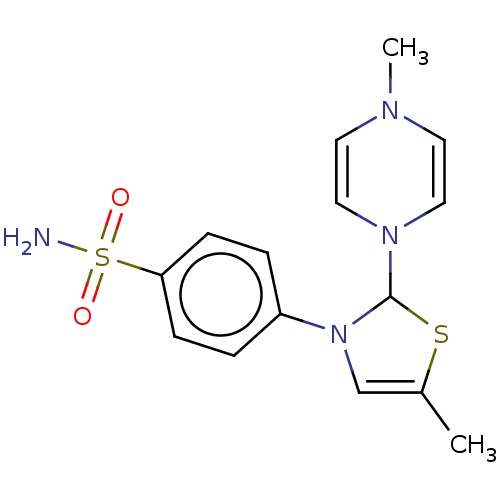
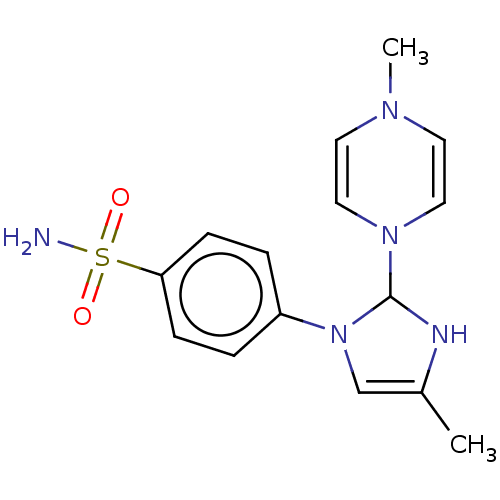
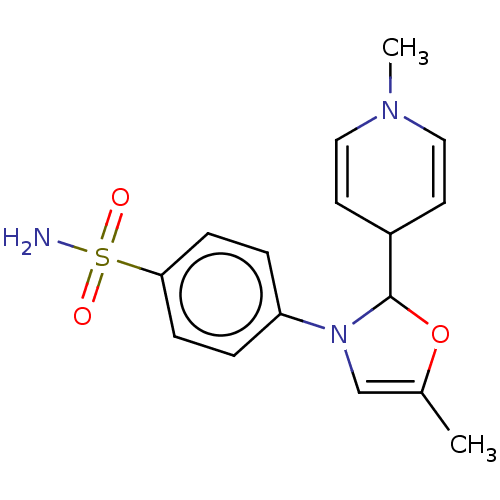
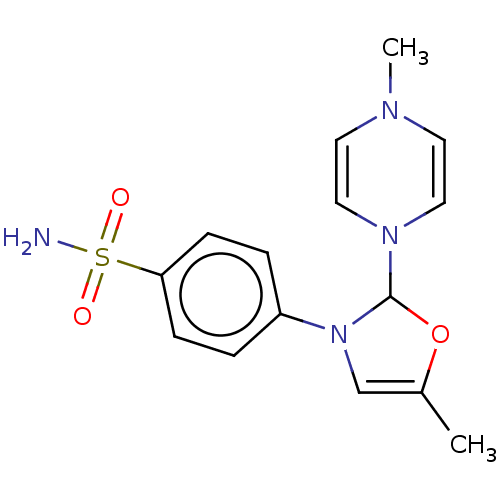
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50000796
Affinity DataIC50: 49nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 79nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 99nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 103nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+3nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of COX-2 in rat peritoneal macrophages assessed as reduction in PGE2 production using radiolabelled-arachidonic acid as substrate pretreat...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 8.90E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 9.50E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 9.80E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90E+4nMAssay Description:Inhibition of COX-1 in rat peritoneal macrophages assessed as reduction in [125I]-6-Keto-PGF1alpha production using arachidonic acid as substrate pre...More data for this Ligand-Target Pair