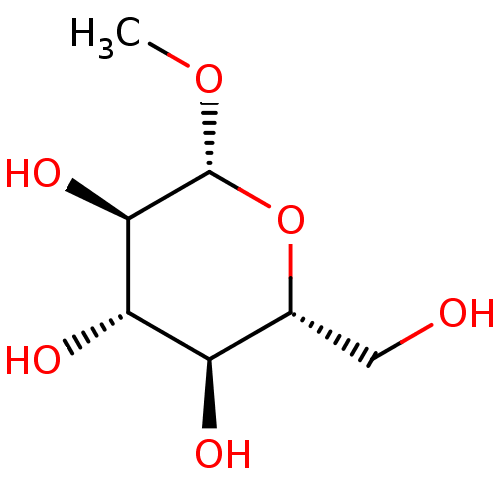
Report error Found 8 Enz. Inhib. hit(s) with all data for entry = 50002186
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 2.80E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 3.40E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 3.70E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 5.00E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 5.20E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 5.70E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 6.50E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair
TargetLipopolysaccharide heptosyltransferase 1(Escherichia coli (strain K12))
Wesleyan University
Curated by ChEMBL
Wesleyan University
Curated by ChEMBL
Affinity DataKi: 6.80E+5nMAssay Description:Inhibition of Escherichia coli Heptosyltransferase I assessed as reduction in ADP release using ODLA and ADP-heptose substrates in presence of phosph...More data for this Ligand-Target Pair